

You can:
| Name | Lysophosphatidic acid receptor 4 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | LPAR4 |
| Synonym | Purinergic receptor 9 P2Y9 P2Y5-like receptor P2Y purinoceptor 9 P2RY9 [ Show all ] |
| Disease | N/A |
| Length | 370 |
| Amino acid sequence | MGDRRFIDFQFQDSNSSLRPRLGNATANNTCIVDDSFKYNLNGAVYSVVFILGLITNSVSLFVFCFRMKMRSETAIFITNLAVSDLLFVCTLPFKIFYNFNRHWPFGDTLCKISGTAFLTNIYGSMLFLTCISVDRFLAIVYPFRSRTIRTRRNSAIVCAGVWILVLSGGISASLFSTTNVNNATTTCFEGFSKRVWKTYLSKITIFIEVVGFIIPLILNVSCSSVVLRTLRKPATLSQIGTNKKKVLKMITVHMAVFVVCFVPYNSVLFLYALVRSQAITNCFLERFAKIMYPITLCLATLNCCFDPFIYYFTLESFQKSFYINAHIRMESLFKTETPLTTKPSLPAIQEEVSDQTTNNGGELMLESTF |
| UniProt | Q99677 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | Q99677 |
| 3D structure model | This predicted structure model is from GPCR-EXP Q99677. |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL5968 |
| IUPHAR | 94 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 553285 | 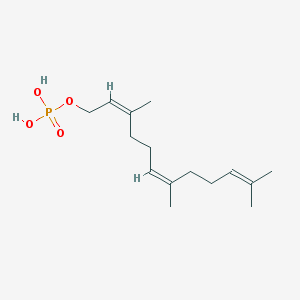 Farnesyl monophosphate Farnesyl monophosphate | C15H27O4P | 302.351 | 4 / 2 | 3.7 | Yes |
| 17530 | 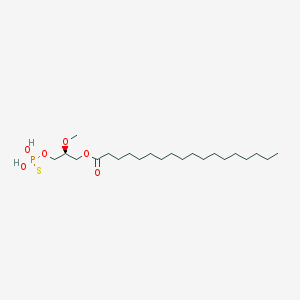 CHEMBL357053 CHEMBL357053 | C22H45O6PS | 468.63 | 7 / 2 | 8.4 | No |
| 17539 | 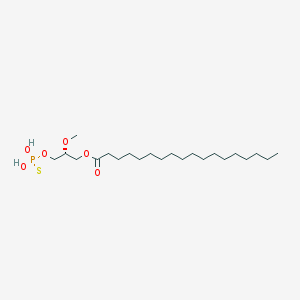 CHEMBL153043 CHEMBL153043 | C22H45O6PS | 468.63 | 7 / 2 | 8.4 | No |
| 17544 | 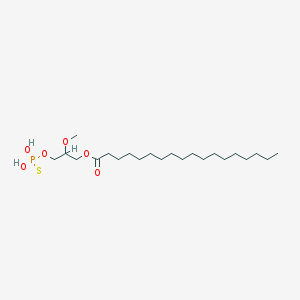 CHEMBL2017139 CHEMBL2017139 | C22H45O6PS | 468.63 | 7 / 2 | 8.4 | No |
| 162859 | 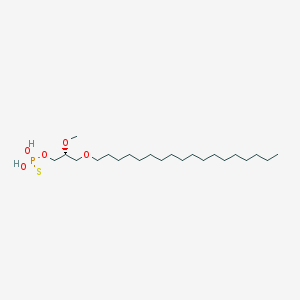 CHEMBL2335048 CHEMBL2335048 | C22H47O5PS | 454.647 | 6 / 2 | 8.8 | No |
| 162861 | 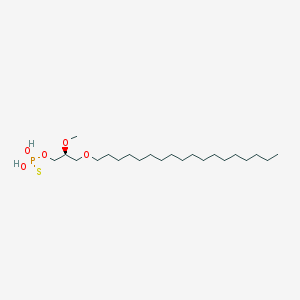 CHEMBL2335047 CHEMBL2335047 | C22H47O5PS | 454.647 | 6 / 2 | 8.8 | No |
| 162870 | 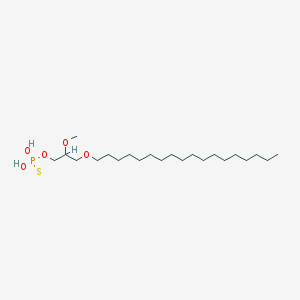 CHEMBL2335052 CHEMBL2335052 | C22H47O5PS | 454.647 | 6 / 2 | 8.8 | No |
| 163623 | 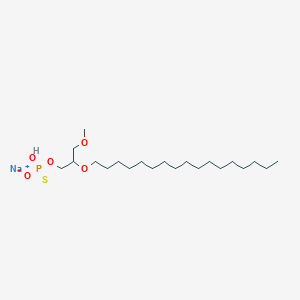 CHEMBL2335050 CHEMBL2335050 | C21H44NaO5PS | 462.602 | 6 / 1 | N/A | N/A |
| 178097 | 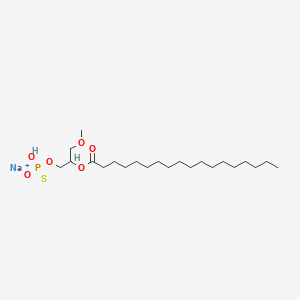 CHEMBL2335051 CHEMBL2335051 | C22H44NaO6PS | 490.612 | 7 / 1 | N/A | N/A |
| 486636 | 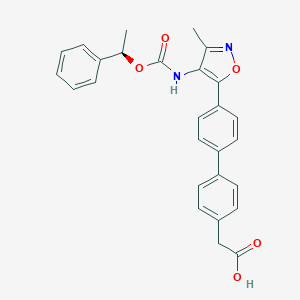 AM095 free acid AM095 free acid | C27H24N2O5 | 456.498 | 6 / 2 | 5.0 | Yes |
| 278364 | 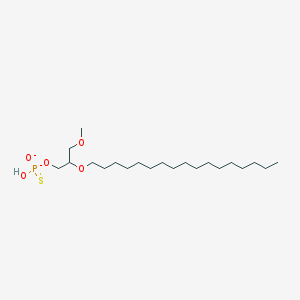 CHEMBL2335050 CHEMBL2335050 | C21H44O5PS- | 439.612 | 6 / 1 | 8.2 | No |
| 290484 | 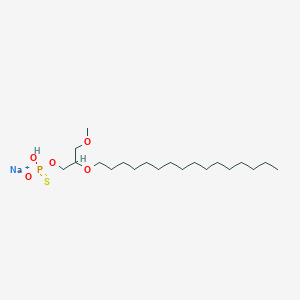 CHEMBL2335049 CHEMBL2335049 | C20H42NaO5PS | 448.575 | 6 / 1 | N/A | N/A |
| 356928 | 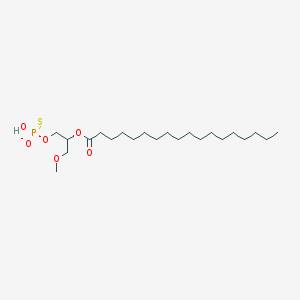 CHEMBL2335051 CHEMBL2335051 | C22H44O6PS- | 467.622 | 7 / 1 | 8.3 | No |
| 555123 | 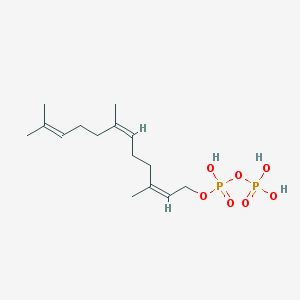 2-cis,6-cis-farnesyl diphosphate 2-cis,6-cis-farnesyl diphosphate | C15H28O7P2 | 382.33 | 7 / 3 | 2.6 | Yes |
| 380363 | 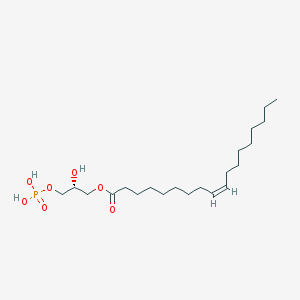 1-Oleoyl Lysophosphatidic Acid 1-Oleoyl Lysophosphatidic Acid | C21H41O7P | 436.526 | 7 / 3 | 5.2 | No |
| 555181 | 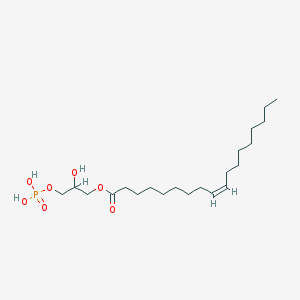 lysophosphatidic acid lysophosphatidic acid | C21H41O7P | 436.526 | 7 / 3 | 5.2 | No |
| 555215 | 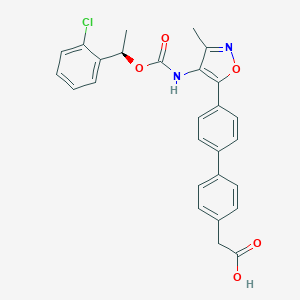 AM966 AM966 | C27H23ClN2O5 | 490.94 | 6 / 2 | 5.6 | No |
| 388658 | 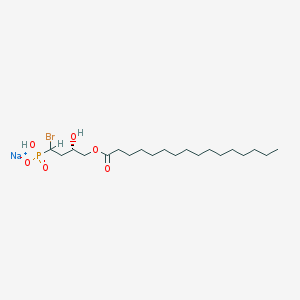 [1-bromo-(3S)-hydrox-4-(palmitoyloxy)butyl]phosphate [1-bromo-(3S)-hydrox-4-(palmitoyloxy)butyl]phosphate | C20H39BrNaO6P | 509.394 | 6 / 2 | N/A | No |
| 414304 | 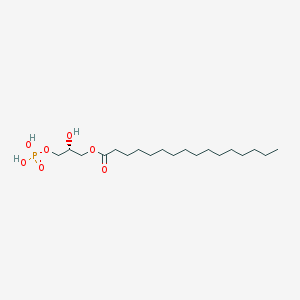 1-Hexadecanoyl-sn-glycero-3-phosphate 1-Hexadecanoyl-sn-glycero-3-phosphate | C19H39O7P | 410.488 | 7 / 3 | 5.0 | Yes |
| 514522 | 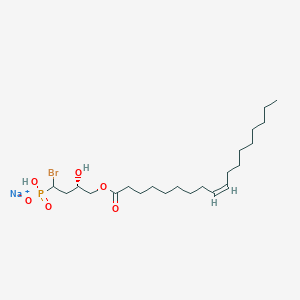 CHEMBL3621353 CHEMBL3621353 | C22H41BrNaO6P | 535.432 | 6 / 2 | N/A | No |
| 515774 | 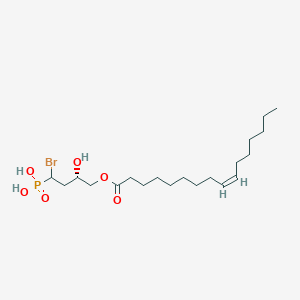 CHEMBL3621964 CHEMBL3621964 | C20H38BrO6P | 485.396 | 6 / 3 | 5.3 | No |
| 435194 | 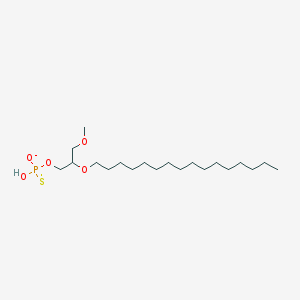 CHEMBL2335049 CHEMBL2335049 | C20H42O5PS- | 425.585 | 6 / 1 | 7.6 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218