

You can:
| Name | Mas-related G-protein coupled receptor member X2 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | MRGPRX2 |
| Synonym | Mrgprb10 MRGPRX2 MRGX2 |
| Disease | N/A |
| Length | 330 |
| Amino acid sequence | MDPTTPAWGTESTTVNGNDQALLLLCGKETLIPVFLILFIALVGLVGNGFVLWLLGFRMRRNAFSVYVLSLAGADFLFLCFQIINCLVYLSNFFCSISINFPSFFTTVMTCAYLAGLSMLSTVSTERCLSVLWPIWYRCRRPRHLSAVVCVLLWALSLLLSILEGKFCGFLFSDGDSGWCQTFDFITAAWLIFLFMVLCGSSLALLVRILCGSRGLPLTRLYLTILLTVLVFLLCGLPFGIQWFLILWIWKDSDVLFCHIHPVSVVLSSLNSSANPIIYFFVGSFRKQWRLQQPILKLALQRALQDIAEVDHSEGCFRQGTPEMSRSSLV |
| UniProt | Q96LB1 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | Q96LB1 |
| 3D structure model | This predicted structure model is from GPCR-EXP Q96LB1. |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL5849 |
| IUPHAR | 157 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 521522 | 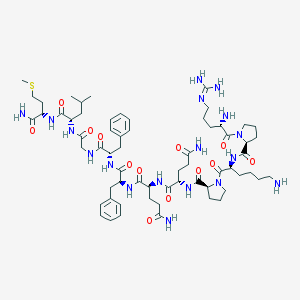 Substance P Substance P | C63H98N18O13S | 1347.65 | 17 / 15 | -2.3 | No |
| 521612 | 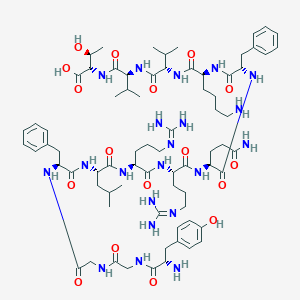 Dynorphin B Dynorphin B | C74H115N21O17 | 1570.86 | 21 / 22 | -3.6 | No |
| 522463 | 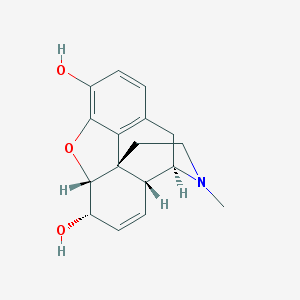 morphine morphine | C17H19NO3 | 285.343 | 4 / 2 | 0.8 | Yes |
| 522464 | 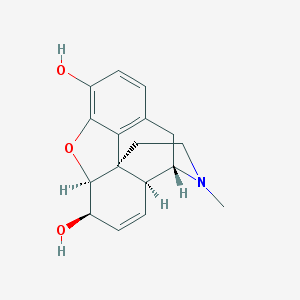 (+)-morphine (+)-morphine | C17H19NO3 | 285.343 | 4 / 2 | 0.8 | Yes |
| 522499 | 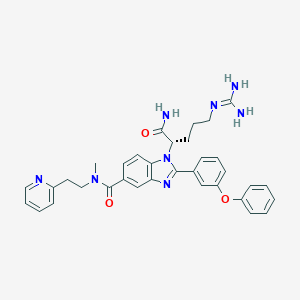 CHEMBL3719266 CHEMBL3719266 | C34H36N8O3 | 604.715 | 6 / 3 | 3.3 | No |
| 522977 | 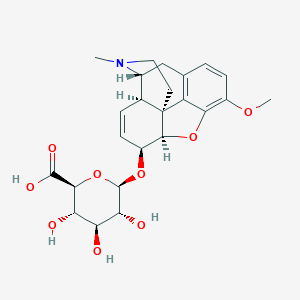 Codeine-6-glucuronide Codeine-6-glucuronide | C24H29NO9 | 475.494 | 10 / 4 | -2.5 | Yes |
| 523187 | 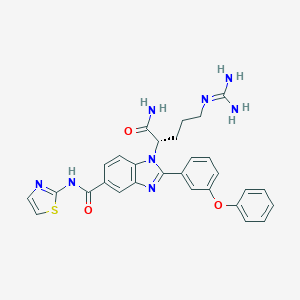 CHEMBL3718857 CHEMBL3718857 | C29H28N8O3S | 568.656 | 7 / 4 | 3.1 | No |
| 57400 | 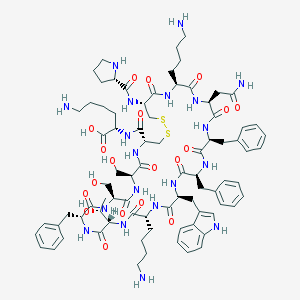 Cortistatin-14 Cortistatin-14 | C81H113N19O19S2 | 1721.03 | 25 / 23 | -2.7 | No |
| 523229 | 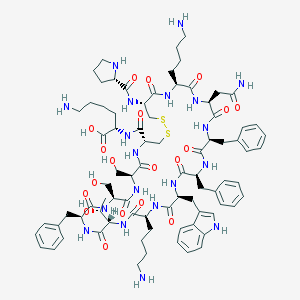 Cortistatin-14 Cortistatin-14 | C81H113N19O19S2 | 1721.03 | 25 / 23 | -2.7 | No |
| 523626 | 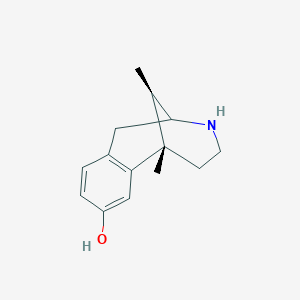 CHEMBL114203 CHEMBL114203 | C14H19NO | 217.312 | 2 / 2 | 1.3 | Yes |
| 523627 | 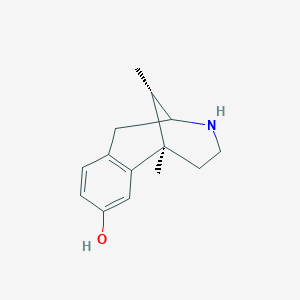 CHEMBL326370 CHEMBL326370 | C14H19NO | 217.312 | 2 / 2 | 1.3 | Yes |
| 553624 | 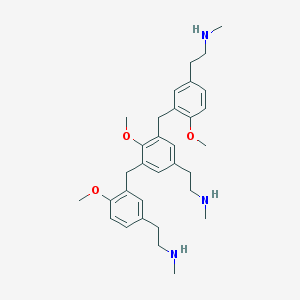 AC1L1EMH AC1L1EMH | C32H45N3O3 | 519.73 | 6 / 3 | 5.2 | No |
| 523649 | 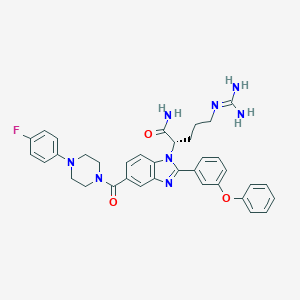 CHEMBL3717645 CHEMBL3717645 | C36H37FN8O3 | 648.743 | 7 / 3 | 4.0 | No |
| 559595 | 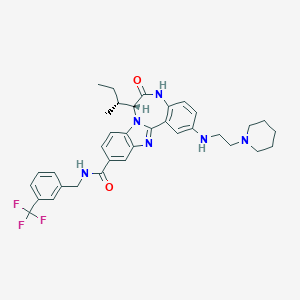 CHEMBL505263 CHEMBL505263 | C35H39F3N6O2 | 632.732 | 8 / 3 | 6.4 | No |
| 559596 | 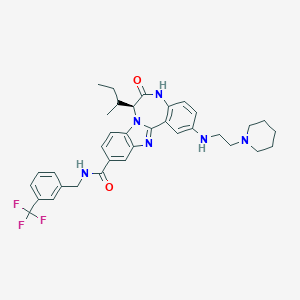 CHEMBL505263 CHEMBL505263 | C35H39F3N6O2 | 632.732 | 8 / 3 | 6.4 | No |
| 559861 | 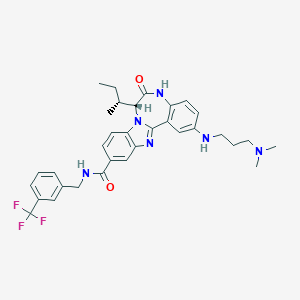 CHEMBL502132 CHEMBL502132 | C33H37F3N6O2 | 606.694 | 8 / 3 | 5.9 | No |
| 559863 | 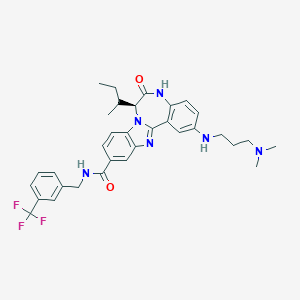 CHEMBL502132 CHEMBL502132 | C33H37F3N6O2 | 606.694 | 8 / 3 | 5.9 | No |
| 524002 | 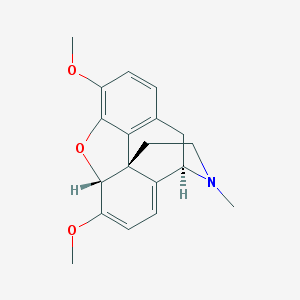 THEBAINE THEBAINE | C19H21NO3 | 311.381 | 4 / 0 | 2.2 | Yes |
| 560177 | 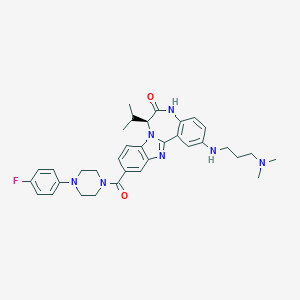 CHEMBL501027 CHEMBL501027 | C34H40FN7O2 | 597.739 | 7 / 2 | 5.1 | No |
| 553726 | 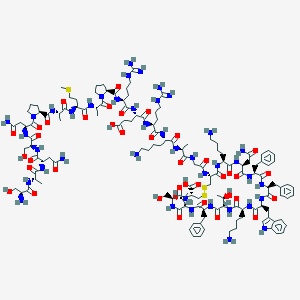 Somatostatin 28 Somatostatin 28 | C137H207N41O39S3 | 3148.59 | 48 / 46 | -15.2 | No |
| 560402 | 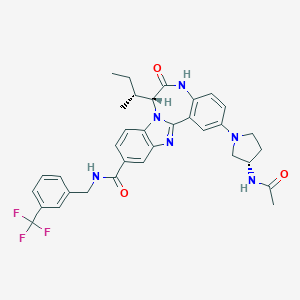 CHEMBL503202 CHEMBL503202 | C34H35F3N6O3 | 632.688 | 8 / 3 | 5.2 | No |
| 560405 | 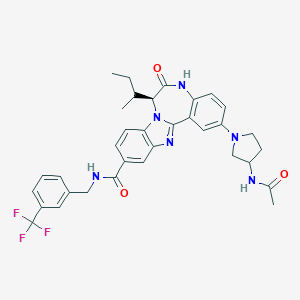 CHEMBL503202 CHEMBL503202 | C34H35F3N6O3 | 632.688 | 8 / 3 | 5.2 | No |
| 524470 | 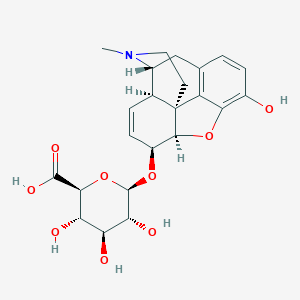 Morphine-6-glucuronide Morphine-6-glucuronide | C23H27NO9 | 461.467 | 10 / 5 | -2.9 | Yes |
| 560462 | 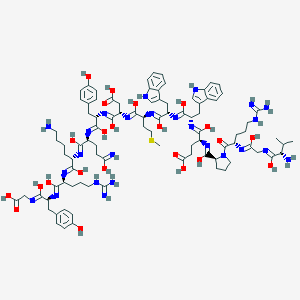 BAM(8-22) BAM(8-22) | C91H127N25O23S | 1971.23 | 42 / 30 | 1.4 | No |
| 524563 | 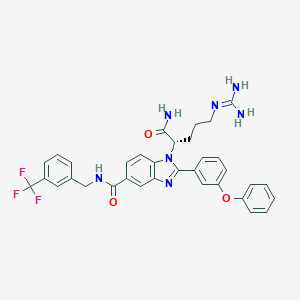 CHEMBL3717941 CHEMBL3717941 | C34H32F3N7O3 | 643.671 | 8 / 4 | 4.5 | No |
| 524770 | 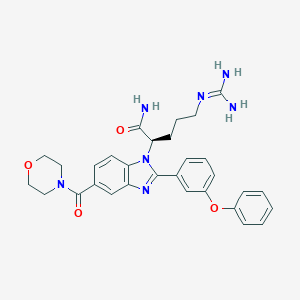 CHEMBL3719296 CHEMBL3719296 | C30H33N7O4 | 555.639 | 6 / 3 | 2.0 | No |
| 524918 | 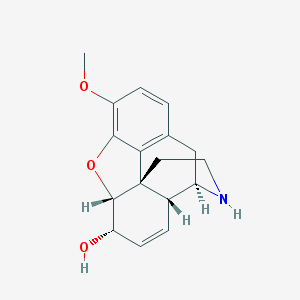 Norcodeine Norcodeine | C17H19NO3 | 285.343 | 4 / 2 | 0.7 | Yes |
| 525367 | 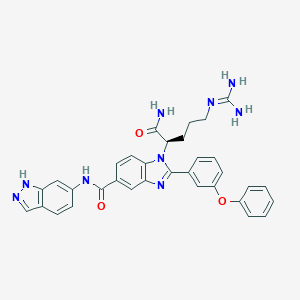 CHEMBL3716995 CHEMBL3716995 | C33H31N9O3 | 601.671 | 6 / 5 | 3.3 | No |
| 525494 | 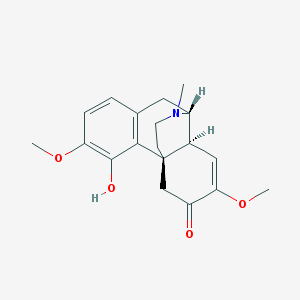 AC1LQMEV AC1LQMEV | C19H23NO4 | 329.396 | 5 / 1 | 2.2 | Yes |
| 525741 | 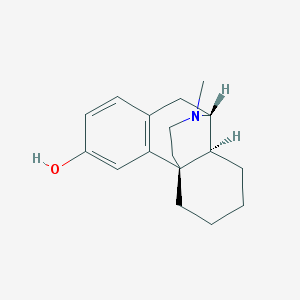 125-73-5 125-73-5 | C17H23NO | 257.377 | 2 / 1 | 3.1 | Yes |
| 525742 | 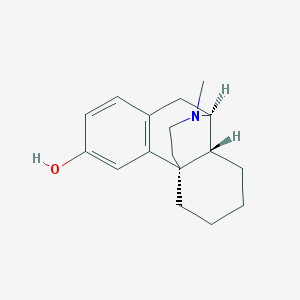 ZINC2433 ZINC2433 | C17H23NO | 257.377 | 2 / 1 | 3.1 | Yes |
| 525828 | 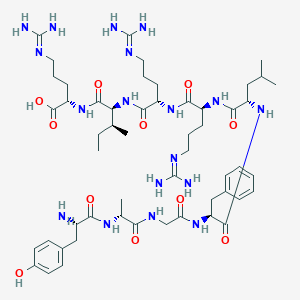 YGGFLRRXR YGGFLRRXR | C53H86N18O11 | 1151.39 | 15 / 17 | -4.8 | No |
| 525882 | 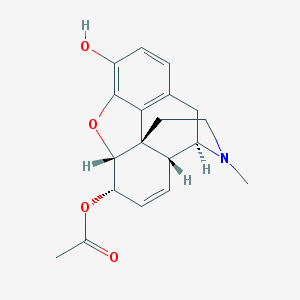 6-Acetylmorphine 6-Acetylmorphine | C19H21NO4 | 327.38 | 5 / 1 | 1.3 | Yes |
| 562142 |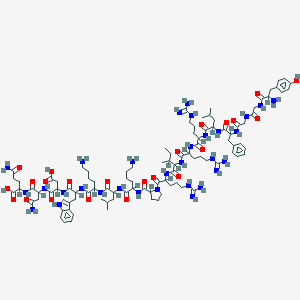  Dynorphins Dynorphins | C99H155N31O23 | 2147.52 | 29 / 33 | -7.8 | No |
| 526695 | 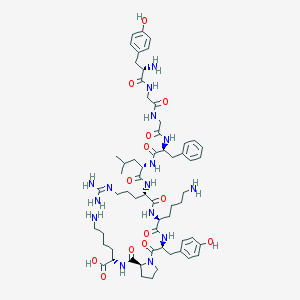 alpha-Neoendorphin alpha-Neoendorphin | C60H89N15O13 | 1228.46 | 17 / 16 | -2.1 | No |
| 526812 | 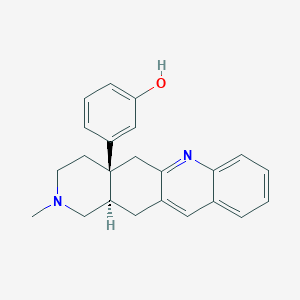 TAN-67 TAN-67 | C23H24N2O | 344.458 | 3 / 1 | 4.3 | Yes |
| 526813 | 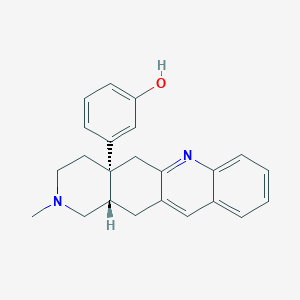 (+)-TAN-67 (+)-TAN-67 | C23H24N2O | 344.458 | 3 / 1 | 4.3 | Yes |
| 526973 | 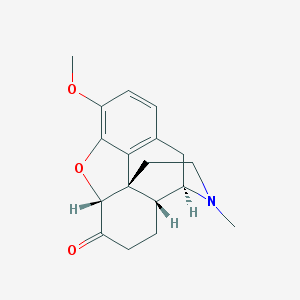 HYDROCODONE HYDROCODONE | C18H21NO3 | 299.37 | 4 / 0 | 2.2 | Yes |
| 527416 | 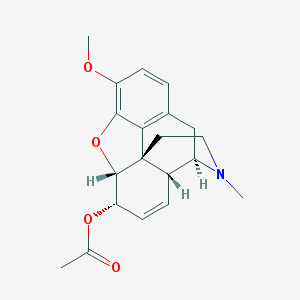 Acetylcodeine Acetylcodeine | C20H23NO4 | 341.407 | 5 / 0 | 1.7 | Yes |
| 527486 | 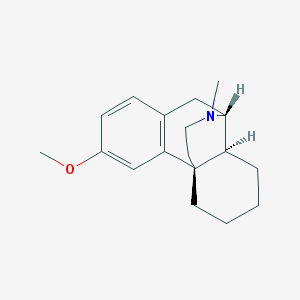 125-71-3 125-71-3 | C18H25NO | 271.404 | 2 / 0 | 3.4 | Yes |
| 564173 | 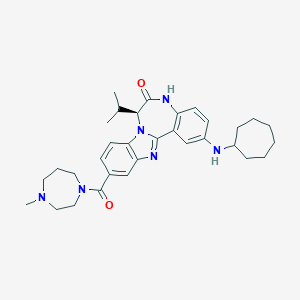 CHEMBL463274 CHEMBL463274 | C32H42N6O2 | 542.728 | 5 / 2 | 5.5 | No |
| 564174 | 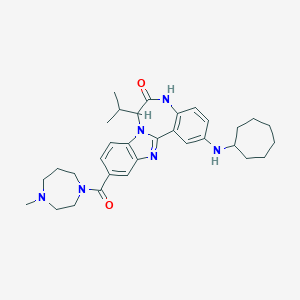 CHEMBL463274 CHEMBL463274 | C32H42N6O2 | 542.728 | 5 / 2 | 5.5 | No |
| 528616 | 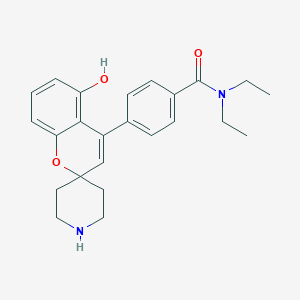 UNII-FA554TW3UP UNII-FA554TW3UP | C24H28N2O3 | 392.499 | 4 / 2 | 3.2 | Yes |
| 528648 | 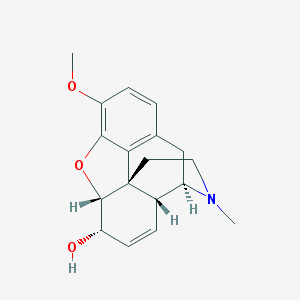 codeine codeine | C18H21NO3 | 299.37 | 4 / 1 | 1.1 | Yes |
| 528649 | 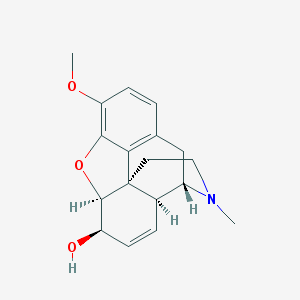 (+)-codeine (+)-codeine | C18H21NO3 | 299.37 | 4 / 1 | 1.1 | Yes |
| 556441 | 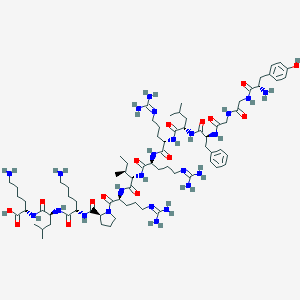 Dynorphin A (1-13) Dynorphin A (1-13) | C75H126N24O15 | 1603.99 | 21 / 22 | -4.2 | No |
| 565428 | 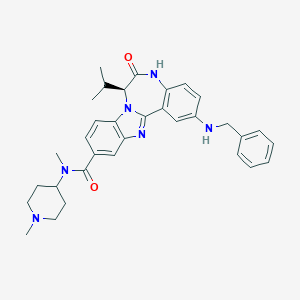 CHEMBL503688 CHEMBL503688 | C33H38N6O2 | 550.707 | 5 / 2 | 5.1 | No |
| 565429 | 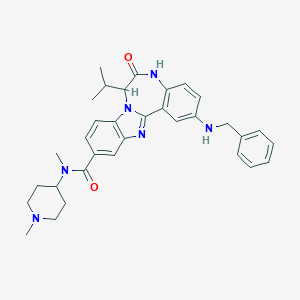 CHEMBL503688 CHEMBL503688 | C33H38N6O2 | 550.707 | 5 / 2 | 5.1 | No |
| 530811 | 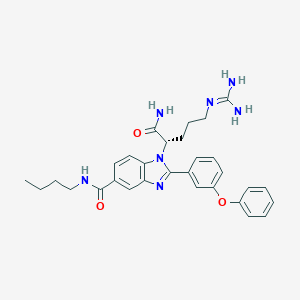 CHEMBL3715864 CHEMBL3715864 | C30H35N7O3 | 541.656 | 5 / 4 | 3.4 | No |
| 531141 | 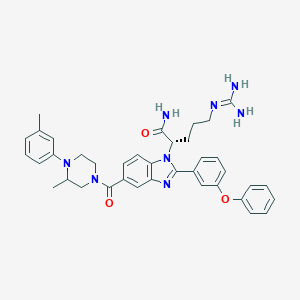 CHEMBL3716029 CHEMBL3716029 | C38H42N8O3 | 658.807 | 6 / 3 | 4.7 | No |
| 531235 | 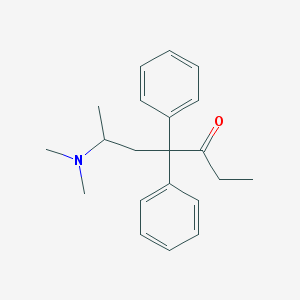 methadone methadone | C21H27NO | 309.453 | 2 / 0 | 3.9 | Yes |
| 568655 | 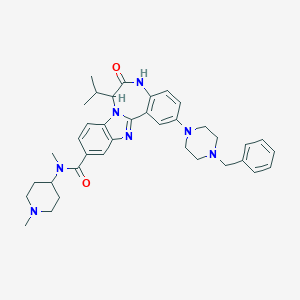 CHEMBL502480 CHEMBL502480 | C37H45N7O2 | 619.814 | 6 / 1 | 5.0 | No |
| 568656 | 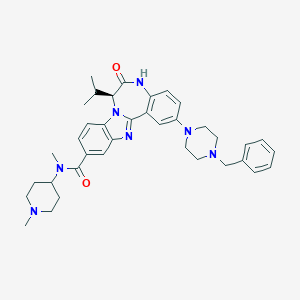 CHEMBL502480 CHEMBL502480 | C37H45N7O2 | 619.814 | 6 / 1 | 5.0 | No |
| 531855 | 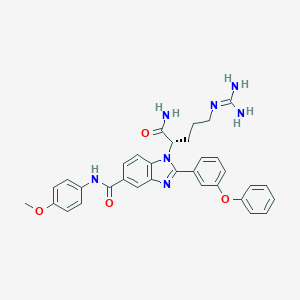 CHEMBL3716921 CHEMBL3716921 | C33H33N7O4 | 591.672 | 6 / 4 | 3.7 | No |
| 531881 | 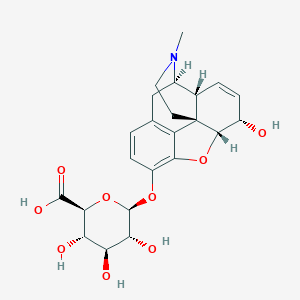 Morphine-3-glucuronide Morphine-3-glucuronide | C23H27NO9 | 461.467 | 10 / 5 | -3.0 | Yes |
| 532045 | 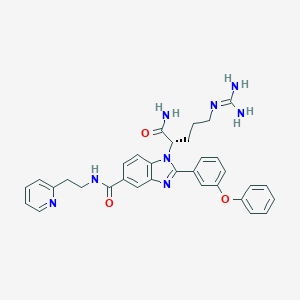 CHEMBL3717681 CHEMBL3717681 | C33H34N8O3 | 590.688 | 6 / 4 | 3.1 | No |
| 555253 | 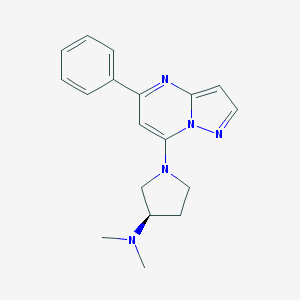 (R)-ZINC-3573 (R)-ZINC-3573 | C18H21N5 | 307.401 | 4 / 0 | 2.7 | Yes |
| 532656 | 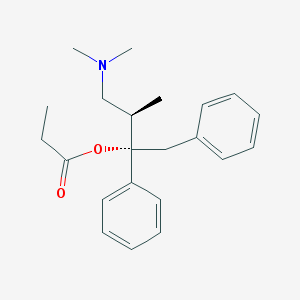 Dextropropoxyphene Dextropropoxyphene | C22H29NO2 | 339.479 | 3 / 0 | 4.2 | Yes |
| 569763 | 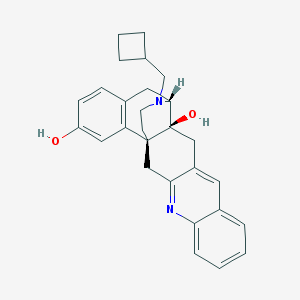 CHEMBL1270383 CHEMBL1270383 | C28H30N2O2 | 426.56 | 4 / 2 | 4.7 | Yes |
| 533120 | 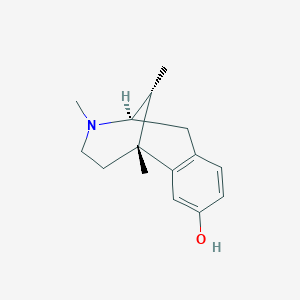 (-)-metazocine (-)-metazocine | C15H21NO | 231.339 | 2 / 1 | 1.8 | Yes |
| 533121 | 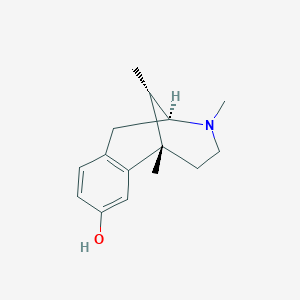 (+)-Metazocine (+)-Metazocine | C15H21NO | 231.339 | 2 / 1 | 1.8 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218