

You can:
| Name | KiSS-1 receptor |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | KISS1R |
| Synonym | Metastin receptor Kisspeptins receptor kisspeptin receptor KiSS1-derived peptide receptor KiSS-1R [ Show all ] |
| Disease | Prostate cancer |
| Length | 398 |
| Amino acid sequence | MHTVATSGPNASWGAPANASGCPGCGANASDGPVPSPRAVDAWLVPLFFAALMLLGLVGNSLVIYVICRHKPMRTVTNFYIANLAATDVTFLLCCVPFTALLYPLPGWVLGDFMCKFVNYIQQVSVQATCATLTAMSVDRWYVTVFPLRALHRRTPRLALAVSLSIWVGSAAVSAPVLALHRLSPGPRAYCSEAFPSRALERAFALYNLLALYLLPLLATCACYAAMLRHLGRVAVRPAPADSALQGQVLAERAGAVRAKVSRLVAAVVLLFAACWGPIQLFLVLQALGPAGSWHPRSYAAYALKTWAHCMSYSNSALNPLLYAFLGSHFRQAFRRVCPCAPRRPRRPRRPGPSDPAAPHAELLRLGSHPAPARAQKPGSSGLAARGLCVLGEDNAPL |
| UniProt | Q969F8 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | Q969F8 |
| 3D structure model | This predicted structure model is from GPCR-EXP Q969F8. |
| BioLiP | N/A |
| Therapeutic Target Database | T39123 |
| ChEMBL | CHEMBL5413 |
| IUPHAR | 266 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 225 | 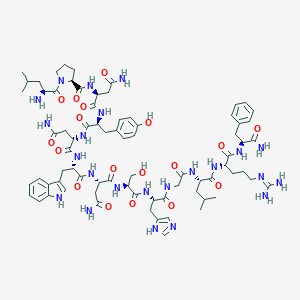 BDBM50203786 BDBM50203786 | C75H105N23O18 | 1616.81 | 21 / 22 | -4.4 | No |
| 513 | 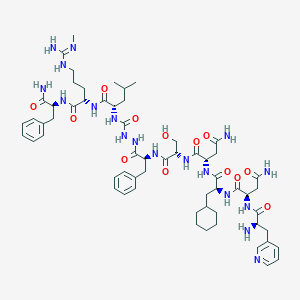 CHEMBL3085807 CHEMBL3085807 | C60H88N18O13 | 1269.48 | 16 / 17 | -1.1 | No |
| 441715 | 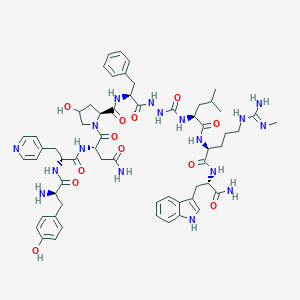 CHEMBL3314214 CHEMBL3314214 | C60H79N17O12 | 1230.4 | 15 / 16 | -0.3 | No |
| 2028 | 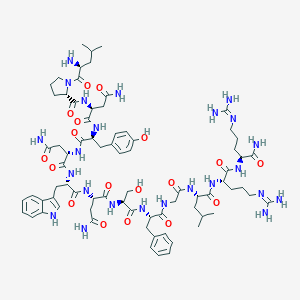 BDBM50203804 BDBM50203804 | C75H110N24O18 | 1635.85 | 21 / 23 | -5.7 | No |
| 2353 | 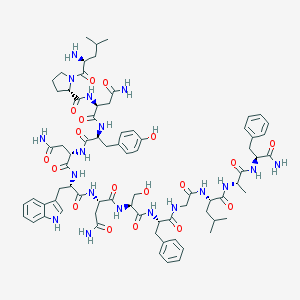 CHEMBL385105 CHEMBL385105 | C75H100N18O18 | 1541.73 | 19 / 19 | -1.7 | No |
| 5264 | 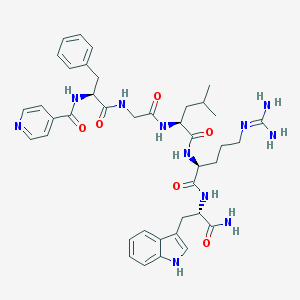 CHEMBL268659 CHEMBL268659 | C40H51N11O6 | 781.919 | 8 / 9 | 1.2 | No |
| 536077 | 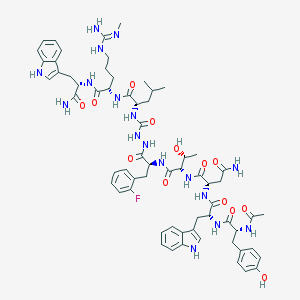 CHEMBL3928465 CHEMBL3928465 | C64H82FN17O13 | 1316.46 | 15 / 18 | 1.3 | No |
| 7154 | 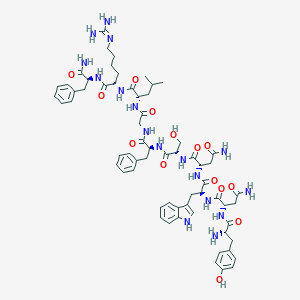 CHEMBL2152060 CHEMBL2152060 | C64H85N17O14 | 1316.49 | 16 / 18 | -1.5 | No |
| 9911 | 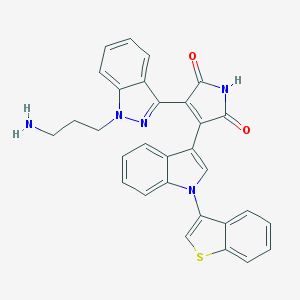 CHEMBL218690 CHEMBL218690 | C30H23N5O2S | 517.607 | 5 / 2 | 4.4 | No |
| 557623 | 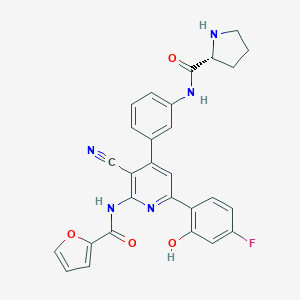 CHEMBL1171102 CHEMBL1171102 | C28H22FN5O4 | 511.513 | 8 / 4 | 4.0 | No |
| 548024 | 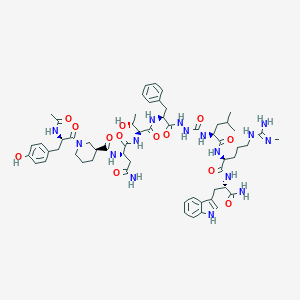 CHEMBL3982675 CHEMBL3982675 | C59H82N16O13 | 1223.4 | 14 / 16 | -0.3 | No |
| 14530 | 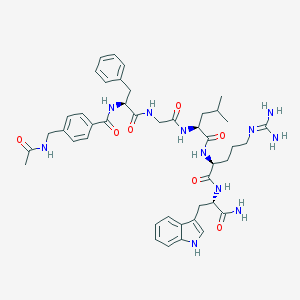 CHEMBL440431 CHEMBL440431 | C44H57N11O7 | 852.01 | 8 / 10 | 1.4 | No |
| 14919 | 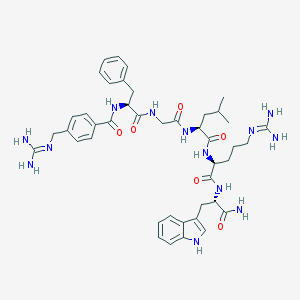 CHEMBL229621 CHEMBL229621 | C43H57N13O6 | 852.014 | 8 / 11 | 0.7 | No |
| 14929 | 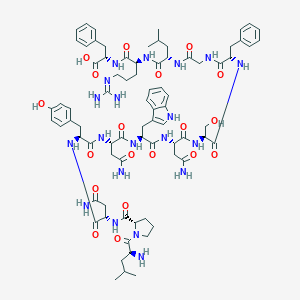 LPNYNWNSFGLRF LPNYNWNSFGLRF | C78H106N20O19 | 1627.83 | 21 / 21 | -4.5 | No |
| 557694 | 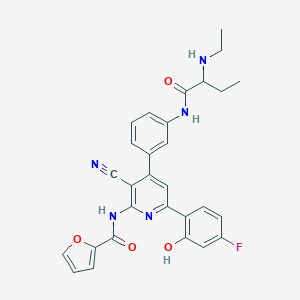 CHEMBL1171104 CHEMBL1171104 | C29H26FN5O4 | 527.556 | 8 / 4 | 4.8 | No |
| 15515 | 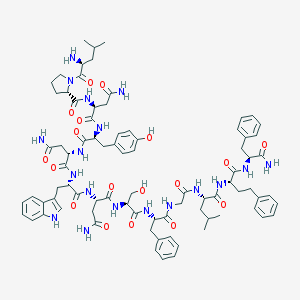 CHEMBL373854 CHEMBL373854 | C82H106N18O18 | 1631.86 | 19 / 19 | 0.3 | No |
| 442501 | 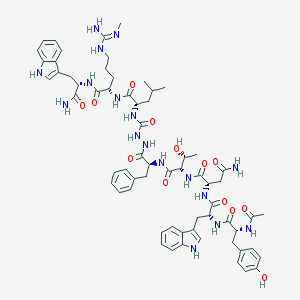 CHEMBL3314224 CHEMBL3314224 | C64H83N17O13 | 1298.47 | 14 / 18 | 1.2 | No |
| 548126 | 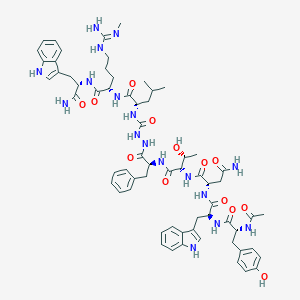 CHEMBL3985112 CHEMBL3985112 | C64H83N17O13 | 1298.47 | 14 / 18 | 1.2 | No |
| 21187 | 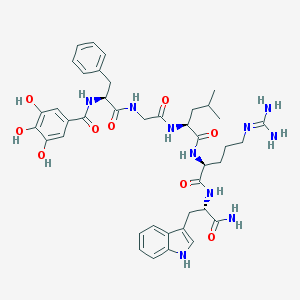 CHEMBL226537 CHEMBL226537 | C41H52N10O9 | 828.928 | 10 / 12 | 1.2 | No |
| 25492 | 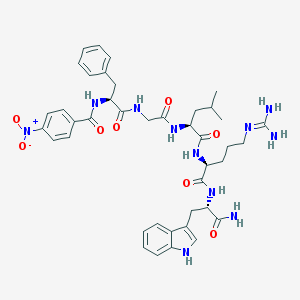 CHEMBL388146 CHEMBL388146 | C41H51N11O8 | 825.928 | 9 / 9 | 2.1 | No |
| 536713 | 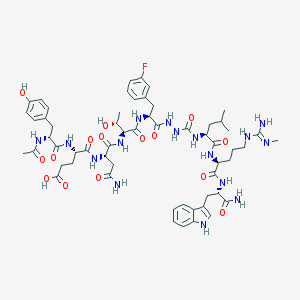 CHEMBL3977314 CHEMBL3977314 | C58H79FN16O15 | 1259.36 | 17 / 18 | -1.0 | No |
| 442825 | 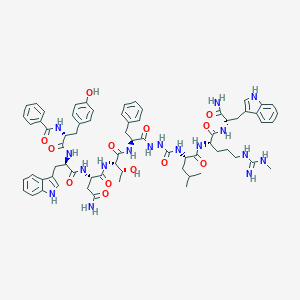 CHEMBL3314227 CHEMBL3314227 | C69H85N17O13 | 1360.55 | 14 / 19 | 3.1 | No |
| 558169 | 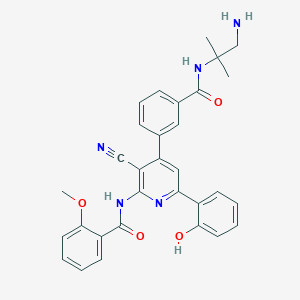 CHEMBL1163808 CHEMBL1163808 | C31H29N5O4 | 535.604 | 7 / 4 | 4.1 | No |
| 558279 | 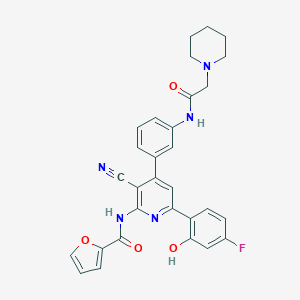 CHEMBL1171296 CHEMBL1171296 | C30H26FN5O4 | 539.567 | 8 / 3 | 4.8 | No |
| 442956 | 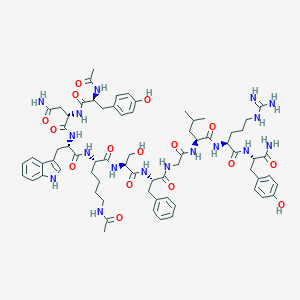 CHEMBL3422510 CHEMBL3422510 | C69H93N17O16 | 1416.61 | 17 / 20 | -0.1 | No |
| 34714 | 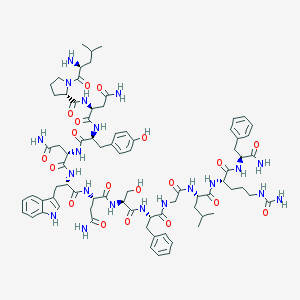 CHEMBL221764 CHEMBL221764 | C78H106N20O19 | 1627.83 | 20 / 21 | -2.6 | No |
| 443045 | 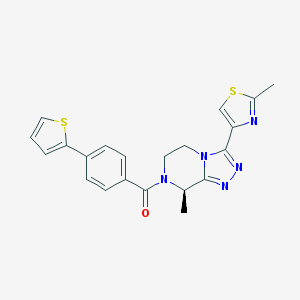 CHEMBL3422010 CHEMBL3422010 | C21H19N5OS2 | 421.537 | 6 / 0 | 3.0 | Yes |
| 558361 | 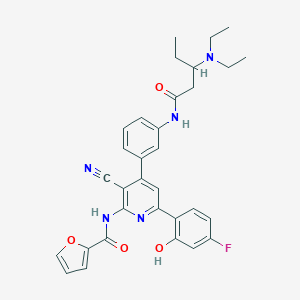 CHEMBL1171797 CHEMBL1171797 | C32H32FN5O4 | 569.637 | 8 / 3 | 5.6 | No |
| 35758 | 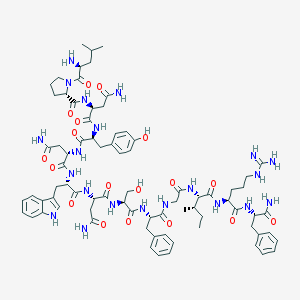 CID 44419669 CID 44419669 | C78H107N21O18 | 1626.84 | 20 / 22 | -2.2 | No |
| 36177 | 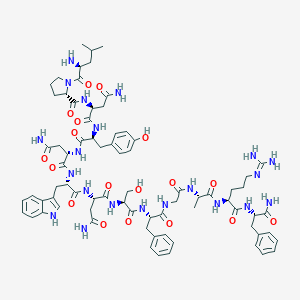 BDBM50203780 BDBM50203780 | C75H101N21O18 | 1584.76 | 20 / 21 | -4.2 | No |
| 36834 | 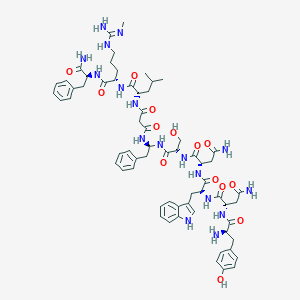 CHEMBL2151645 CHEMBL2151645 | C64H85N17O14 | 1316.49 | 16 / 18 | -1.2 | No |
| 443161 | 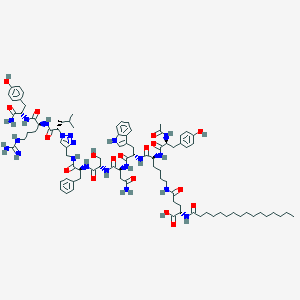 CHEMBL3422516 CHEMBL3422516 | C89H128N20O18 | 1766.12 | 21 / 21 | 6.5 | No |
| 536952 | 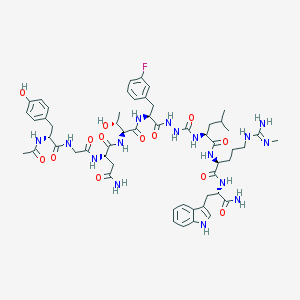 CHEMBL3939287 CHEMBL3939287 | C55H75FN16O13 | 1187.3 | 15 / 17 | -0.9 | No |
| 536984 | 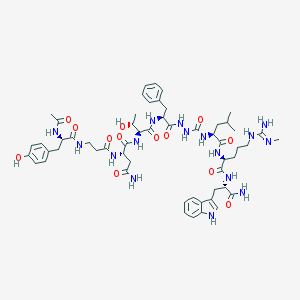 CHEMBL3985323 CHEMBL3985323 | C56H78N16O13 | 1183.34 | 14 / 17 | -1.1 | No |
| 443244 | 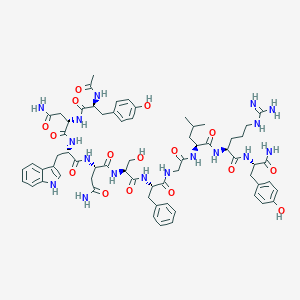 CHEMBL3422407 CHEMBL3422407 | C65H85N17O16 | 1360.5 | 17 / 20 | -1.9 | No |
| 40352 | 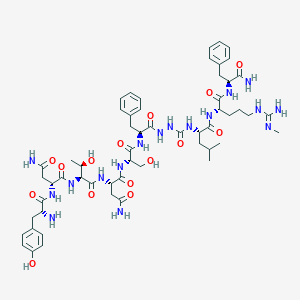 CHEMBL3087930 CHEMBL3087930 | C56H81N17O15 | 1232.37 | 17 / 19 | -3.8 | No |
| 40353 | 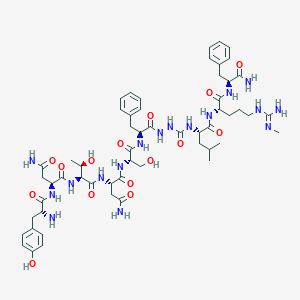 CHEMBL2151648 CHEMBL2151648 | C56H81N17O15 | 1232.37 | 17 / 19 | -3.8 | No |
| 558540 | 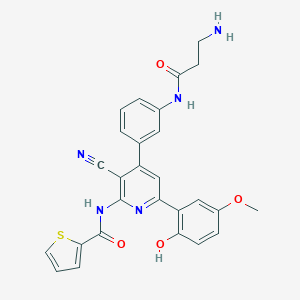 CHEMBL1165103 CHEMBL1165103 | C27H23N5O4S | 513.572 | 8 / 4 | 3.9 | No |
| 537091 | 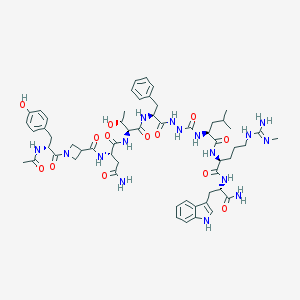 CHEMBL3931718 CHEMBL3931718 | C57H78N16O13 | 1195.35 | 14 / 16 | -1.1 | No |
| 43464 | 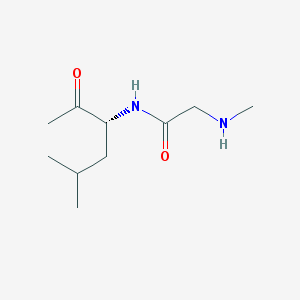 CHEMBL3645505 CHEMBL3645505 | C10H20N2O2 | 200.282 | 3 / 2 | 0.6 | Yes |
| 43465 | 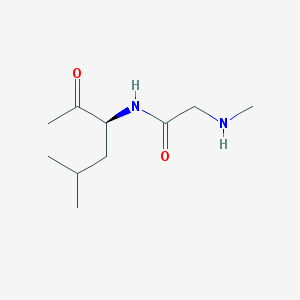 CHEMBL3645498 CHEMBL3645498 | C10H20N2O2 | 200.282 | 3 / 2 | 0.6 | Yes |
| 43520 | 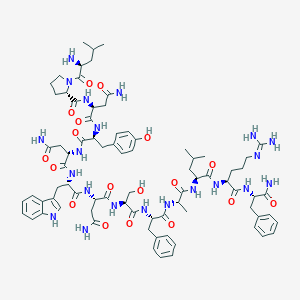 BDBM50203814 BDBM50203814 | C79H109N21O18 | 1640.87 | 20 / 21 | -2.1 | No |
| 443576 | 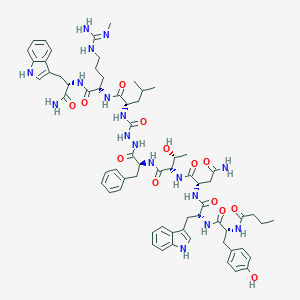 CHEMBL3314225 CHEMBL3314225 | C66H87N17O13 | 1326.53 | 14 / 18 | 2.0 | No |
| 49372 | 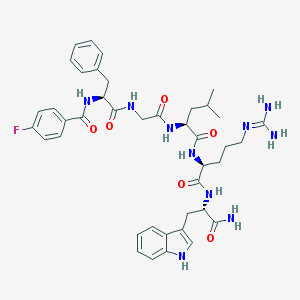 CHEMBL388586 CHEMBL388586 | C41H51FN10O6 | 798.921 | 8 / 9 | 2.4 | No |
| 49375 | 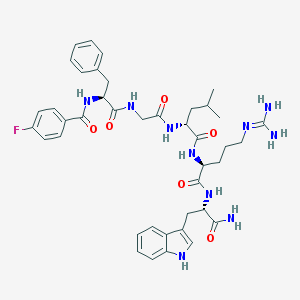 Peptide analogue, 26 Peptide analogue, 26 | C41H51FN10O6 | 798.921 | 8 / 9 | 2.4 | No |
| 56872 | 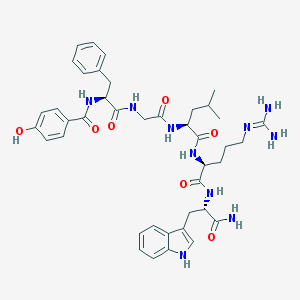 CHEMBL389708 CHEMBL389708 | C41H52N10O7 | 796.93 | 8 / 10 | 1.9 | No |
| 444023 | 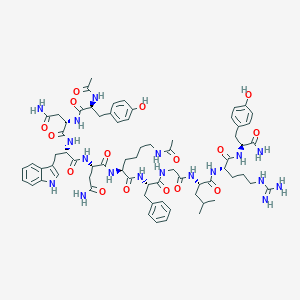 CHEMBL3422511 CHEMBL3422511 | C70H94N18O16 | 1443.63 | 17 / 20 | -0.8 | No |
| 444065 | 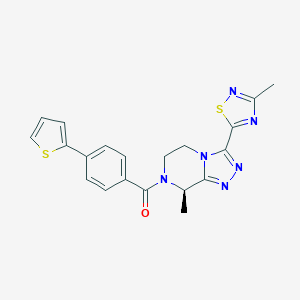 CHEMBL3422018 CHEMBL3422018 | C20H18N6OS2 | 422.525 | 7 / 0 | 2.8 | Yes |
| 62321 | 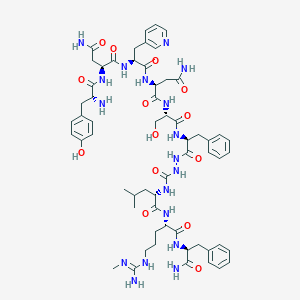 CHEMBL2151651 CHEMBL2151651 | C60H82N18O14 | 1279.43 | 17 / 18 | -2.6 | No |
| 444203 | 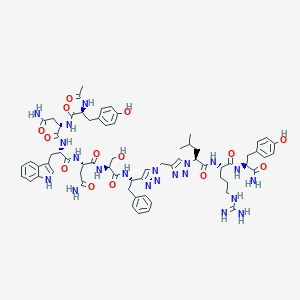 CHEMBL3422410 CHEMBL3422410 | C67H85N21O14 | 1408.55 | 19 / 18 | -1.5 | No |
| 559331 | 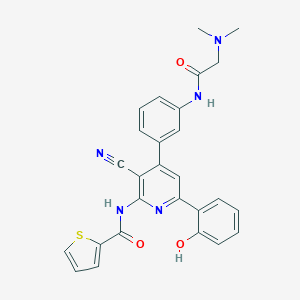 CHEMBL1169893 CHEMBL1169893 | C27H23N5O3S | 497.573 | 7 / 3 | 4.5 | Yes |
| 559441 | 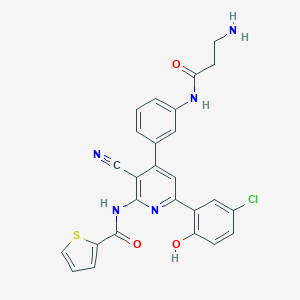 CHEMBL1164988 CHEMBL1164988 | C26H20ClN5O3S | 517.988 | 7 / 4 | 4.6 | No |
| 444446 | 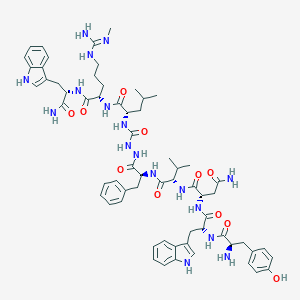 CHEMBL3314217 CHEMBL3314217 | C63H83N17O11 | 1254.46 | 13 / 17 | 2.5 | No |
| 71110 | 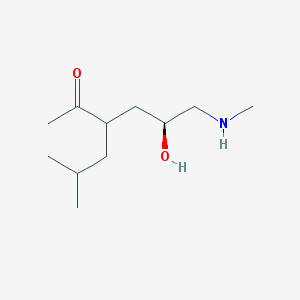 US8592379, 37 US8592379, 37 | C11H23NO2 | 201.31 | 3 / 2 | 0.9 | Yes |
| 71111 | 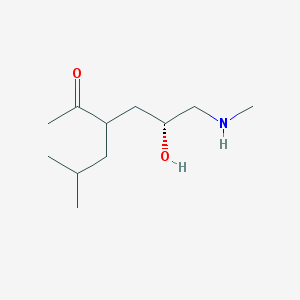 US8592379, 35 US8592379, 35 | C11H23NO2 | 201.31 | 3 / 2 | 0.9 | Yes |
| 71876 | 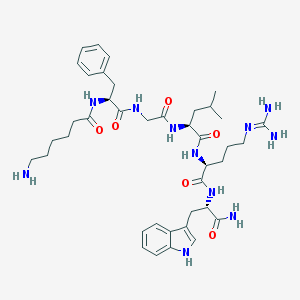 CHEMBL389262 CHEMBL389262 | C40H59N11O6 | 789.983 | 8 / 10 | 0.7 | No |
| 444601 | 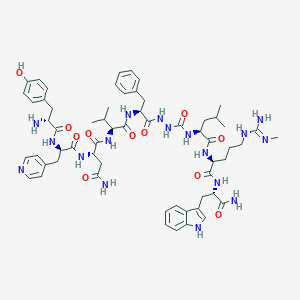 CHEMBL3314209 CHEMBL3314209 | C60H81N17O11 | 1216.42 | 14 / 16 | 1.3 | No |
| 78730 | 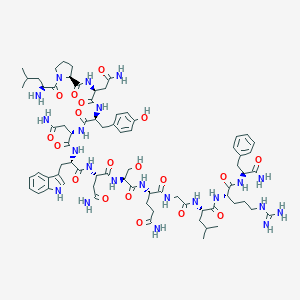 CID 44419652 CID 44419652 | C74H106N22O19 | 1607.8 | 21 / 23 | -5.3 | No |
| 444923 | 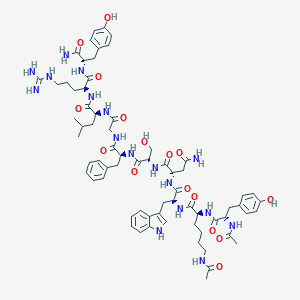 CHEMBL3422413 CHEMBL3422413 | C69H93N17O16 | 1416.61 | 17 / 20 | -0.1 | No |
| 81779 | 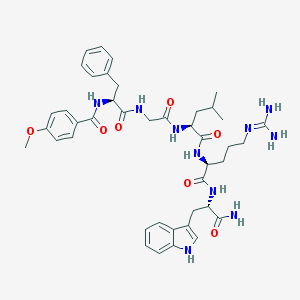 CHEMBL223390 CHEMBL223390 | C42H54N10O7 | 810.957 | 8 / 9 | 2.2 | No |
| 559877 | 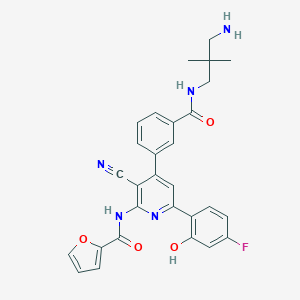 CHEMBL1165432 CHEMBL1165432 | C29H26FN5O4 | 527.556 | 8 / 4 | 4.2 | No |
| 559897 | 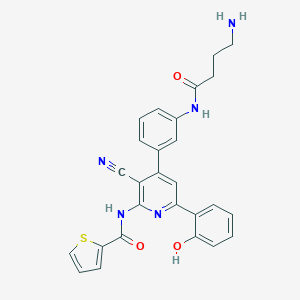 CHEMBL1164939 CHEMBL1164939 | C27H23N5O3S | 497.573 | 7 / 4 | 3.8 | Yes |
| 548939 | 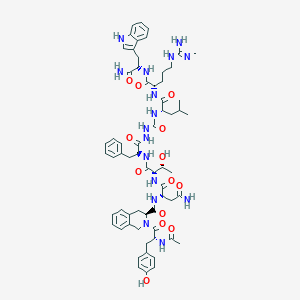 CHEMBL3964403 CHEMBL3964403 | C63H82N16O13 | 1271.45 | 14 / 16 | 0.8 | No |
| 560074 | 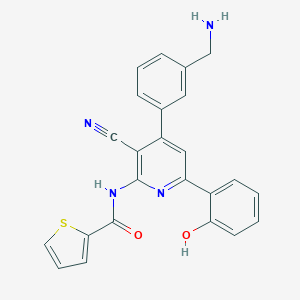 CHEMBL1164673 CHEMBL1164673 | C24H18N4O2S | 426.494 | 6 / 3 | 4.1 | Yes |
| 91857 | 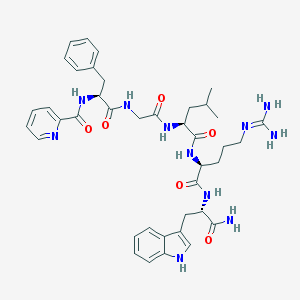 CHEMBL268579 CHEMBL268579 | C40H51N11O6 | 781.919 | 8 / 9 | 1.5 | No |
| 92004 | 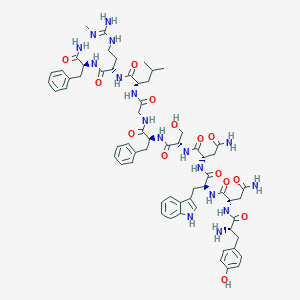 CHEMBL2152063 CHEMBL2152063 | C63H83N17O14 | 1302.46 | 16 / 18 | -1.8 | No |
| 92354 | 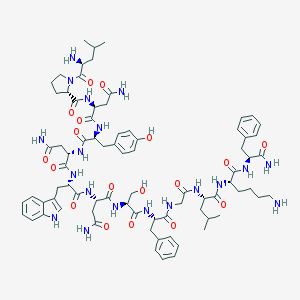 CHEMBL375432 CHEMBL375432 | C78H107N19O18 | 1598.83 | 20 / 20 | -1.9 | No |
| 548990 | 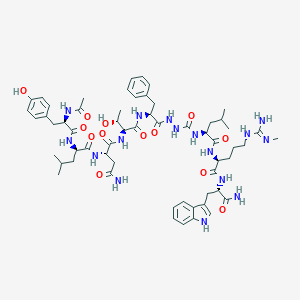 CHEMBL3976038 CHEMBL3976038 | C59H84N16O13 | 1225.42 | 14 / 17 | 0.8 | No |
| 92801 | 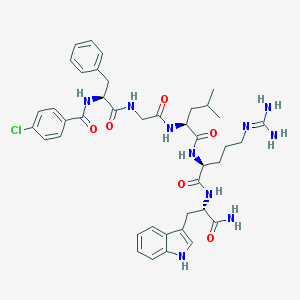 CHEMBL388585 CHEMBL388585 | C41H51ClN10O6 | 815.373 | 7 / 9 | 2.9 | No |
| 93156 | 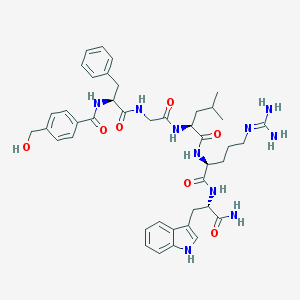 CHEMBL415004 CHEMBL415004 | C42H54N10O7 | 810.957 | 8 / 10 | 1.4 | No |
| 93161 | 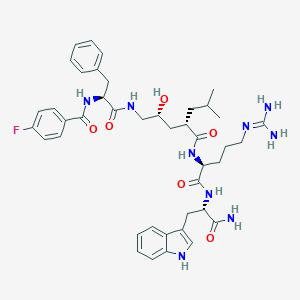 Peptide analogue, 24a Peptide analogue, 24a | C42H54FN9O6 | 799.949 | 8 / 9 | 3.0 | No |
| 93162 | 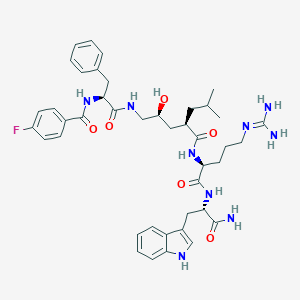 Peptide analogue, 25b Peptide analogue, 25b | C42H54FN9O6 | 799.949 | 8 / 9 | 3.0 | No |
| 93163 | 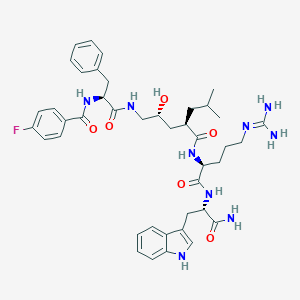 Peptide analogue, 24b Peptide analogue, 24b | C42H54FN9O6 | 799.949 | 8 / 9 | 3.0 | No |
| 93164 | 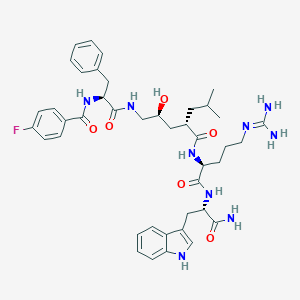 Peptide analogue, 25a Peptide analogue, 25a | C42H54FN9O6 | 799.949 | 8 / 9 | 3.0 | No |
| 445503 | 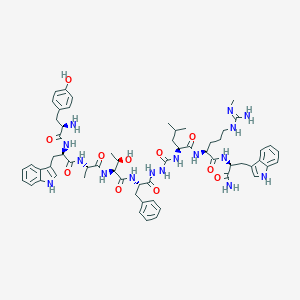 CHEMBL3314221 CHEMBL3314221 | C61H80N16O11 | 1213.41 | 13 / 17 | 2.4 | No |
| 445522 | 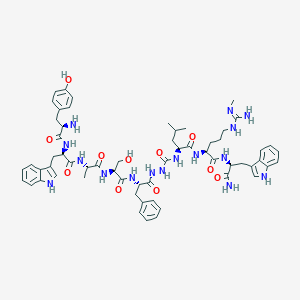 CHEMBL3314220 CHEMBL3314220 | C60H78N16O11 | 1199.39 | 13 / 17 | 2.0 | No |
| 560321 | 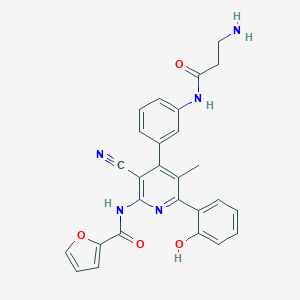 CHEMBL1163739 CHEMBL1163739 | C27H23N5O4 | 481.512 | 7 / 4 | 3.7 | Yes |
| 99859 | 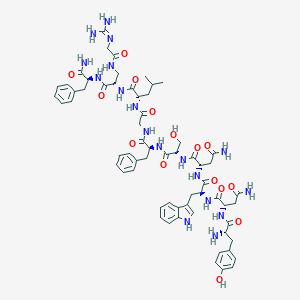 BDBM50392498 BDBM50392498 | C63H82N18O15 | 1331.46 | 17 / 19 | -3.1 | No |
| 517846 | 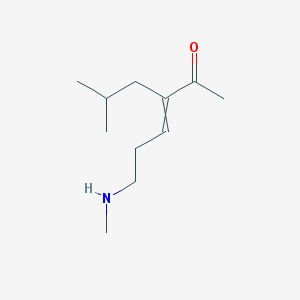 CHEMBL3926886 CHEMBL3926886 | C11H21NO | 183.295 | 2 / 1 | 1.8 | Yes |
| 445634 | 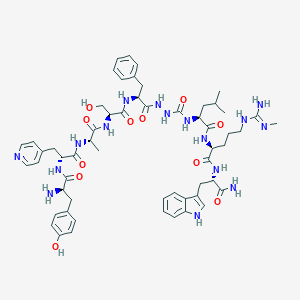 CHEMBL3314218 CHEMBL3314218 | C57H76N16O11 | 1161.34 | 14 / 16 | 0.8 | No |
| 100981 | 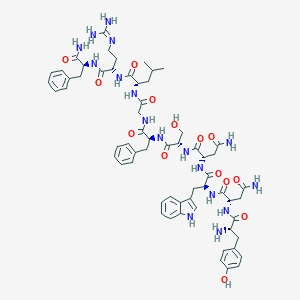 CHEMBL2152062 CHEMBL2152062 | C62H81N17O14 | 1288.44 | 16 / 18 | -2.4 | No |
| 445747 | 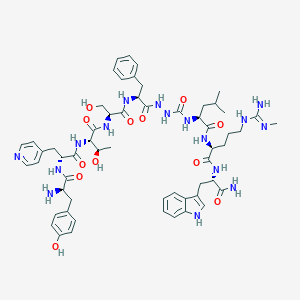 CHEMBL3314219 CHEMBL3314219 | C58H78N16O12 | 1191.36 | 15 / 17 | 0.2 | No |
| 560506 | 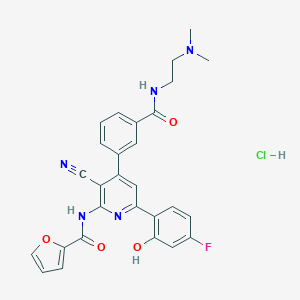 CHEMBL1171483 CHEMBL1171483 | C28H25ClFN5O4 | 549.987 | 8 / 4 | N/A | No |
| 103483 | 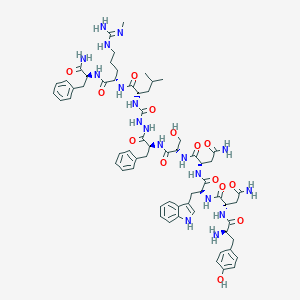 CHEMBL2151646 CHEMBL2151646 | C63H84N18O14 | 1317.48 | 16 / 19 | -1.4 | No |
| 103485 | 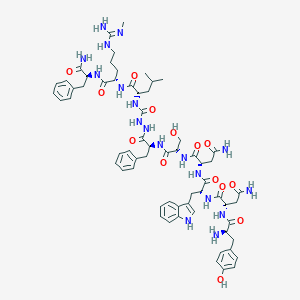 CHEMBL2151654 CHEMBL2151654 | C63H84N18O14 | 1317.48 | 16 / 19 | -1.4 | No |
| 103487 | 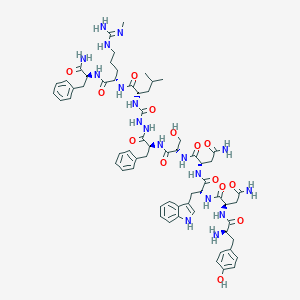 CHEMBL3085804 CHEMBL3085804 | C63H84N18O14 | 1317.48 | 16 / 19 | -1.4 | No |
| 103490 | 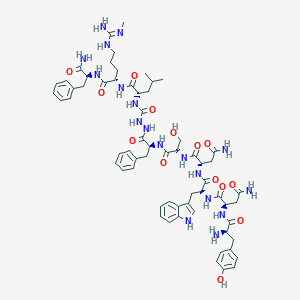 CHEMBL3087925 CHEMBL3087925 | C63H84N18O14 | 1317.48 | 16 / 19 | -1.4 | No |
| 103611 | 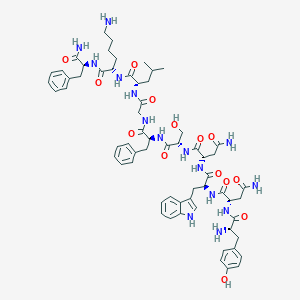 CHEMBL2152052 CHEMBL2152052 | C63H83N15O14 | 1274.45 | 16 / 17 | -1.1 | No |
| 107726 | 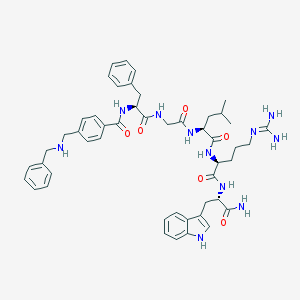 CHEMBL387648 CHEMBL387648 | C49H61N11O6 | 900.098 | 8 / 10 | 3.1 | No |
| 109663 | 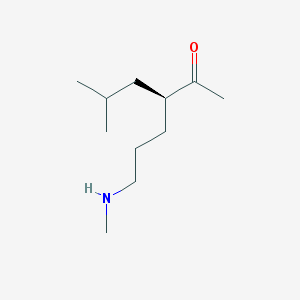 CHEMBL3645500 CHEMBL3645500 | C11H23NO | 185.311 | 2 / 1 | 1.9 | Yes |
| 560822 | 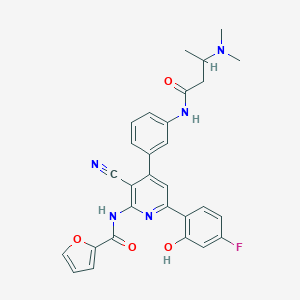 CHEMBL1170894 CHEMBL1170894 | C29H26FN5O4 | 527.556 | 8 / 3 | 4.3 | No |
| 116180 | 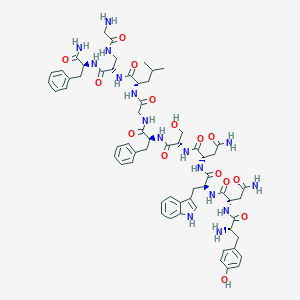 CHEMBL2152067 CHEMBL2152067 | C62H80N16O15 | 1289.42 | 17 / 18 | -2.6 | No |
| 560927 | 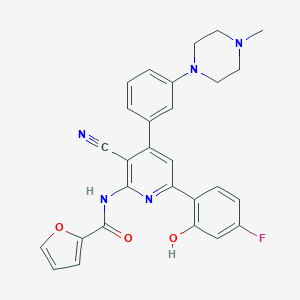 CHEMBL1172940 CHEMBL1172940 | C28H24FN5O3 | 497.53 | 8 / 2 | 4.7 | Yes |
| 122482 | 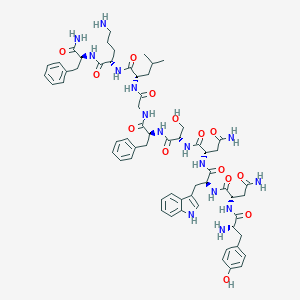 CHEMBL2152054 CHEMBL2152054 | C62H81N15O14 | 1260.42 | 16 / 17 | -1.4 | No |
| 122634 | 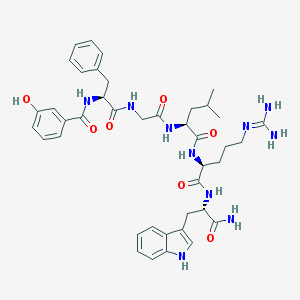 CHEMBL226566 CHEMBL226566 | C41H52N10O7 | 796.93 | 8 / 10 | 1.9 | No |
| 123519 | 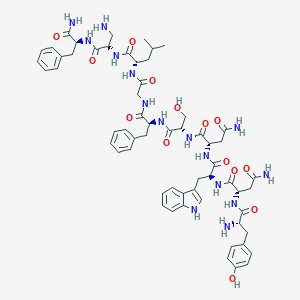 CHEMBL2152066 CHEMBL2152066 | C60H77N15O14 | 1232.37 | 16 / 17 | -2.1 | No |
| 446596 | 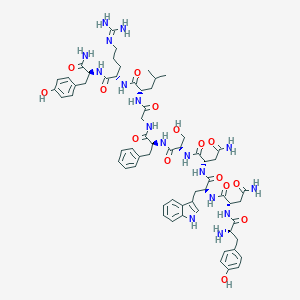 CHEMBL3422406 CHEMBL3422406 | C63H83N17O15 | 1318.46 | 17 / 19 | -2.4 | No |
| 561239 | 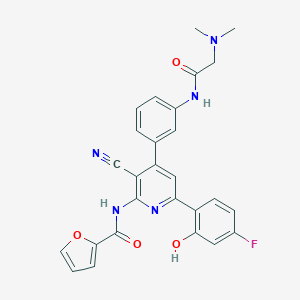 CHEMBL1171977 CHEMBL1171977 | C27H22FN5O4 | 499.502 | 8 / 3 | 4.0 | Yes |
| 561254 | 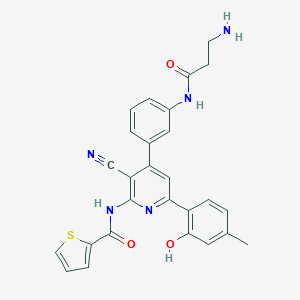 CHEMBL1163888 CHEMBL1163888 | C27H23N5O3S | 497.573 | 7 / 4 | 4.3 | Yes |
| 128470 | 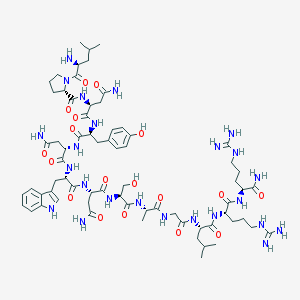 CID 44419665 CID 44419665 | C69H106N24O18 | 1559.76 | 21 / 25 | -6.5 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218