

You can:
| Name | KiSS-1 receptor |
|---|---|
| Species | Rattus norvegicus (Rat) |
| Gene | Kiss1r |
| Synonym | Metastin receptor Kisspeptins receptor kisspeptin receptor KiSS1-derived peptide receptor KiSS-1R [ Show all ] |
| Disease | N/A for non-human GPCRs |
| Length | 396 |
| Amino acid sequence | MAAEATLGPNVSWWAPSNASGCPGCGVNASDGPGSAPRPLDAWLVPLFFAALMLLGLVGNSLVIFVICRHKHMQTVTNFYIANLAATDVTFLLCCVPFTALLYPLPTWVLGDFMCKFVNYIQQVSVQATCATLTAMSVDRWYVTVFPLRALHRRTPRLALTVSLSIWVGSAAVSAPVLALHRLSPGPHTYCSEAFPSRALERAFALYNLLALYLLPLLATCACYGAMLRHLGRAAVRPAPTDGALQGQLLAQRAGAVRTKVSRLVAAVVLLFAACWGPIQLFLVLQALGPSGAWHPRSYAAYALKIWAHCMSYSNSALNPLLYAFLGSHFRQAFCRVCPCGPQRQRRPHASAHSDRAAPHSVPHSRAAHPVRVRTPEPGNPVRRSPSVQDEHTAPL |
| UniProt | Q924U1 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL1169599 |
| IUPHAR | 266 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 514 | 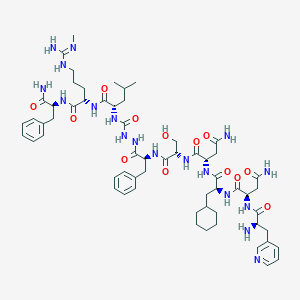 CHEMBL3085807 CHEMBL3085807 | C60H88N18O13 | 1269.48 | 16 / 17 | -1.1 | No |
| 441716 | 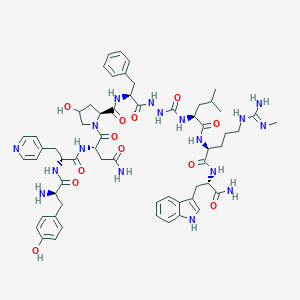 CHEMBL3314214 CHEMBL3314214 | C60H79N17O12 | 1230.4 | 15 / 16 | -0.3 | No |
| 536076 | 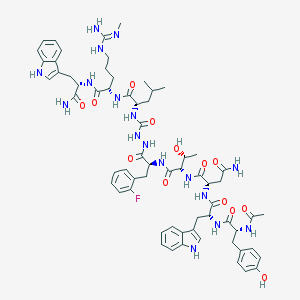 CHEMBL3928465 CHEMBL3928465 | C64H82FN17O13 | 1316.46 | 15 / 18 | 1.3 | No |
| 548025 | 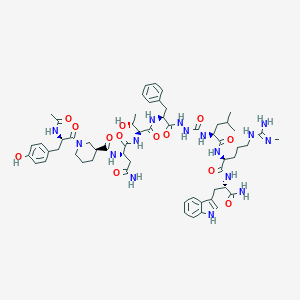 CHEMBL3982675 CHEMBL3982675 | C59H82N16O13 | 1223.4 | 14 / 16 | -0.3 | No |
| 442502 | 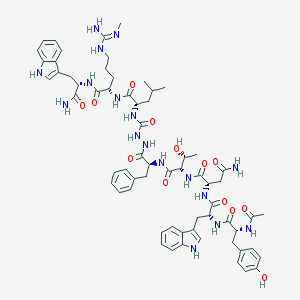 CHEMBL3314224 CHEMBL3314224 | C64H83N17O13 | 1298.47 | 14 / 18 | 1.2 | No |
| 548125 | 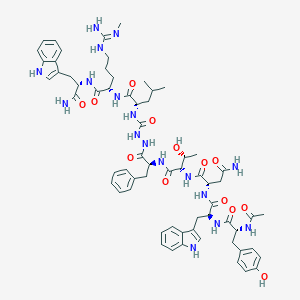 CHEMBL3985112 CHEMBL3985112 | C64H83N17O13 | 1298.47 | 14 / 18 | 1.2 | No |
| 536712 | 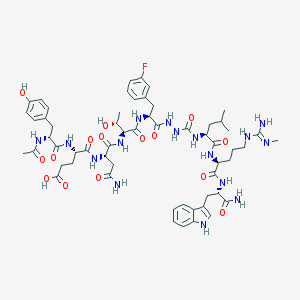 CHEMBL3977314 CHEMBL3977314 | C58H79FN16O15 | 1259.36 | 17 / 18 | -1.0 | No |
| 442826 | 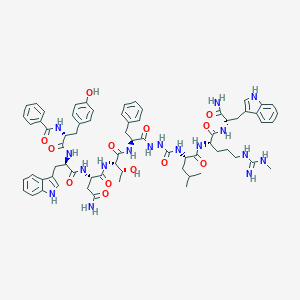 CHEMBL3314227 CHEMBL3314227 | C69H85N17O13 | 1360.55 | 14 / 19 | 3.1 | No |
| 36835 | 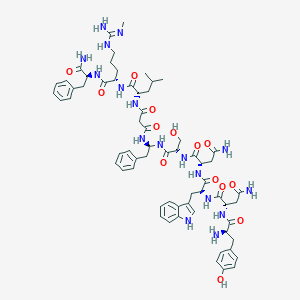 CHEMBL2151645 CHEMBL2151645 | C64H85N17O14 | 1316.49 | 16 / 18 | -1.2 | No |
| 536953 | 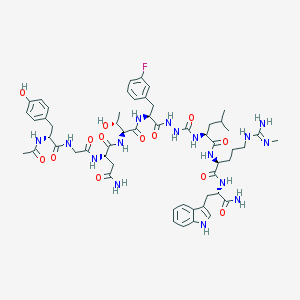 CHEMBL3939287 CHEMBL3939287 | C55H75FN16O13 | 1187.3 | 15 / 17 | -0.9 | No |
| 536983 | 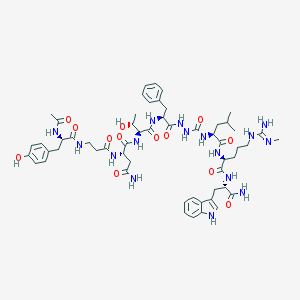 CHEMBL3985323 CHEMBL3985323 | C56H78N16O13 | 1183.34 | 14 / 17 | -1.1 | No |
| 40351 | 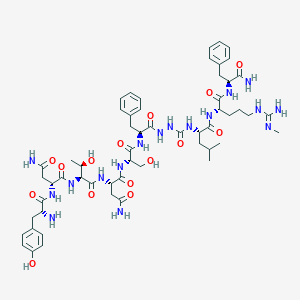 CHEMBL3087930 CHEMBL3087930 | C56H81N17O15 | 1232.37 | 17 / 19 | -3.8 | No |
| 40354 | 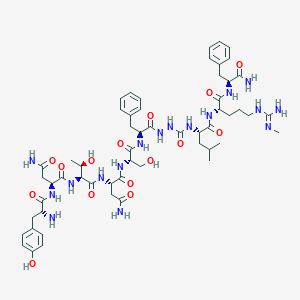 CHEMBL2151648 CHEMBL2151648 | C56H81N17O15 | 1232.37 | 17 / 19 | -3.8 | No |
| 537090 | 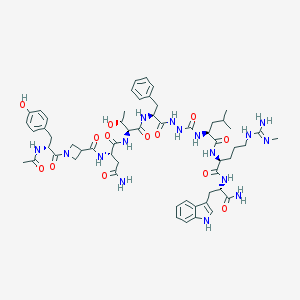 CHEMBL3931718 CHEMBL3931718 | C57H78N16O13 | 1195.35 | 14 / 16 | -1.1 | No |
| 443575 | 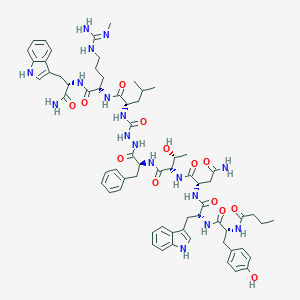 CHEMBL3314225 CHEMBL3314225 | C66H87N17O13 | 1326.53 | 14 / 18 | 2.0 | No |
| 62322 | 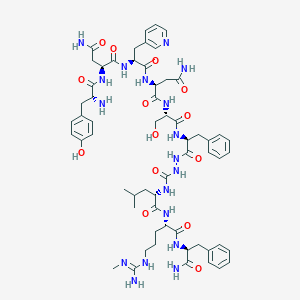 CHEMBL2151651 CHEMBL2151651 | C60H82N18O14 | 1279.43 | 17 / 18 | -2.6 | No |
| 444447 | 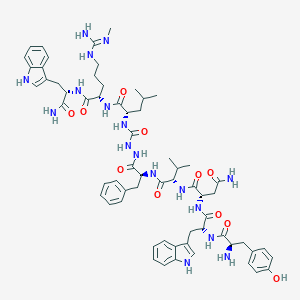 CHEMBL3314217 CHEMBL3314217 | C63H83N17O11 | 1254.46 | 13 / 17 | 2.5 | No |
| 444602 | 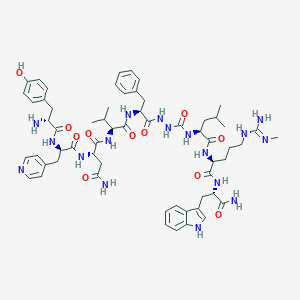 CHEMBL3314209 CHEMBL3314209 | C60H81N17O11 | 1216.42 | 14 / 16 | 1.3 | No |
| 548938 | 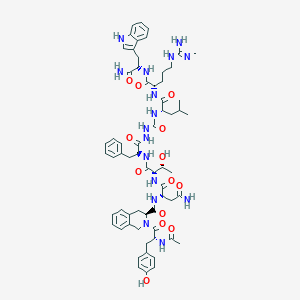 CHEMBL3964403 CHEMBL3964403 | C63H82N16O13 | 1271.45 | 14 / 16 | 0.8 | No |
| 548989 | 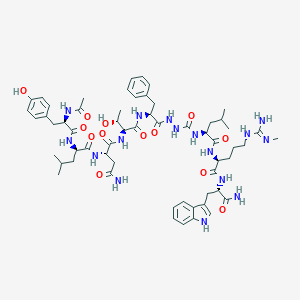 CHEMBL3976038 CHEMBL3976038 | C59H84N16O13 | 1225.42 | 14 / 17 | 0.8 | No |
| 445502 | 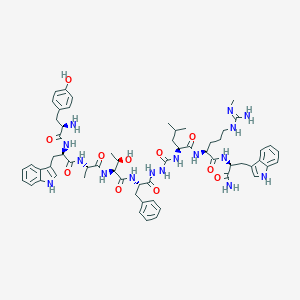 CHEMBL3314221 CHEMBL3314221 | C61H80N16O11 | 1213.41 | 13 / 17 | 2.4 | No |
| 445523 | 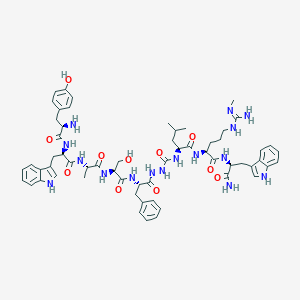 CHEMBL3314220 CHEMBL3314220 | C60H78N16O11 | 1199.39 | 13 / 17 | 2.0 | No |
| 445635 | 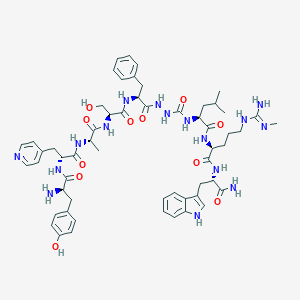 CHEMBL3314218 CHEMBL3314218 | C57H76N16O11 | 1161.34 | 14 / 16 | 0.8 | No |
| 445748 | 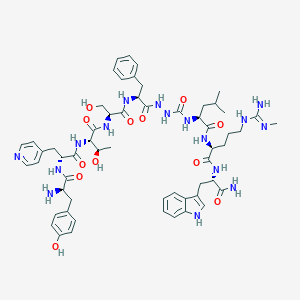 CHEMBL3314219 CHEMBL3314219 | C58H78N16O12 | 1191.36 | 15 / 17 | 0.2 | No |
| 103484 | 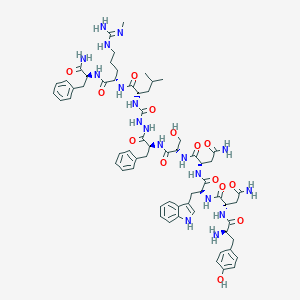 CHEMBL2151646 CHEMBL2151646 | C63H84N18O14 | 1317.48 | 16 / 19 | -1.4 | No |
| 103486 | 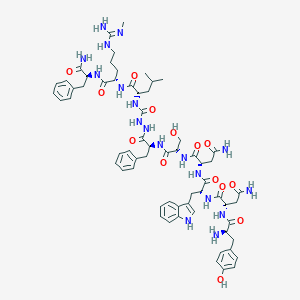 CHEMBL2151654 CHEMBL2151654 | C63H84N18O14 | 1317.48 | 16 / 19 | -1.4 | No |
| 103488 | 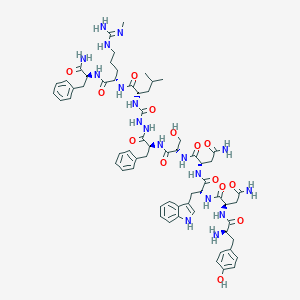 CHEMBL3085804 CHEMBL3085804 | C63H84N18O14 | 1317.48 | 16 / 19 | -1.4 | No |
| 103489 | 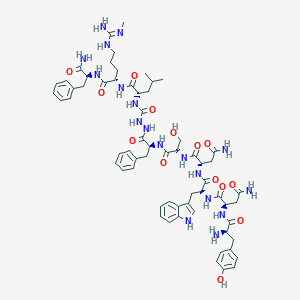 CHEMBL3087925 CHEMBL3087925 | C63H84N18O14 | 1317.48 | 16 / 19 | -1.4 | No |
| 561238 | 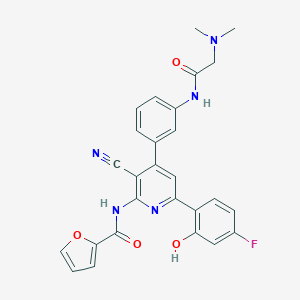 CHEMBL1171977 CHEMBL1171977 | C27H22FN5O4 | 499.502 | 8 / 3 | 4.0 | Yes |
| 133224 | 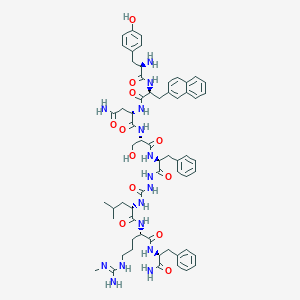 CHEMBL3086282 CHEMBL3086282 | C61H79N15O12 | 1214.4 | 14 / 16 | 1.5 | No |
| 147150 | 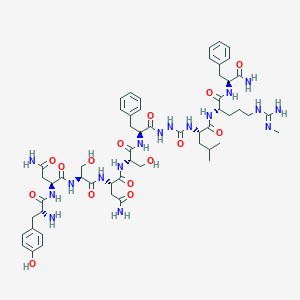 CHEMBL2151647 CHEMBL2151647 | C55H79N17O15 | 1218.34 | 17 / 19 | -4.2 | No |
| 539771 | 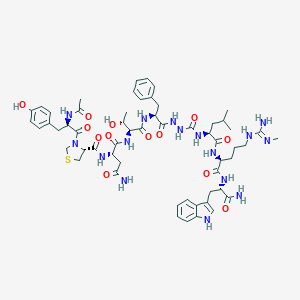 CHEMBL3956072 CHEMBL3956072 | C57H78N16O13S | 1227.41 | 15 / 16 | -0.2 | No |
| 561974 | 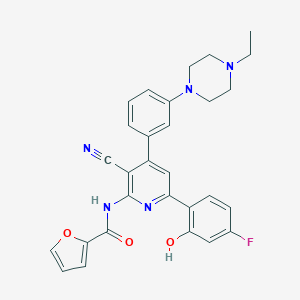 CHEMBL1173008 CHEMBL1173008 | C29H26FN5O3 | 511.557 | 8 / 2 | 5.0 | No |
| 539918 | 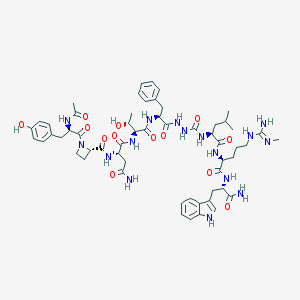 CHEMBL3967058 CHEMBL3967058 | C57H78N16O13 | 1195.35 | 14 / 16 | -0.6 | No |
| 158936 | 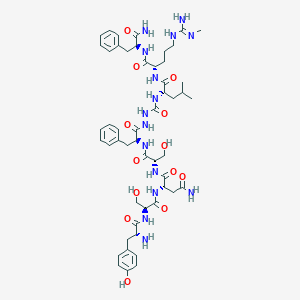 CHEMBL3085810 CHEMBL3085810 | C51H73N15O13 | 1104.24 | 15 / 17 | -2.4 | No |
| 540147 | 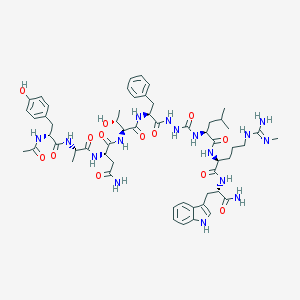 CHEMBL3915251 CHEMBL3915251 | C56H78N16O13 | 1183.34 | 14 / 17 | -0.6 | No |
| 549937 | 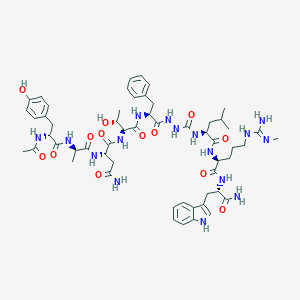 CHEMBL3904889 CHEMBL3904889 | C56H78N16O13 | 1183.34 | 14 / 17 | -0.6 | No |
| 540214 | 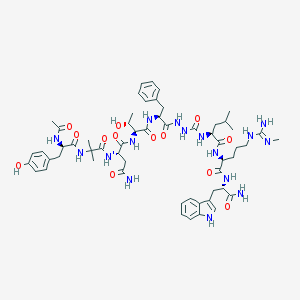 CHEMBL3964945 CHEMBL3964945 | C57H80N16O13 | 1197.37 | 14 / 17 | -0.4 | No |
| 448232 | 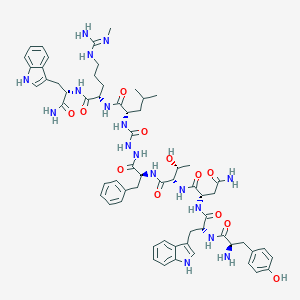 CHEMBL3314216 CHEMBL3314216 | C62H81N17O12 | 1256.44 | 14 / 18 | 0.9 | No |
| 167307 | 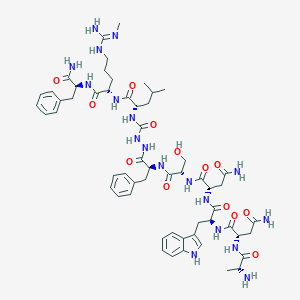 CHEMBL3085805 CHEMBL3085805 | C57H80N18O13 | 1225.38 | 15 / 18 | -2.7 | No |
| 540376 | 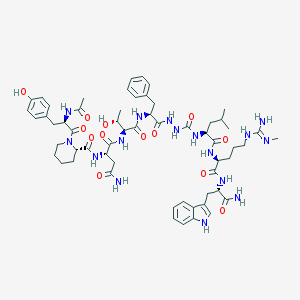 CHEMBL3894943 CHEMBL3894943 | C59H82N16O13 | 1223.4 | 14 / 16 | 0.1 | No |
| 448491 | 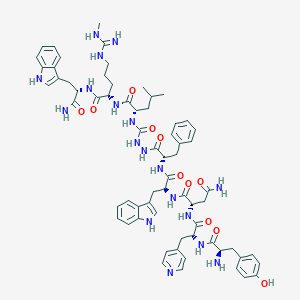 CHEMBL3314212 CHEMBL3314212 | C66H82N18O11 | 1303.5 | 14 / 18 | 2.4 | No |
| 174046 | 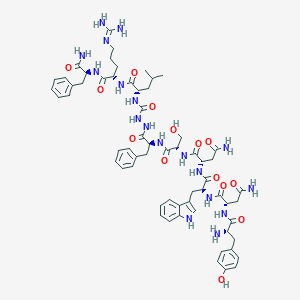 CHEMBL3087793 CHEMBL3087793 | C62H82N18O14 | 1303.45 | 16 / 19 | -1.9 | No |
| 176245 | 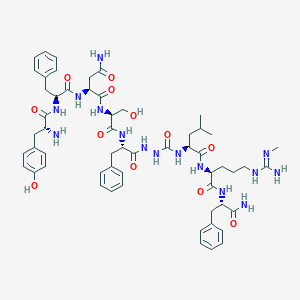 CHEMBL3086281 CHEMBL3086281 | C57H77N15O12 | 1164.34 | 14 / 16 | 0.2 | No |
| 178500 | 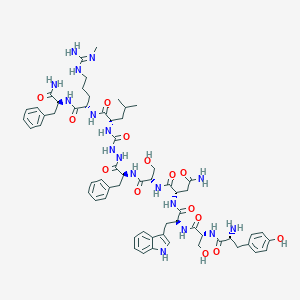 CHEMBL3087926 CHEMBL3087926 | C62H83N17O14 | 1290.45 | 16 / 19 | -1.0 | No |
| 563096 | 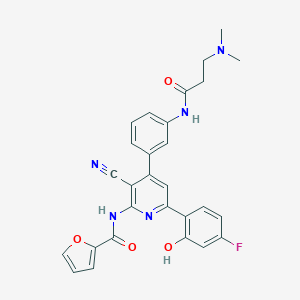 CHEMBL1169902 CHEMBL1169902 | C28H24FN5O4 | 513.529 | 8 / 3 | 3.9 | No |
| 541000 | 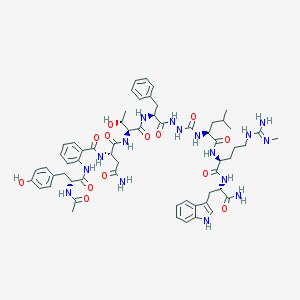 CHEMBL3938759 CHEMBL3938759 | C60H78N16O13 | 1231.38 | 14 / 17 | 1.1 | No |
| 449424 | 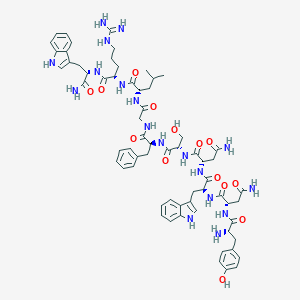 CHEMBL3315315 CHEMBL3315315 | C65H84N18O14 | 1341.5 | 16 / 20 | -1.4 | No |
| 201328 | 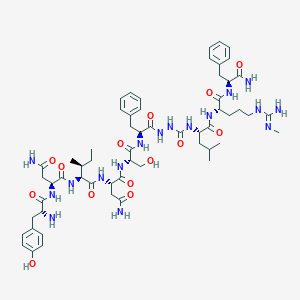 CHEMBL2151649 CHEMBL2151649 | C58H85N17O14 | 1244.42 | 16 / 18 | -1.8 | No |
| 541657 | 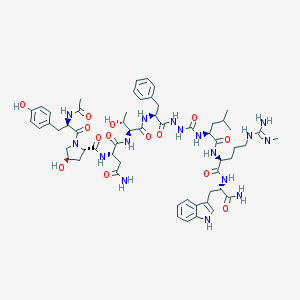 UNII-YO029HR229 UNII-YO029HR229 | C58H80N16O14 | 1225.38 | 15 / 17 | -1.2 | No |
| 217834 | 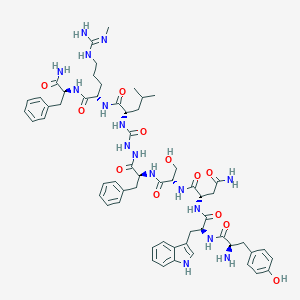 CHEMBL3085809 CHEMBL3085809 | C59H78N16O12 | 1203.37 | 14 / 17 | 0.3 | No |
| 217836 | 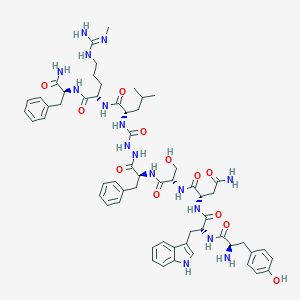 CHEMBL3086283 CHEMBL3086283 | C59H78N16O12 | 1203.37 | 14 / 17 | 0.3 | No |
| 550599 | 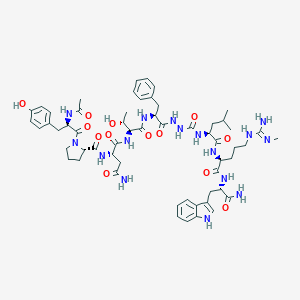 CHEMBL3911692 CHEMBL3911692 | C58H80N16O13 | 1209.38 | 14 / 16 | -0.3 | No |
| 550600 | 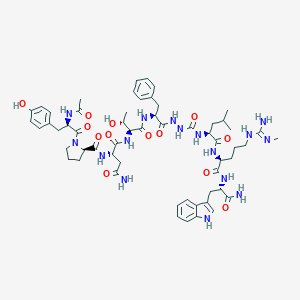 CHEMBL3966187 CHEMBL3966187 | C58H80N16O13 | 1209.38 | 14 / 16 | -0.3 | No |
| 542107 | 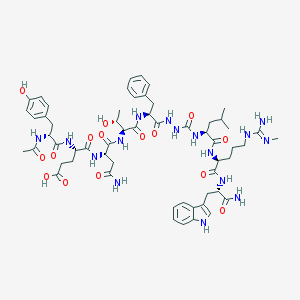 CHEMBL3958495 CHEMBL3958495 | C58H80N16O15 | 1241.38 | 16 / 18 | -1.1 | No |
| 550757 | 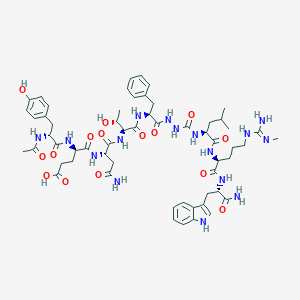 CHEMBL3984058 CHEMBL3984058 | C58H80N16O15 | 1241.38 | 16 / 18 | -1.1 | No |
| 237411 | 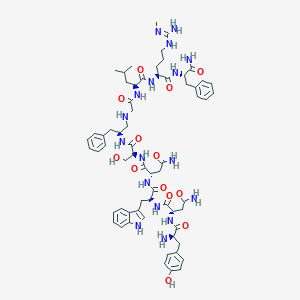 CHEMBL2151644 CHEMBL2151644 | C64H87N17O13 | 1302.51 | 16 / 18 | -1.2 | No |
| 451492 | 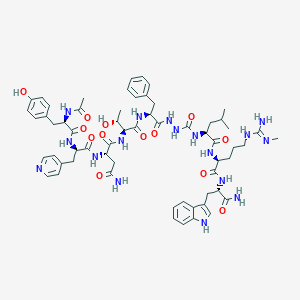 CHEMBL3314228 CHEMBL3314228 | C61H81N17O13 | 1260.43 | 15 / 17 | 0.0 | No |
| 451520 | 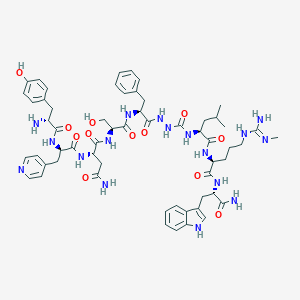 CHEMBL3314207 CHEMBL3314207 | C58H77N17O12 | 1204.36 | 15 / 17 | -0.7 | No |
| 451836 | 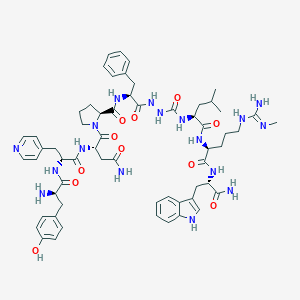 CHEMBL3314213 CHEMBL3314213 | C60H79N17O11 | 1214.4 | 14 / 15 | 0.6 | No |
| 265525 | 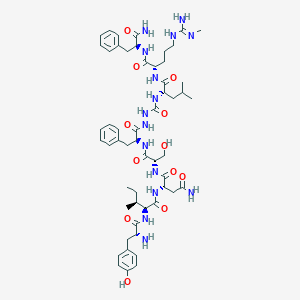 CHEMBL3086280 CHEMBL3086280 | C54H79N15O12 | 1130.32 | 14 / 16 | -0.1 | No |
| 543087 | 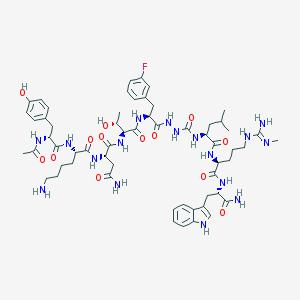 CHEMBL3949224 CHEMBL3949224 | C59H84FN17O13 | 1258.43 | 16 / 18 | -0.7 | No |
| 271174 | 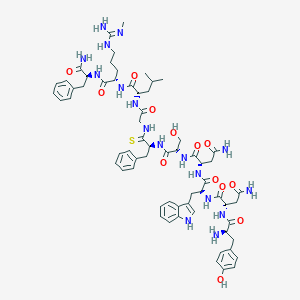 CHEMBL2151643 CHEMBL2151643 | C64H85N17O13S | 1332.55 | 16 / 18 | -0.8 | No |
| 543346 | 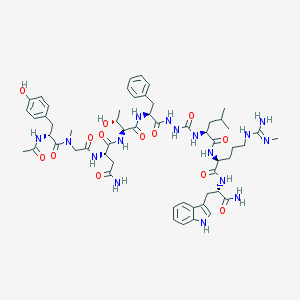 CHEMBL3956626 CHEMBL3956626 | C56H78N16O13 | 1183.34 | 14 / 16 | -0.8 | No |
| 281556 | 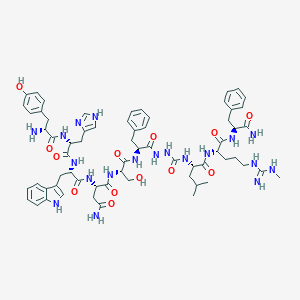 CID 72711590 CID 72711590 | C65H85N19O13 | 1340.52 | 16 / 20 | 0.2 | No |
| 452800 | 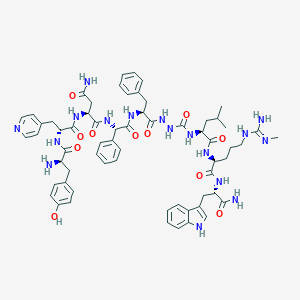 CHEMBL3314210 CHEMBL3314210 | C63H79N17O11 | 1250.43 | 14 / 16 | 1.6 | No |
| 285672 | 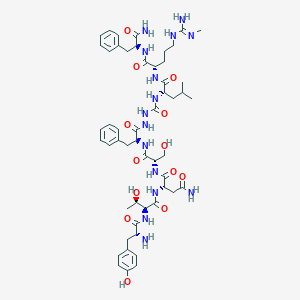 CHEMBL3085811 CHEMBL3085811 | C52H75N15O13 | 1118.26 | 15 / 17 | -2.0 | No |
| 297196 | 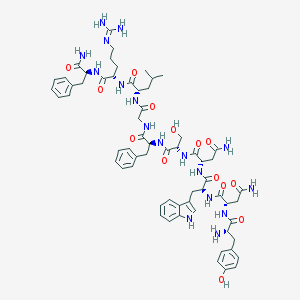 Kisspeptin-10 Kisspeptin-10 | C63H83N17O14 | 1302.46 | 16 / 18 | -1.9 | No |
| 544069 | 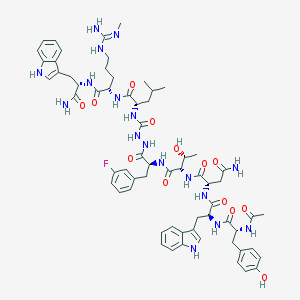 CHEMBL3949159 CHEMBL3949159 | C64H82FN17O13 | 1316.46 | 15 / 18 | 1.3 | No |
| 544071 | 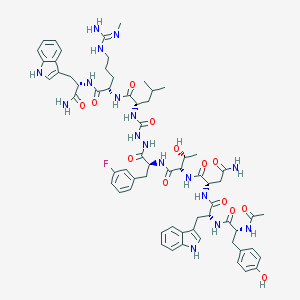 CHEMBL3930297 CHEMBL3930297 | C64H82FN17O13 | 1316.46 | 15 / 18 | 1.3 | No |
| 453661 | 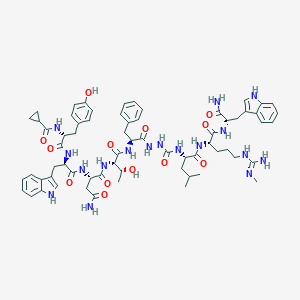 CHEMBL3314226 CHEMBL3314226 | C66H85N17O13 | 1324.51 | 14 / 18 | 1.6 | No |
| 566795 | 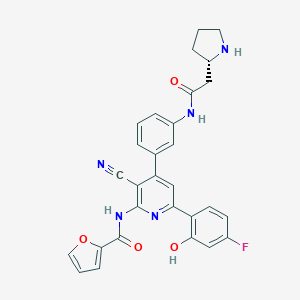 CHEMBL1170903 CHEMBL1170903 | C29H24FN5O4 | 525.54 | 8 / 4 | 4.0 | No |
| 566797 | 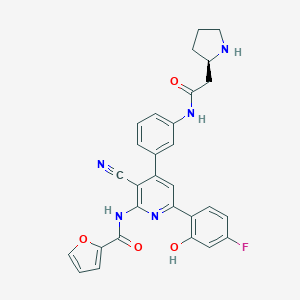 CHEMBL1170904 CHEMBL1170904 | C29H24FN5O4 | 525.54 | 8 / 4 | 4.0 | No |
| 544584 | 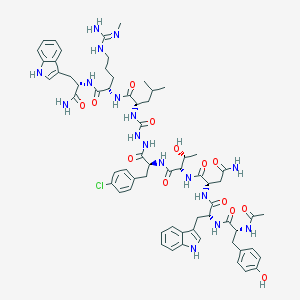 CHEMBL3921327 CHEMBL3921327 | C64H82ClN17O13 | 1332.92 | 14 / 18 | 1.8 | No |
| 551906 | 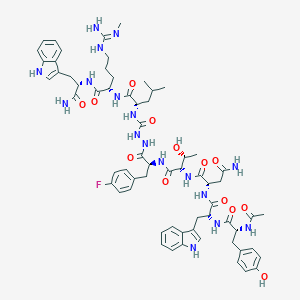 CHEMBL3957784 CHEMBL3957784 | C64H82FN17O13 | 1316.46 | 15 / 18 | 1.3 | No |
| 324484 | 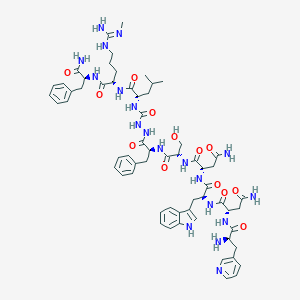 CHEMBL3085806 CHEMBL3085806 | C62H83N19O13 | 1302.47 | 16 / 18 | -2.1 | No |
| 567738 | 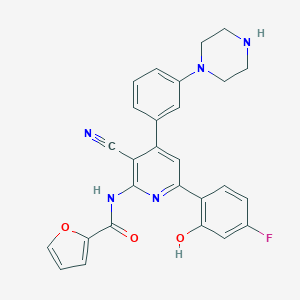 CHEMBL1173643 CHEMBL1173643 | C27H22FN5O3 | 483.503 | 8 / 3 | 4.2 | Yes |
| 338066 | 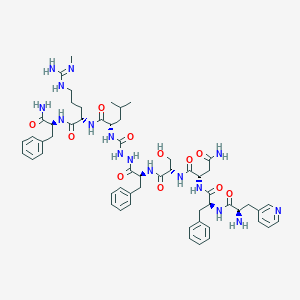 CHEMBL3086285 CHEMBL3086285 | C56H76N16O11 | 1149.32 | 14 / 15 | -0.5 | No |
| 455266 | 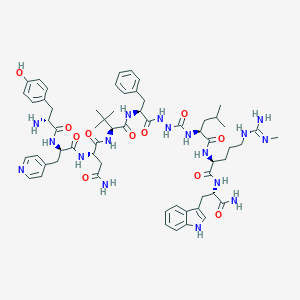 CHEMBL3314211 CHEMBL3314211 | C61H83N17O11 | 1230.44 | 14 / 16 | 1.7 | No |
| 552246 | 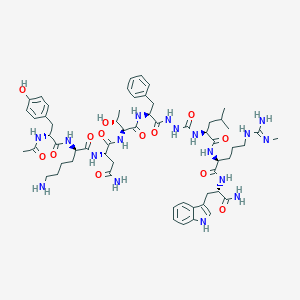 CHEMBL3914238 CHEMBL3914238 | C59H85N17O13 | 1240.44 | 15 / 18 | -0.8 | No |
| 545361 | 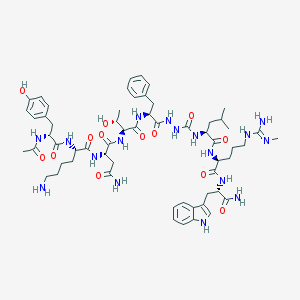 CHEMBL3956437 CHEMBL3956437 | C59H85N17O13 | 1240.44 | 15 / 18 | -0.8 | No |
| 545392 | 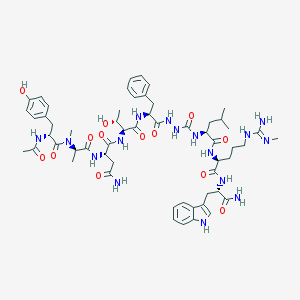 CHEMBL3977235 CHEMBL3977235 | C57H80N16O13 | 1197.37 | 14 / 16 | -0.4 | No |
| 545395 | 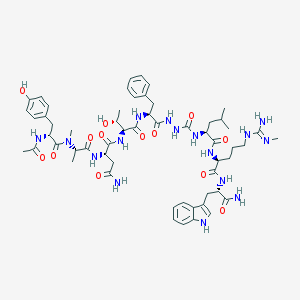 CHEMBL3947975 CHEMBL3947975 | C57H80N16O13 | 1197.37 | 14 / 16 | -0.4 | No |
| 455687 | 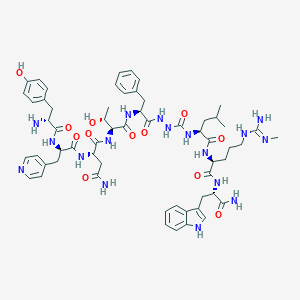 CHEMBL3314208 CHEMBL3314208 | C59H79N17O12 | 1218.39 | 15 / 17 | -0.3 | No |
| 545804 | 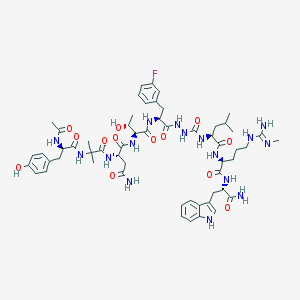 CHEMBL3968824 CHEMBL3968824 | C57H79FN16O13 | 1215.36 | 15 / 17 | -0.3 | No |
| 368145 | 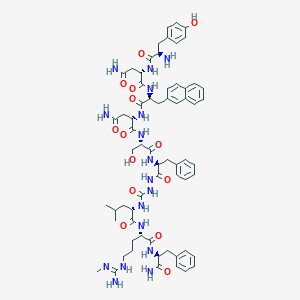 CHEMBL2151653 CHEMBL2151653 | C65H85N17O14 | 1328.5 | 16 / 18 | -0.3 | No |
| 377629 | 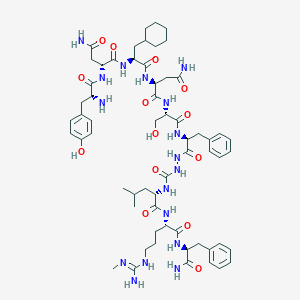 CHEMBL3087929 CHEMBL3087929 | C61H89N17O14 | 1284.49 | 16 / 18 | -0.4 | No |
| 377631 | 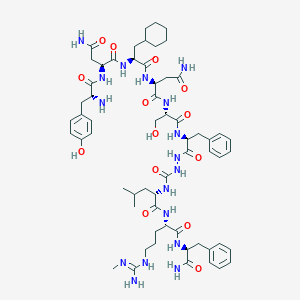 CHEMBL2151650 CHEMBL2151650 | C61H89N17O14 | 1284.49 | 16 / 18 | -0.4 | No |
| 457343 | 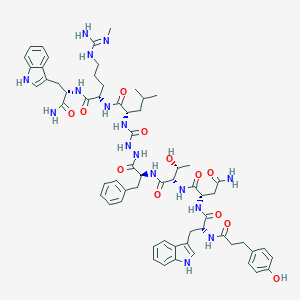 CHEMBL3314223 CHEMBL3314223 | C62H80N16O12 | 1241.42 | 13 / 17 | 1.7 | No |
| 393559 | 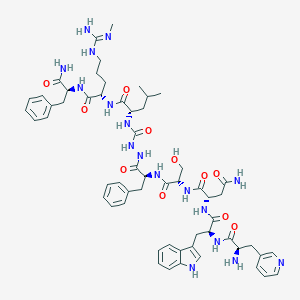 CHEMBL3086286 CHEMBL3086286 | C58H77N17O11 | 1188.36 | 14 / 16 | -0.4 | No |
| 400268 | 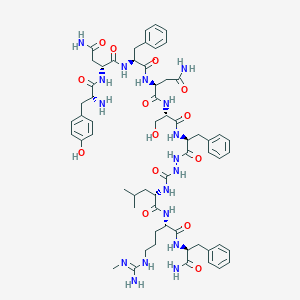 CHEMBL3087928 CHEMBL3087928 | C61H83N17O14 | 1278.44 | 16 / 18 | -1.6 | No |
| 457839 | 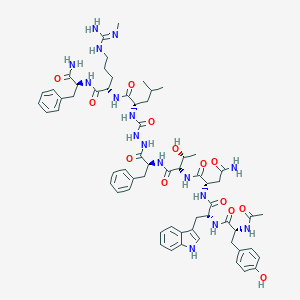 CHEMBL3314229 CHEMBL3314229 | C62H82N16O13 | 1259.44 | 14 / 17 | 1.0 | No |
| 547066 | 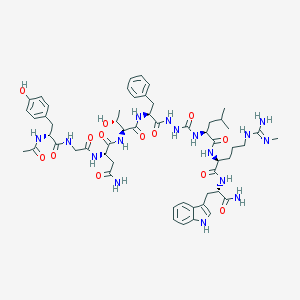 CHEMBL3896533 CHEMBL3896533 | C55H76N16O13 | 1169.31 | 14 / 17 | -1.0 | No |
| 547112 | 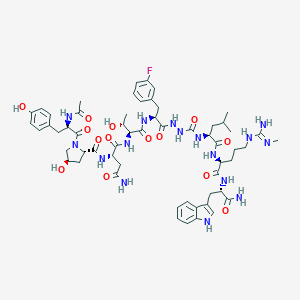 CHEMBL3967782 CHEMBL3967782 | C58H79FN16O14 | 1243.37 | 16 / 17 | -1.1 | No |
| 418463 | 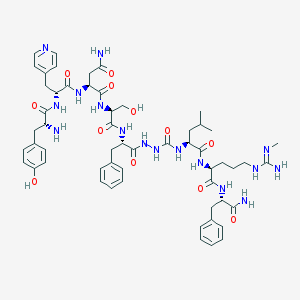 CHEMBL3086284 CHEMBL3086284 | C56H76N16O12 | 1165.32 | 15 / 16 | -0.9 | No |
| 423400 | 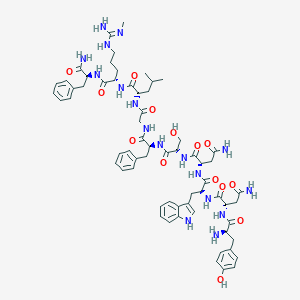 CHEMBL2151642 CHEMBL2151642 | C64H85N17O14 | 1316.49 | 16 / 18 | -1.4 | No |
| 458540 | 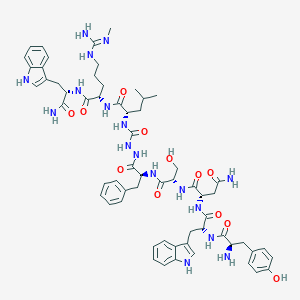 CHEMBL3314215 CHEMBL3314215 | C61H79N17O12 | 1242.41 | 14 / 18 | 0.5 | No |
| 570527 | 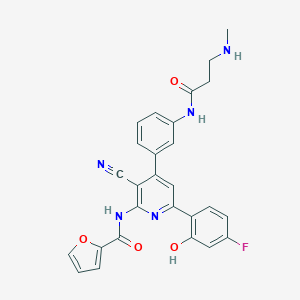 CHEMBL1169901 CHEMBL1169901 | C27H22FN5O4 | 499.502 | 8 / 4 | 4.0 | Yes |
| 427583 | 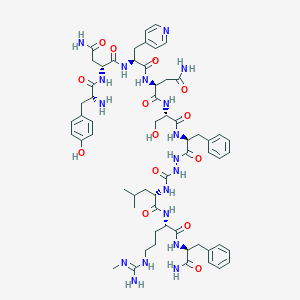 CHEMBL3087931 CHEMBL3087931 | C60H82N18O14 | 1279.43 | 17 / 18 | -2.6 | No |
| 427584 | 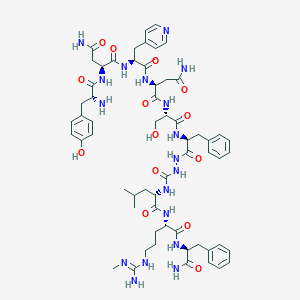 CHEMBL2151652 CHEMBL2151652 | C60H82N18O14 | 1279.43 | 17 / 18 | -2.6 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218