

You can:
| Name | Melanin-concentrating hormone receptor 2 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | MCHR2 |
| Synonym | G-protein coupled receptor 145 melanin-concentrating hormone receptor 2 MCHR-2 MCH2R MCH2 receptor [ Show all ] |
| Disease | N/A |
| Length | 340 |
| Amino acid sequence | MNPFHASCWNTSAELLNKSWNKEFAYQTASVVDTVILPSMIGIICSTGLVGNILIVFTIIRSRKKTVPDIYICNLAVADLVHIVGMPFLIHQWARGGEWVFGGPLCTIITSLDTCNQFACSAIMTVMSVDRYFALVQPFRLTRWRTRYKTIRINLGLWAASFILALPVWVYSKVIKFKDGVESCAFDLTSPDDVLWYTLYLTITTFFFPLPLILVCYILILCYTWEMYQQNKDARCCNPSVPKQRVMKLTKMVLVLVVVFILSAAPYHVIQLVNLQMEQPTLAFYVGYYLSICLSYASSSINPFLYILLSGNFQKRLPQIQRRATEKEINNMGNTLKSHF |
| UniProt | Q969V1 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | Q969V1 |
| 3D structure model | This predicted structure model is from GPCR-EXP Q969V1. |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL5038 |
| IUPHAR | 281 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 414 | 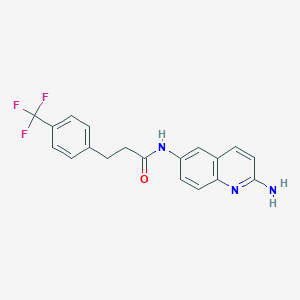 CHEMBL217751 CHEMBL217751 | C19H16F3N3O | 359.352 | 6 / 2 | 3.8 | Yes |
| 794 | 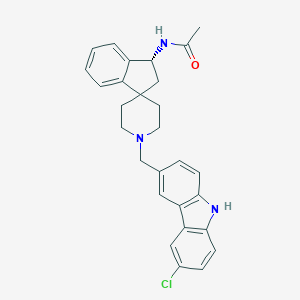 CHEMBL1934127 CHEMBL1934127 | C28H28ClN3O | 458.002 | 2 / 2 | 5.4 | No |
| 1571 | 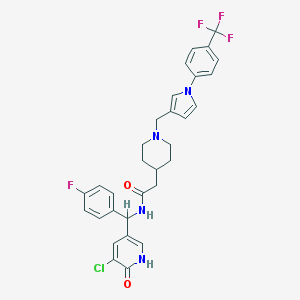 CHEMBL540930 CHEMBL540930 | C31H29ClF4N4O2 | 601.043 | 7 / 2 | 5.0 | No |
| 4931 | 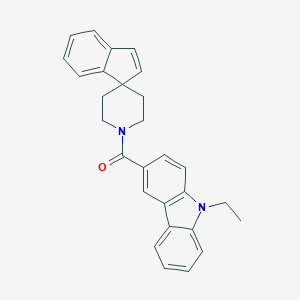 CHEMBL1934105 CHEMBL1934105 | C28H26N2O | 406.529 | 1 / 0 | 5.9 | No |
| 5010 | 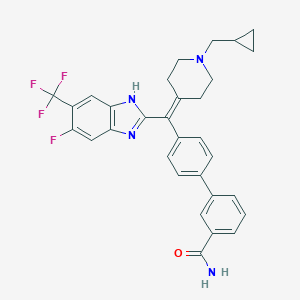 CHEMBL377270 CHEMBL377270 | C31H28F4N4O | 548.586 | 7 / 2 | 6.2 | No |
| 6231 | 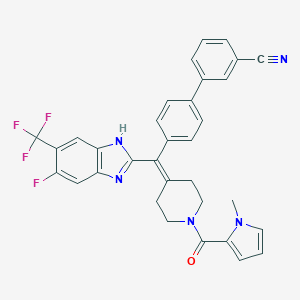 CHEMBL268938 CHEMBL268938 | C33H25F4N5O | 583.591 | 7 / 1 | 6.5 | No |
| 7515 | 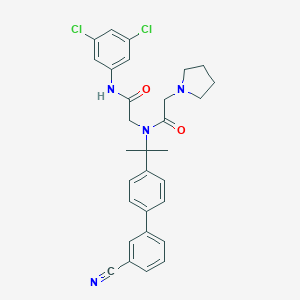 CHEMBL198724 CHEMBL198724 | C30H30Cl2N4O2 | 549.496 | 4 / 1 | 5.9 | No |
| 9149 | 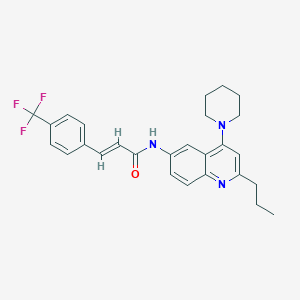 CHEMBL213873 CHEMBL213873 | C27H28F3N3O | 467.536 | 6 / 1 | 6.5 | No |
| 9519 | 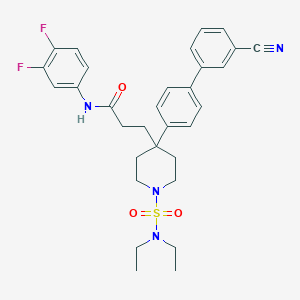 CHEMBL212671 CHEMBL212671 | C31H34F2N4O3S | 580.695 | 8 / 1 | 5.1 | No |
| 12310 | 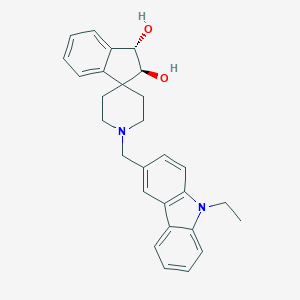 CHEMBL1934118 CHEMBL1934118 | C28H30N2O2 | 426.56 | 3 / 2 | 4.0 | Yes |
| 12311 | 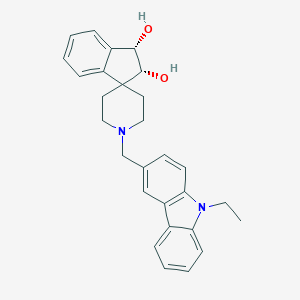 CHEMBL1934124 CHEMBL1934124 | C28H30N2O2 | 426.56 | 3 / 2 | 4.0 | Yes |
| 21552 | 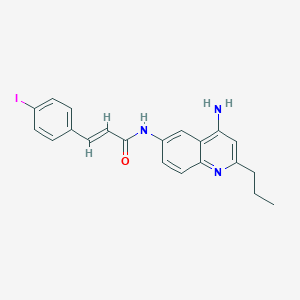 CHEMBL216510 CHEMBL216510 | C21H20IN3O | 457.315 | 3 / 2 | 4.6 | Yes |
| 21951 | 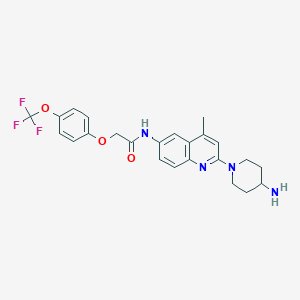 CHEMBL197026 CHEMBL197026 | C24H25F3N4O3 | 474.484 | 9 / 2 | 4.7 | Yes |
| 22429 | 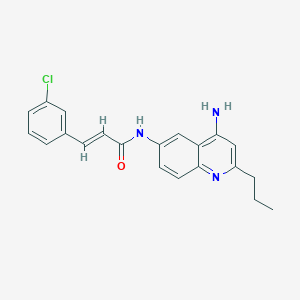 CHEMBL214843 CHEMBL214843 | C21H20ClN3O | 365.861 | 3 / 2 | 4.6 | Yes |
| 22663 | 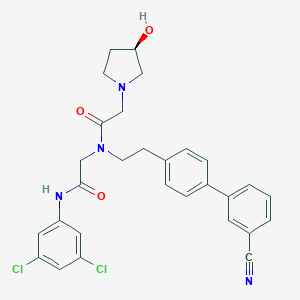 CHEMBL198945 CHEMBL198945 | C29H28Cl2N4O3 | 551.468 | 5 / 2 | 4.8 | No |
| 22665 | 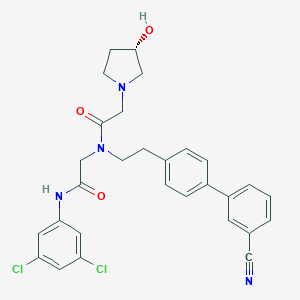 CHEMBL372551 CHEMBL372551 | C29H28Cl2N4O3 | 551.468 | 5 / 2 | 4.8 | No |
| 24358 | 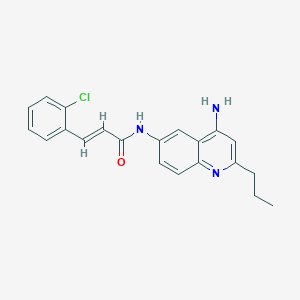 CHEMBL216342 CHEMBL216342 | C21H20ClN3O | 365.861 | 3 / 2 | 4.6 | Yes |
| 24508 | 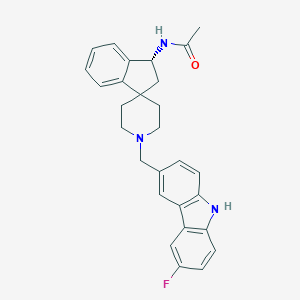 CHEMBL1934126 CHEMBL1934126 | C28H28FN3O | 441.55 | 3 / 2 | 4.8 | Yes |
| 25299 | 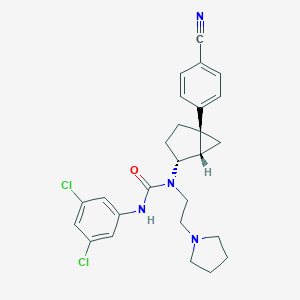 CHEMBL372721 CHEMBL372721 | C26H28Cl2N4O | 483.437 | 3 / 1 | 5.3 | No |
| 522348 | 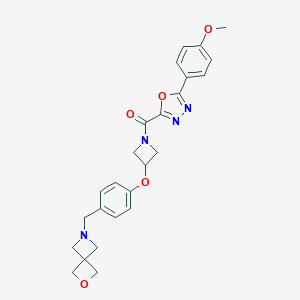 AZD-1979 AZD-1979 | C25H26N4O5 | 462.506 | 8 / 0 | 2.0 | Yes |
| 26899 | 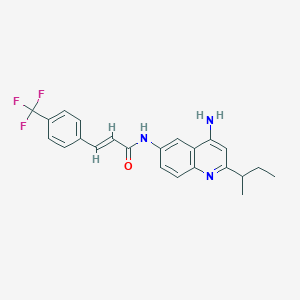 CHEMBL412831 CHEMBL412831 | C23H22F3N3O | 413.444 | 6 / 2 | 5.2 | No |
| 27816 | 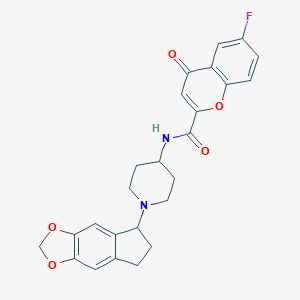 CHEMBL214585 CHEMBL214585 | C25H23FN2O5 | 450.466 | 7 / 1 | 3.5 | Yes |
| 29500 | 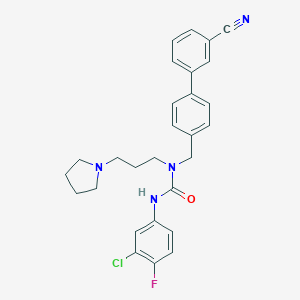 CHEMBL197429 CHEMBL197429 | C28H28ClFN4O | 491.007 | 4 / 1 | 5.6 | No |
| 31741 | 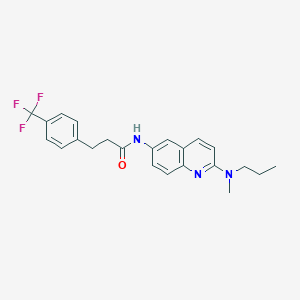 CHEMBL384848 CHEMBL384848 | C23H24F3N3O | 415.46 | 6 / 1 | 5.5 | No |
| 33013 | 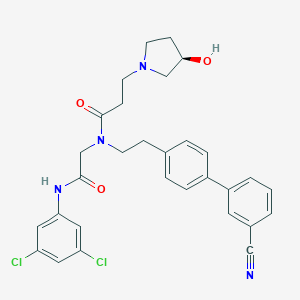 CHEMBL371099 CHEMBL371099 | C30H30Cl2N4O3 | 565.495 | 5 / 2 | 4.7 | No |
| 33014 | 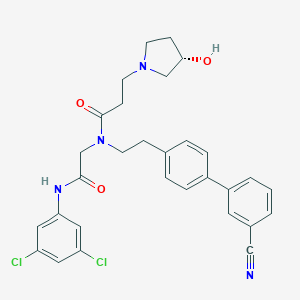 CHEMBL198562 CHEMBL198562 | C30H30Cl2N4O3 | 565.495 | 5 / 2 | 4.7 | No |
| 36555 | 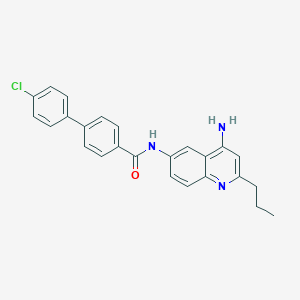 CHEMBL385957 CHEMBL385957 | C25H22ClN3O | 415.921 | 3 / 2 | 5.8 | No |
| 36675 | 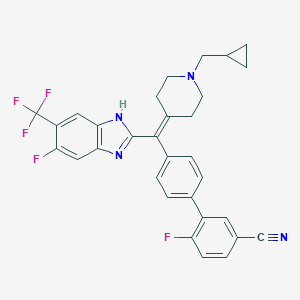 CHEMBL211672 CHEMBL211672 | C31H25F5N4 | 548.561 | 8 / 1 | 7.2 | No |
| 37338 | 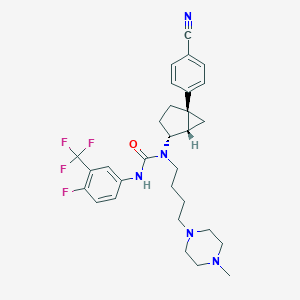 CHEMBL190842 CHEMBL190842 | C30H35F4N5O | 557.638 | 8 / 1 | 5.1 | No |
| 45395 | 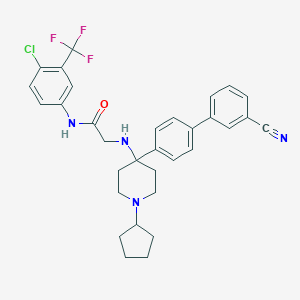 CHEMBL378180 CHEMBL378180 | C32H32ClF3N4O | 581.08 | 7 / 2 | 6.7 | No |
| 46351 | 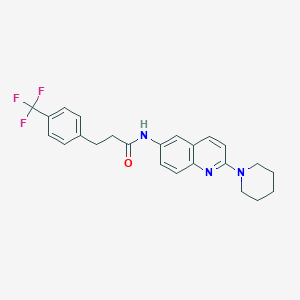 CHEMBL215662 CHEMBL215662 | C24H24F3N3O | 427.471 | 6 / 1 | 5.4 | No |
| 48335 | 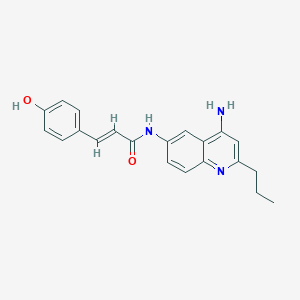 CHEMBL215586 CHEMBL215586 | C21H21N3O2 | 347.418 | 4 / 3 | 3.6 | Yes |
| 48518 | 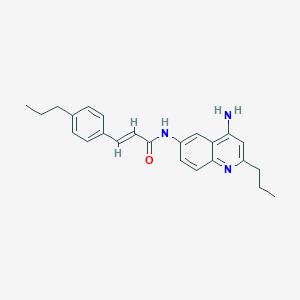 CHEMBL217462 CHEMBL217462 | C24H27N3O | 373.5 | 3 / 2 | 5.3 | No |
| 52962 | 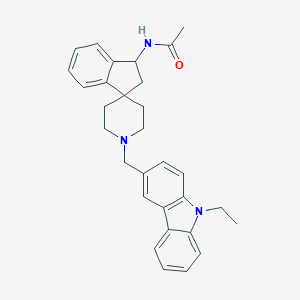 CHEMBL1934121 CHEMBL1934121 | C30H33N3O | 451.614 | 2 / 1 | 5.0 | Yes |
| 53129 | 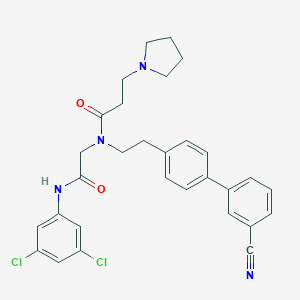 CHEMBL199274 CHEMBL199274 | C30H30Cl2N4O2 | 549.496 | 4 / 1 | 5.6 | No |
| 53863 | 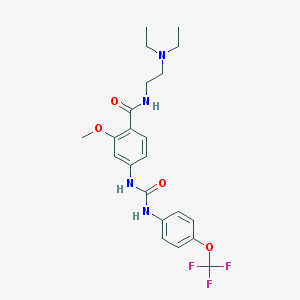 CHEMBL365055 CHEMBL365055 | C22H27F3N4O4 | 468.477 | 8 / 3 | 3.8 | Yes |
| 56506 | 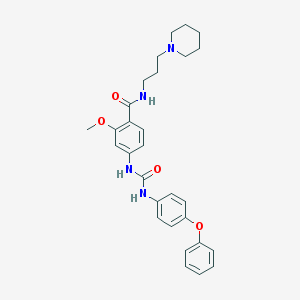 CHEMBL182554 CHEMBL182554 | C29H34N4O4 | 502.615 | 5 / 3 | 4.5 | No |
| 61008 | 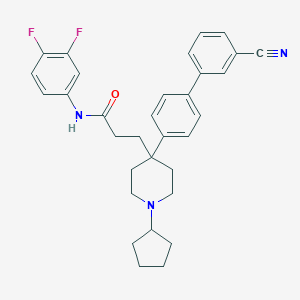 CHEMBL209089 CHEMBL209089 | C32H33F2N3O | 513.633 | 5 / 1 | 6.5 | No |
| 61510 | 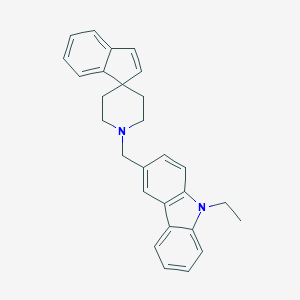 CHEMBL1934102 CHEMBL1934102 | C28H28N2 | 392.546 | 1 / 0 | 6.2 | No |
| 62673 | 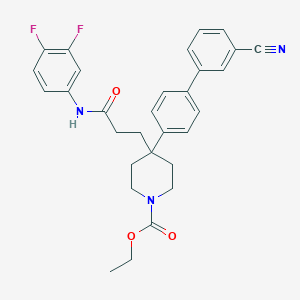 CHEMBL209059 CHEMBL209059 | C30H29F2N3O3 | 517.577 | 6 / 1 | 5.4 | No |
| 65472 | 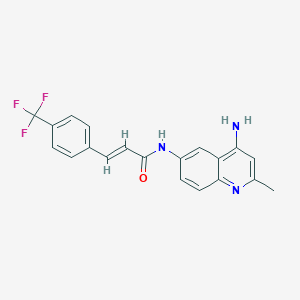 CHEMBL87370 CHEMBL87370 | C20H16F3N3O | 371.363 | 6 / 2 | 4.1 | Yes |
| 66560 | 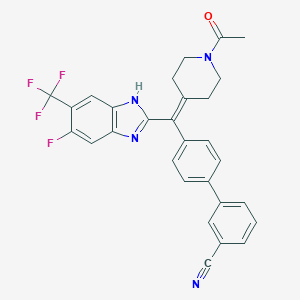 CHEMBL208573 CHEMBL208573 | C29H22F4N4O | 518.516 | 7 / 1 | 5.8 | No |
| 68717 | 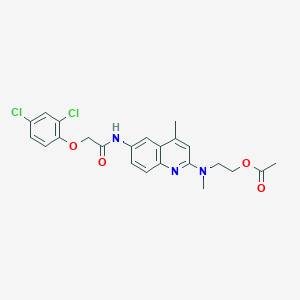 CHEMBL196567 CHEMBL196567 | C23H23Cl2N3O4 | 476.354 | 6 / 1 | 5.0 | Yes |
| 71792 | 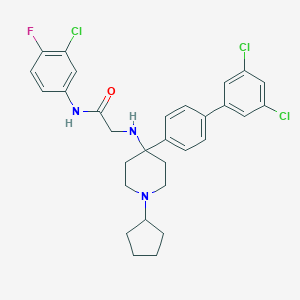 CHEMBL211504 CHEMBL211504 | C30H31Cl3FN3O | 574.946 | 4 / 2 | 7.5 | No |
| 72489 | 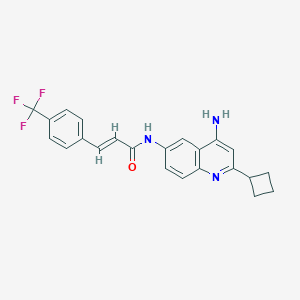 CHEMBL384547 CHEMBL384547 | C23H20F3N3O | 411.428 | 6 / 2 | 4.8 | Yes |
| 74795 | 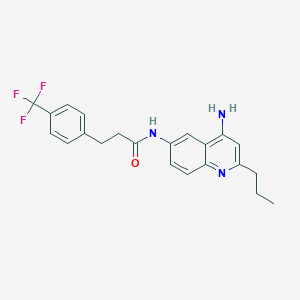 CHEMBL424721 CHEMBL424721 | C22H22F3N3O | 401.433 | 6 / 2 | 4.6 | Yes |
| 75681 | 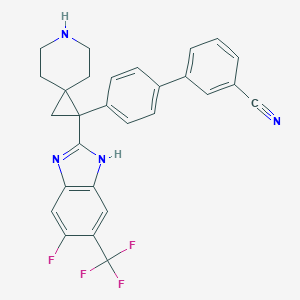 CHEMBL212256 CHEMBL212256 | C28H22F4N4 | 490.506 | 7 / 2 | 5.9 | No |
| 77154 | 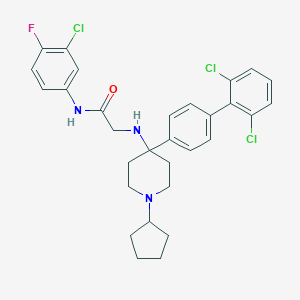 CHEMBL210255 CHEMBL210255 | C30H31Cl3FN3O | 574.946 | 4 / 2 | 7.5 | No |
| 79256 | 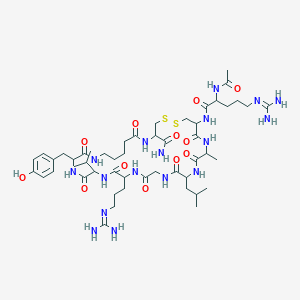 Ala8,Ava14,15 Ala8,Ava14,15 | C50H83N17O12S2 | 1178.44 | 16 / 16 | -2.7 | No |
| 79849 | 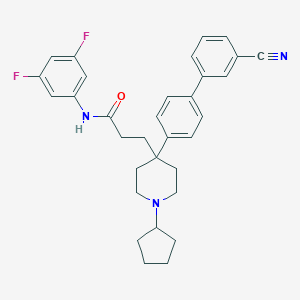 CHEMBL378972 CHEMBL378972 | C32H33F2N3O | 513.633 | 5 / 1 | 6.5 | No |
| 79921 | 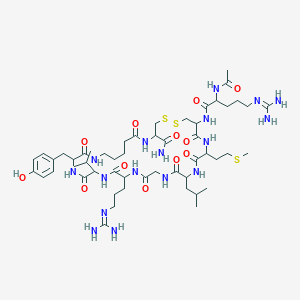 Ava14,15 Ava14,15 | C52H87N17O12S3 | 1238.56 | 17 / 16 | -2.1 | No |
| 79922 | 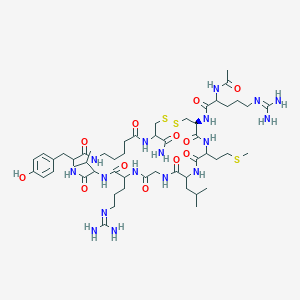 D-Arg6,Ava14,15 D-Arg6,Ava14,15 | C52H87N17O12S3 | 1238.56 | 17 / 16 | -2.1 | No |
| 80599 | 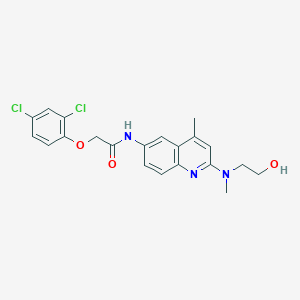 CHEMBL197245 CHEMBL197245 | C21H21Cl2N3O3 | 434.317 | 5 / 2 | 4.5 | Yes |
| 80835 | 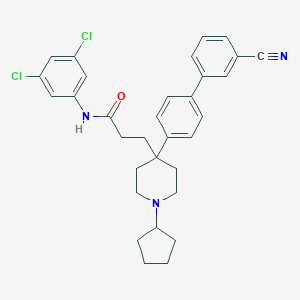 CHEMBL210343 CHEMBL210343 | C32H33Cl2N3O | 546.536 | 3 / 1 | 7.5 | No |
| 83774 | 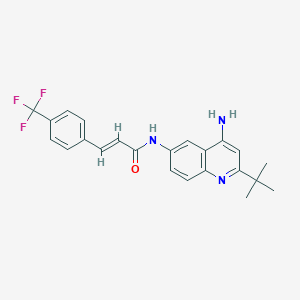 CHEMBL439358 CHEMBL439358 | C23H22F3N3O | 413.444 | 6 / 2 | 5.4 | No |
| 84217 | 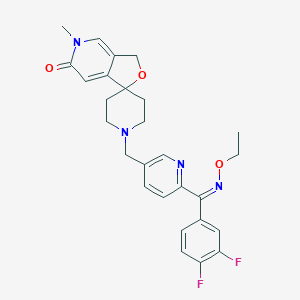 CHEMBL585664 CHEMBL585664 | C27H28F2N4O3 | 494.543 | 8 / 0 | 2.3 | Yes |
| 85154 | 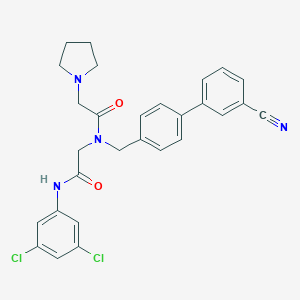 CHEMBL196819 CHEMBL196819 | C28H26Cl2N4O2 | 521.442 | 4 / 1 | 5.3 | No |
| 89179 | 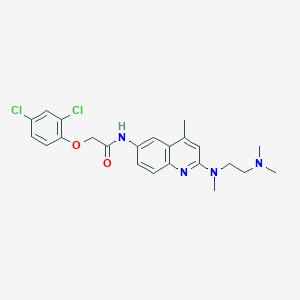 CHEMBL194408 CHEMBL194408 | C23H26Cl2N4O2 | 461.387 | 5 / 1 | 5.2 | No |
| 89490 | 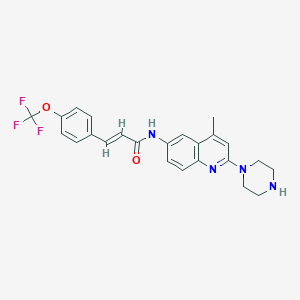 CHEMBL195351 CHEMBL195351 | C24H23F3N4O2 | 456.469 | 8 / 2 | 4.8 | Yes |
| 90090 | 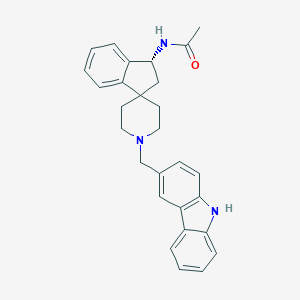 CHEMBL1934123 CHEMBL1934123 | C28H29N3O | 423.56 | 2 / 2 | 4.7 | Yes |
| 90091 | 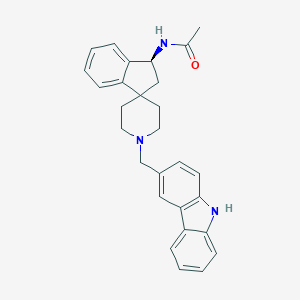 CHEMBL1934122 CHEMBL1934122 | C28H29N3O | 423.56 | 2 / 2 | 4.7 | Yes |
| 90133 | 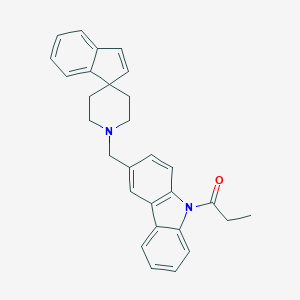 CHEMBL1934113 CHEMBL1934113 | C29H28N2O | 420.556 | 2 / 0 | 6.3 | No |
| 90593 | 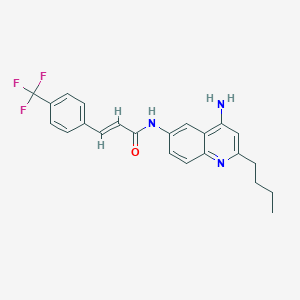 CHEMBL384765 CHEMBL384765 | C23H22F3N3O | 413.444 | 6 / 2 | 5.4 | No |
| 90635 | 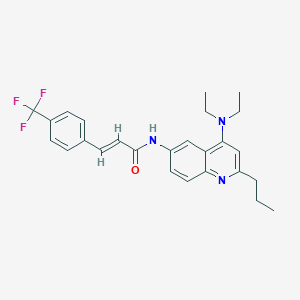 CHEMBL425979 CHEMBL425979 | C26H28F3N3O | 455.525 | 6 / 1 | 6.4 | No |
| 91711 | 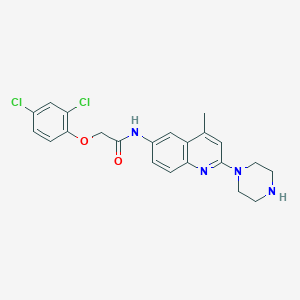 CHEMBL196667 CHEMBL196667 | C22H22Cl2N4O2 | 445.344 | 5 / 2 | 4.5 | Yes |
| 91974 | 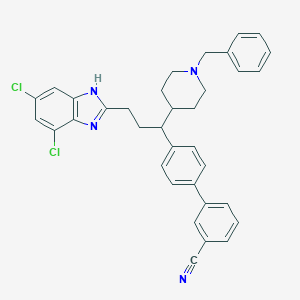 CHEMBL377874 CHEMBL377874 | C35H32Cl2N4 | 579.569 | 3 / 1 | 8.8 | No |
| 92949 | 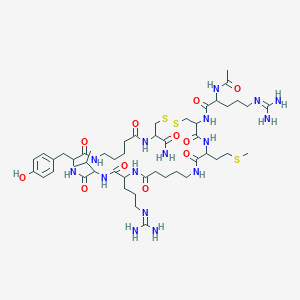 Ava9,10,Ava14,15 Ava9,10,Ava14,15 | C49H82N16O11S3 | 1167.48 | 16 / 15 | -2.5 | No |
| 94363 | 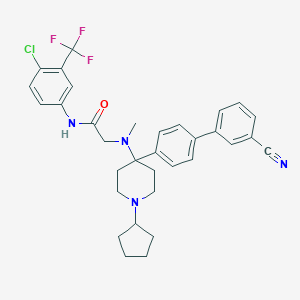 CHEMBL379489 CHEMBL379489 | C33H34ClF3N4O | 595.107 | 7 / 1 | 7.2 | No |
| 95063 | 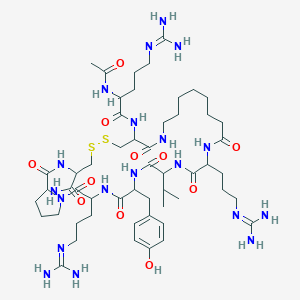 Aoct8,9,10 Aoct8,9,10 | C53H89N19O11S2 | 1232.54 | 16 / 16 | -2.9 | No |
| 96964 | 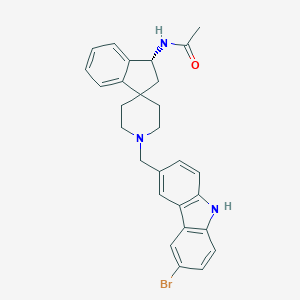 CHEMBL1934128 CHEMBL1934128 | C28H28BrN3O | 502.456 | 2 / 2 | 5.4 | No |
| 97395 | 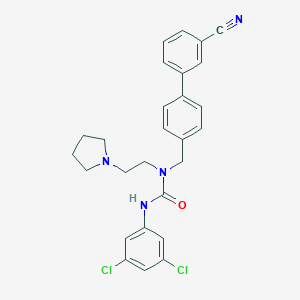 CHEMBL197280 CHEMBL197280 | C27H26Cl2N4O | 493.432 | 3 / 1 | 5.7 | No |
| 97562 | 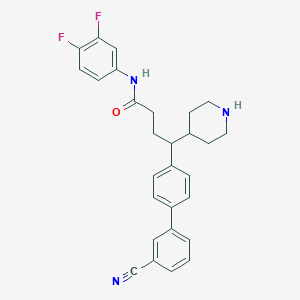 CHEMBL377321 CHEMBL377321 | C28H27F2N3O | 459.541 | 5 / 2 | 5.0 | Yes |
| 98507 | 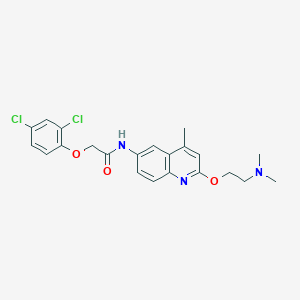 CHEMBL194837 CHEMBL194837 | C22H23Cl2N3O3 | 448.344 | 5 / 1 | 5.0 | Yes |
| 100470 | 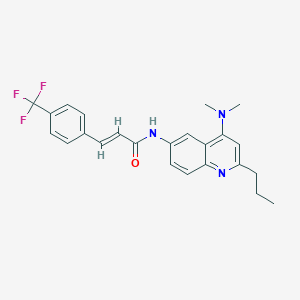 CHEMBL217575 CHEMBL217575 | C24H24F3N3O | 427.471 | 6 / 1 | 5.6 | No |
| 100766 | 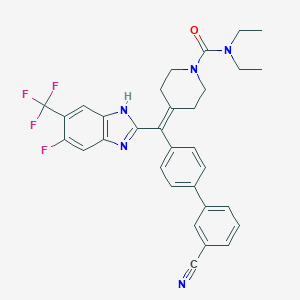 CHEMBL211615 CHEMBL211615 | C32H29F4N5O | 575.612 | 7 / 1 | 6.5 | No |
| 102294 | 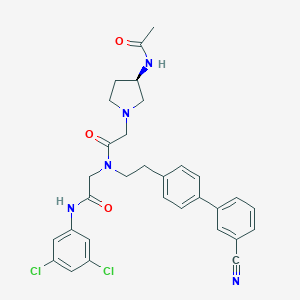 CHEMBL372803 CHEMBL372803 | C31H31Cl2N5O3 | 592.521 | 5 / 2 | 4.8 | No |
| 104778 | 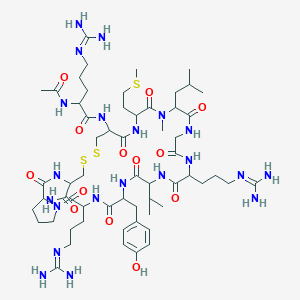 N-Me-Leu9 N-Me-Leu9 | C59H99N21O13S3 | 1406.76 | 19 / 17 | -3.3 | No |
| 105164 | 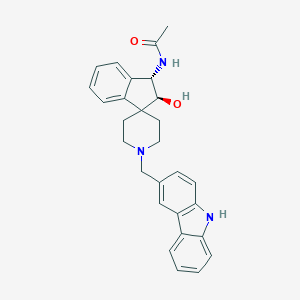 CHEMBL1934125 CHEMBL1934125 | C28H29N3O2 | 439.559 | 3 / 3 | 3.8 | Yes |
| 115381 | 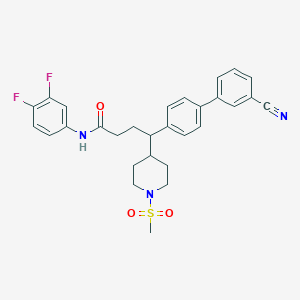 CHEMBL208081 CHEMBL208081 | C29H29F2N3O3S | 537.626 | 7 / 1 | 4.7 | No |
| 117239 | 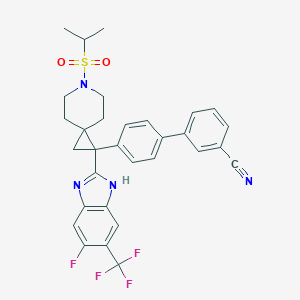 CHEMBL208688 CHEMBL208688 | C31H28F4N4O2S | 596.645 | 9 / 1 | 6.4 | No |
| 117535 | 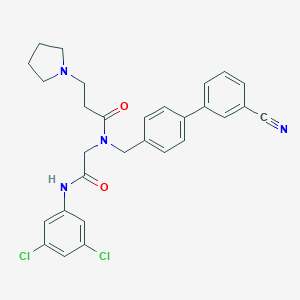 CHEMBL198689 CHEMBL198689 | C29H28Cl2N4O2 | 535.469 | 4 / 1 | 5.2 | No |
| 118711 | 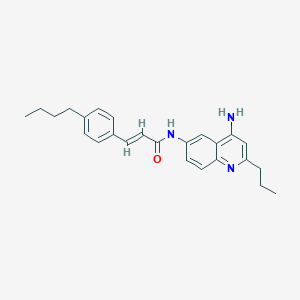 CHEMBL385221 CHEMBL385221 | C25H29N3O | 387.527 | 3 / 2 | 5.8 | No |
| 121387 | 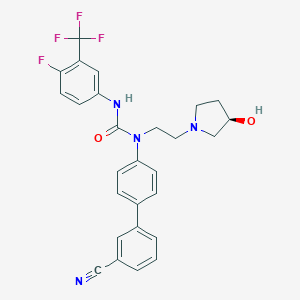 CHEMBL191393 CHEMBL191393 | C27H24F4N4O2 | 512.509 | 8 / 2 | 4.5 | No |
| 123560 | 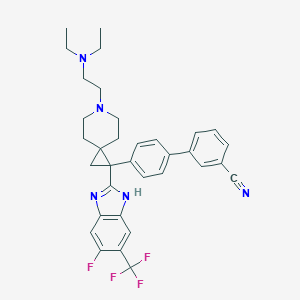 CHEMBL209754 CHEMBL209754 | C34H35F4N5 | 589.683 | 8 / 1 | 7.1 | No |
| 124132 | 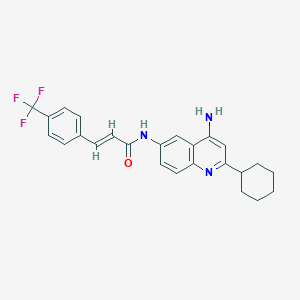 CHEMBL386320 CHEMBL386320 | C25H24F3N3O | 439.482 | 6 / 2 | 5.9 | No |
| 125641 | 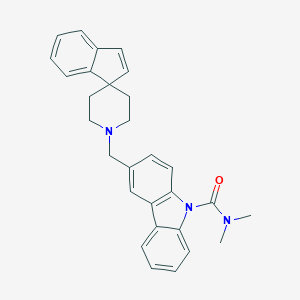 CHEMBL1934109 CHEMBL1934109 | C29H29N3O | 435.571 | 2 / 0 | 5.8 | No |
| 126741 | 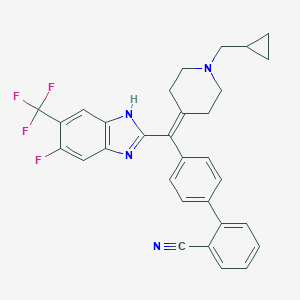 CHEMBL384449 CHEMBL384449 | C31H26F4N4 | 530.571 | 7 / 1 | 7.1 | No |
| 128970 | 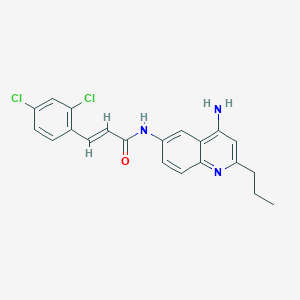 CHEMBL217614 CHEMBL217614 | C21H19Cl2N3O | 400.303 | 3 / 2 | 5.2 | No |
| 130394 | 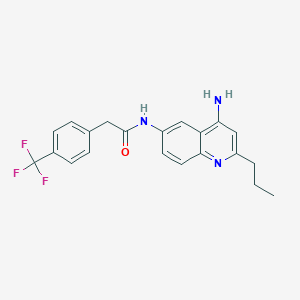 CHEMBL386509 CHEMBL386509 | C21H20F3N3O | 387.406 | 6 / 2 | 4.4 | Yes |
| 131897 | 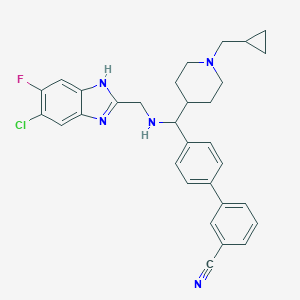 CHEMBL209510 CHEMBL209510 | C31H31ClFN5 | 528.072 | 5 / 2 | 6.0 | No |
| 132874 | 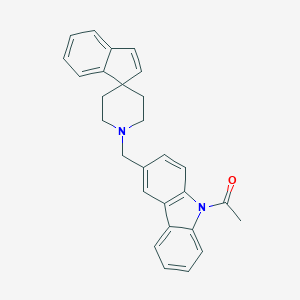 CHEMBL1934108 CHEMBL1934108 | C28H26N2O | 406.529 | 2 / 0 | 5.8 | No |
| 134632 | 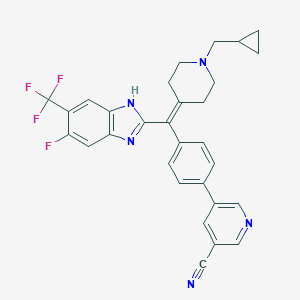 CHEMBL377372 CHEMBL377372 | C30H25F4N5 | 531.559 | 8 / 1 | 6.0 | No |
| 136757 | 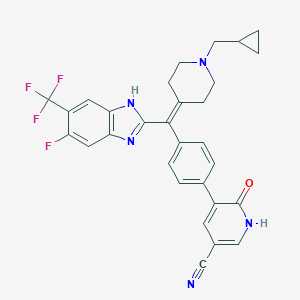 CHEMBL209590 CHEMBL209590 | C30H25F4N5O | 547.558 | 8 / 2 | 5.1 | No |
| 139685 | 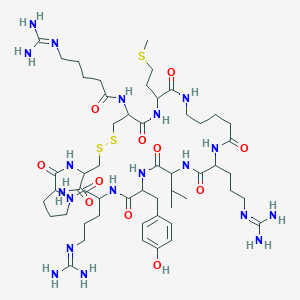 Gva6,Ava9,10 Gva6,Ava9,10 | C53H89N19O11S3 | 1264.6 | 17 / 16 | -3.4 | No |
| 139692 | 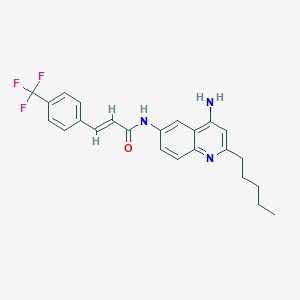 CHEMBL216428 CHEMBL216428 | C24H24F3N3O | 427.471 | 6 / 2 | 5.9 | No |
| 139753 | 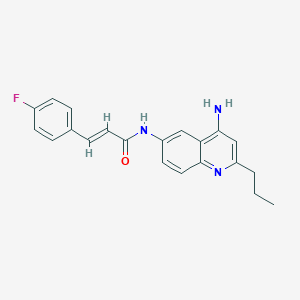 CHEMBL384550 CHEMBL384550 | C21H20FN3O | 349.409 | 4 / 2 | 4.1 | Yes |
| 140341 | 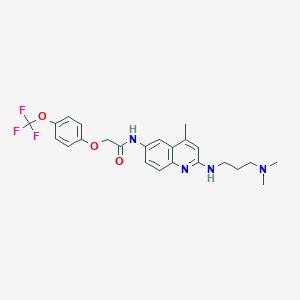 CHEMBL196581 CHEMBL196581 | C24H27F3N4O3 | 476.5 | 9 / 2 | 5.3 | No |
| 148047 | 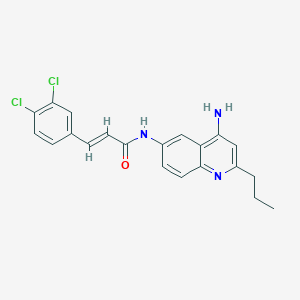 CHEMBL215241 CHEMBL215241 | C21H19Cl2N3O | 400.303 | 3 / 2 | 5.2 | No |
| 149741 | 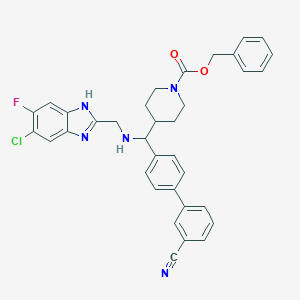 CHEMBL211311 CHEMBL211311 | C35H31ClFN5O2 | 608.114 | 6 / 2 | 6.6 | No |
| 149747 | 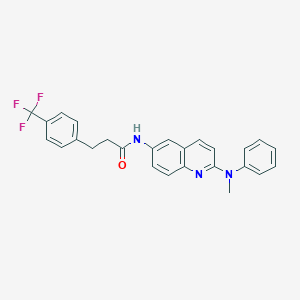 CHEMBL215989 CHEMBL215989 | C26H22F3N3O | 449.477 | 6 / 1 | 6.2 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218