

You can:
| Name | Succinate receptor 1 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | SUCNR1 |
| Synonym | succinate receptor 1 succinate receptor P2Y purinoceptor 1-like P2Y purinoceptor 1 GPR91 [ Show all ] |
| Disease | N/A |
| Length | 334 |
| Amino acid sequence | MLGIMAWNATCKNWLAAEAALEKYYLSIFYGIEFVVGVLGNTIVVYGYIFSLKNWNSSNIYLFNLSVSDLAFLCTLPMLIRSYANGNWIYGDVLCISNRYVLHANLYTSILFLTFISIDRYLIIKYPFREHLLQKKEFAILISLAIWVLVTLELLPILPLINPVITDNGTTCNDFASSGDPNYNLIYSMCLTLLGFLIPLFVMCFFYYKIALFLKQRNRQVATALPLEKPLNLVIMAVVIFSVLFTPYHVMRNVRIASRLGSWKQYQCTQVVINSFYIVTRPLAFLNSVINPVFYFLLGDHFRDMLMNQLRHNFKSLTSFSRWAHELLLSFREK |
| UniProt | Q9BXA5 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | Q9BXA5 |
| 3D structure model | This predicted structure model is from GPCR-EXP Q9BXA5. |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL2150838 |
| IUPHAR | 166 |
| DrugBank | BE0002258 |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 12676 | 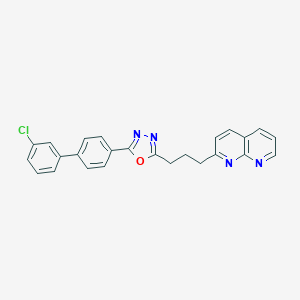 CHEMBL2153586 CHEMBL2153586 | C25H19ClN4O | 426.904 | 5 / 0 | 5.8 | No |
| 13164 | 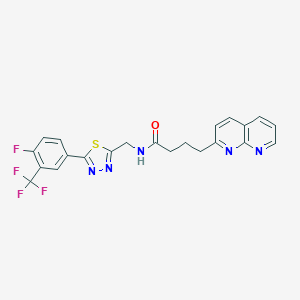 CHEMBL2153585 CHEMBL2153585 | C22H17F4N5OS | 475.466 | 10 / 1 | 4.0 | Yes |
| 17287 | 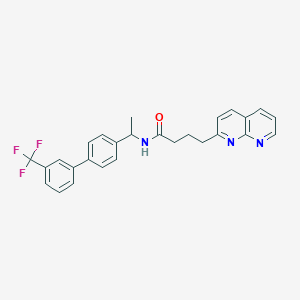 CHEMBL2153445 CHEMBL2153445 | C27H24F3N3O | 463.504 | 6 / 1 | 5.8 | No |
| 18005 | 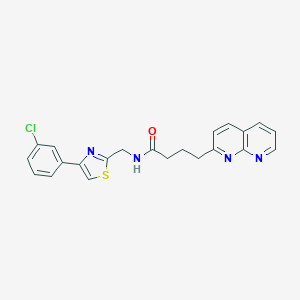 CHEMBL2153583 CHEMBL2153583 | C22H19ClN4OS | 422.931 | 5 / 1 | 4.2 | Yes |
| 23956 | 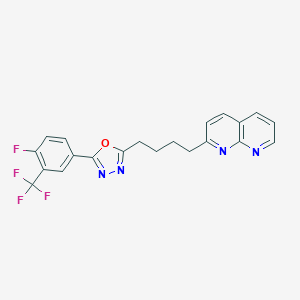 CHEMBL2153597 CHEMBL2153597 | C21H16F4N4O | 416.38 | 9 / 0 | 4.9 | Yes |
| 44766 | 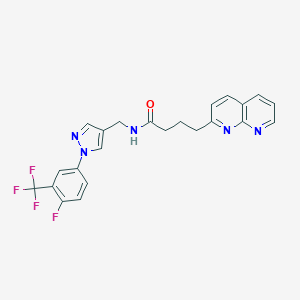 CHEMBL2153472 CHEMBL2153472 | C23H19F4N5O | 457.433 | 8 / 1 | 3.8 | Yes |
| 50278 | 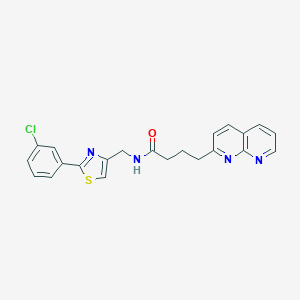 CHEMBL2153582 CHEMBL2153582 | C22H19ClN4OS | 422.931 | 5 / 1 | 4.2 | Yes |
| 553514 | 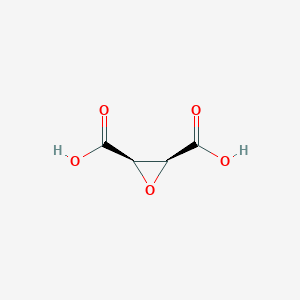 cis-Epoxysuccinic acid cis-Epoxysuccinic acid | C4H4O5 | 132.071 | 5 / 2 | -0.9 | Yes |
| 58115 | 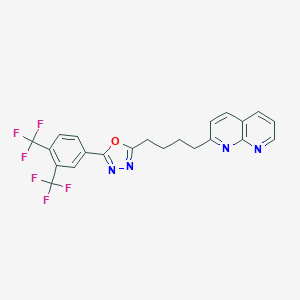 CHEMBL2153594 CHEMBL2153594 | C22H16F6N4O | 466.387 | 11 / 0 | 5.7 | No |
| 70441 | 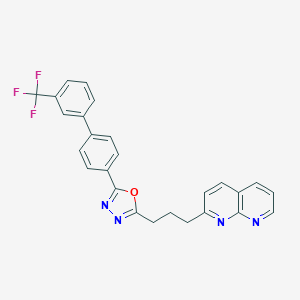 CHEMBL2153588 CHEMBL2153588 | C26H19F3N4O | 460.46 | 8 / 0 | 6.1 | No |
| 80859 | 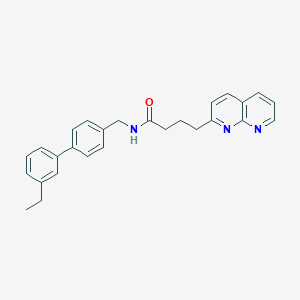 CHEMBL2153444 CHEMBL2153444 | C27H27N3O | 409.533 | 3 / 1 | 5.3 | No |
| 82835 | 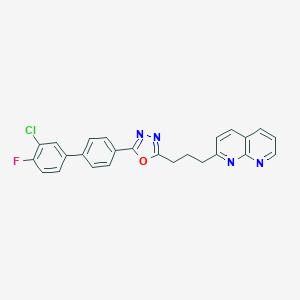 CHEMBL2153587 CHEMBL2153587 | C25H18ClFN4O | 444.894 | 6 / 0 | 5.9 | No |
| 115223 | 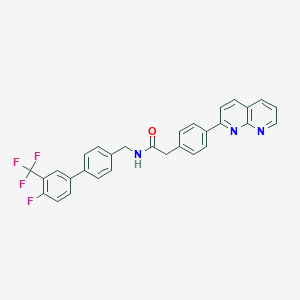 CHEMBL2153460 CHEMBL2153460 | C30H21F4N3O | 515.512 | 7 / 1 | 6.4 | No |
| 131167 | 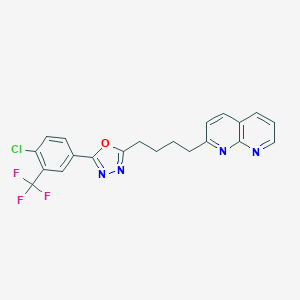 CHEMBL2153596 CHEMBL2153596 | C21H16ClF3N4O | 432.831 | 8 / 0 | 5.5 | No |
| 132335 | 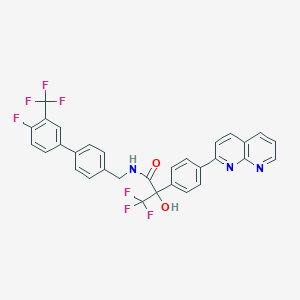 CHEMBL2153463 CHEMBL2153463 | C31H20F7N3O2 | 599.509 | 11 / 2 | 6.8 | No |
| 140135 | 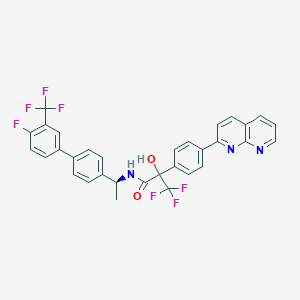 CHEMBL2153468 CHEMBL2153468 | C32H22F7N3O2 | 613.536 | 11 / 2 | 7.2 | No |
| 141049 | 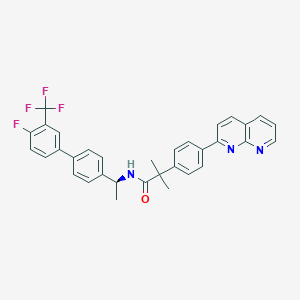 CHEMBL2153465 CHEMBL2153465 | C33H27F4N3O | 557.593 | 7 / 1 | 7.8 | No |
| 144740 | 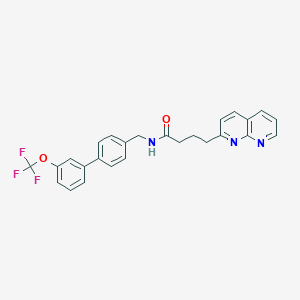 CHEMBL2153448 CHEMBL2153448 | C26H22F3N3O2 | 465.476 | 7 / 1 | 5.7 | No |
| 147354 | 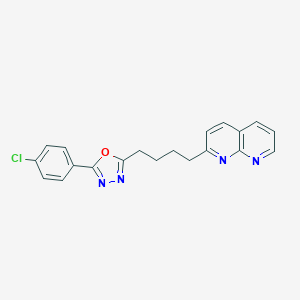 CHEMBL2153593 CHEMBL2153593 | C20H17ClN4O | 364.833 | 5 / 0 | 4.6 | Yes |
| 150673 | 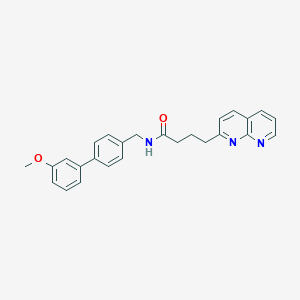 CHEMBL2153447 CHEMBL2153447 | C26H25N3O2 | 411.505 | 4 / 1 | 4.4 | Yes |
| 163866 | 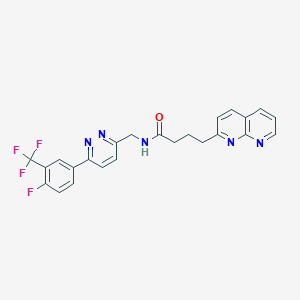 CHEMBL2153471 CHEMBL2153471 | C24H19F4N5O | 469.444 | 9 / 1 | 3.5 | Yes |
| 166811 | 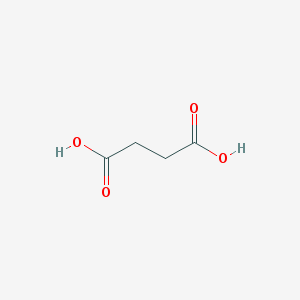 succinic acid succinic acid | C4H6O4 | 118.088 | 4 / 2 | -0.6 | Yes |
| 174196 | 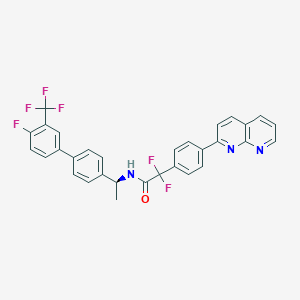 CHEMBL2153466 CHEMBL2153466 | C31H21F6N3O | 565.519 | 9 / 1 | 7.6 | No |
| 184492 | 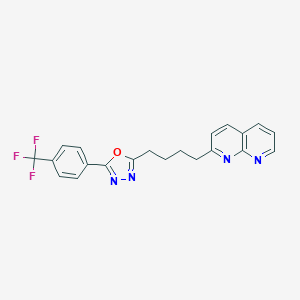 CHEMBL2153591 CHEMBL2153591 | C21H17F3N4O | 398.389 | 8 / 0 | 4.8 | Yes |
| 191500 | 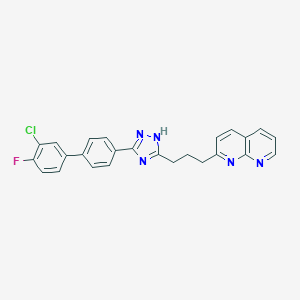 CHEMBL2153590 CHEMBL2153590 | C25H19ClFN5 | 443.91 | 5 / 1 | 6.2 | No |
| 216692 | 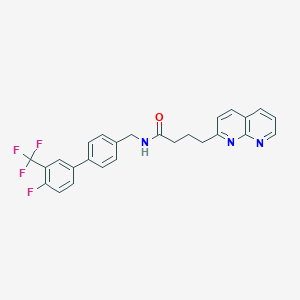 CHEMBL2153441 CHEMBL2153441 | C26H21F4N3O | 467.468 | 7 / 1 | 5.5 | No |
| 218522 | 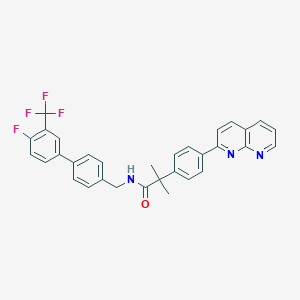 CHEMBL2153462 CHEMBL2153462 | C32H25F4N3O | 543.566 | 7 / 1 | 7.4 | No |
| 247144 | 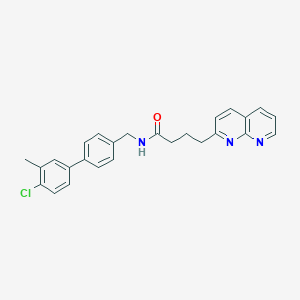 CHEMBL2153457 CHEMBL2153457 | C26H24ClN3O | 429.948 | 3 / 1 | 5.5 | No |
| 257268 | 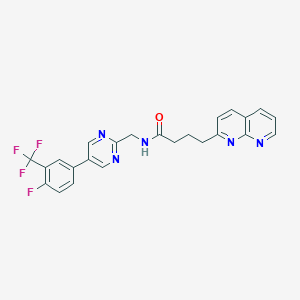 CHEMBL2153470 CHEMBL2153470 | C24H19F4N5O | 469.444 | 9 / 1 | 3.8 | Yes |
| 261233 | 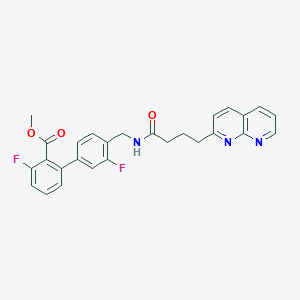 CHEMBL2153436 CHEMBL2153436 | C27H23F2N3O3 | 475.496 | 7 / 1 | 4.5 | Yes |
| 265459 | 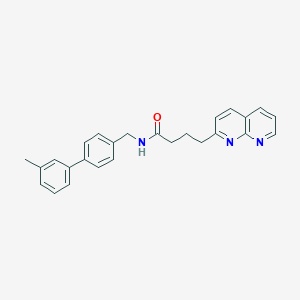 CHEMBL2153439 CHEMBL2153439 | C26H25N3O | 395.506 | 3 / 1 | 4.8 | Yes |
| 267338 | 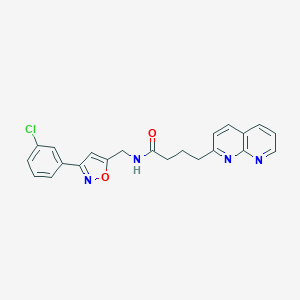 CHEMBL2153579 CHEMBL2153579 | C22H19ClN4O2 | 406.87 | 5 / 1 | 3.6 | Yes |
| 554643 | 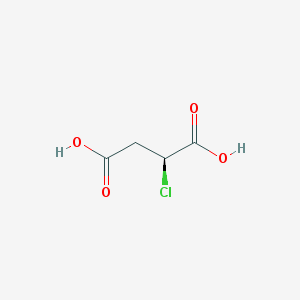 (S)-2-chlorosuccinic acid (S)-2-chlorosuccinic acid | C4H5ClO4 | 152.53 | 4 / 2 | -0.1 | Yes |
| 290045 | 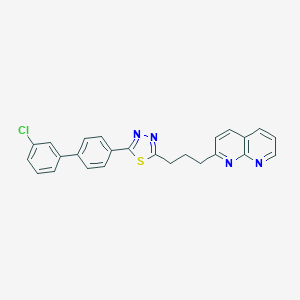 CHEMBL2153589 CHEMBL2153589 | C25H19ClN4S | 442.965 | 5 / 0 | 6.5 | No |
| 554729 | 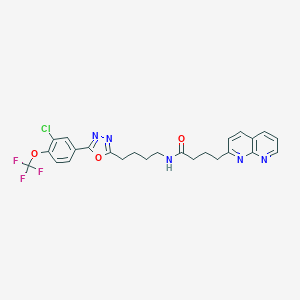 D0GC5Y D0GC5Y | C25H23ClF3N5O3 | 533.936 | 10 / 1 | 5.4 | No |
| 554747 | 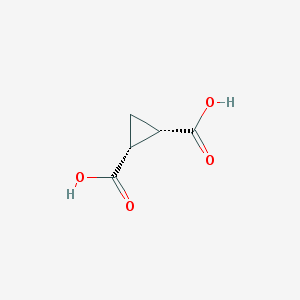 696-74-2 696-74-2 | C5H6O4 | 130.099 | 4 / 2 | -0.6 | Yes |
| 310814 | 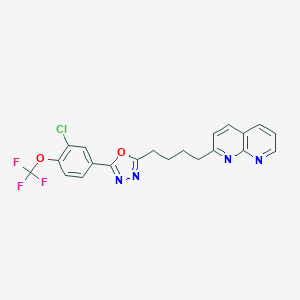 CHEMBL2153595 CHEMBL2153595 | C21H16ClF3N4O2 | 448.83 | 9 / 0 | 5.8 | No |
| 312688 | 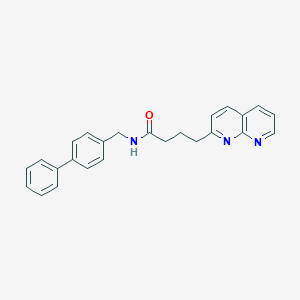 CHEMBL2153438 CHEMBL2153438 | C25H23N3O | 381.479 | 3 / 1 | 4.5 | Yes |
| 313673 | 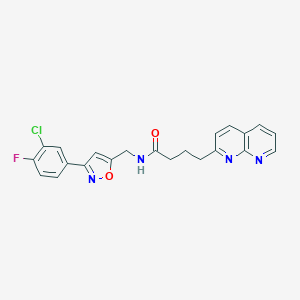 CHEMBL2153580 CHEMBL2153580 | C22H18ClFN4O2 | 424.86 | 6 / 1 | 3.7 | Yes |
| 322244 | 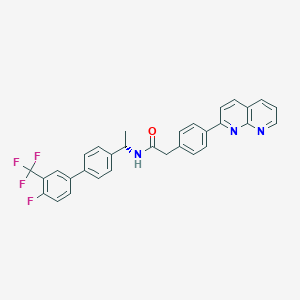 CHEMBL2153461 CHEMBL2153461 | C31H23F4N3O | 529.539 | 7 / 1 | 6.8 | No |
| 324370 | 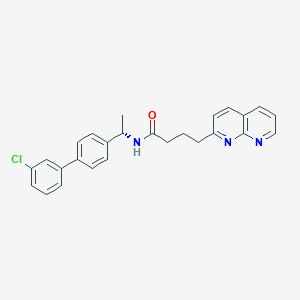 CHEMBL2153446 CHEMBL2153446 | C26H24ClN3O | 429.948 | 3 / 1 | 5.5 | No |
| 324963 | 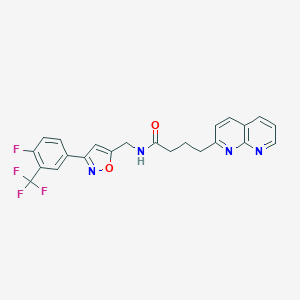 CHEMBL2153581 CHEMBL2153581 | C23H18F4N4O2 | 458.417 | 9 / 1 | 4.0 | Yes |
| 331759 | 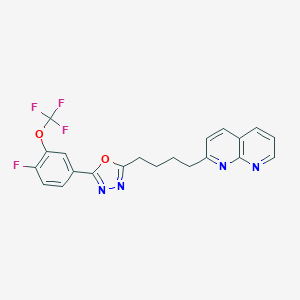 CHEMBL2153598 CHEMBL2153598 | C21H16F4N4O2 | 432.379 | 10 / 0 | 5.2 | No |
| 333665 | 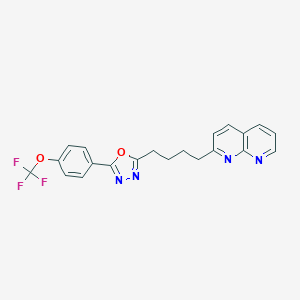 CHEMBL2153592 CHEMBL2153592 | C21H17F3N4O2 | 414.388 | 9 / 0 | 5.1 | No |
| 335238 | 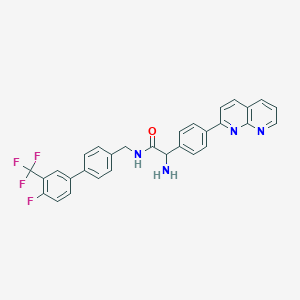 CHEMBL2153464 CHEMBL2153464 | C30H22F4N4O | 530.527 | 8 / 2 | 5.6 | No |
| 336808 | 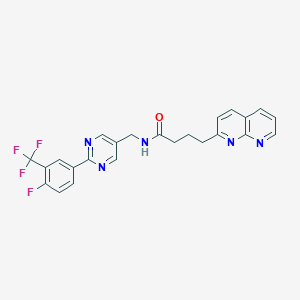 CHEMBL2153469 CHEMBL2153469 | C24H19F4N5O | 469.444 | 9 / 1 | 3.8 | Yes |
| 352318 | 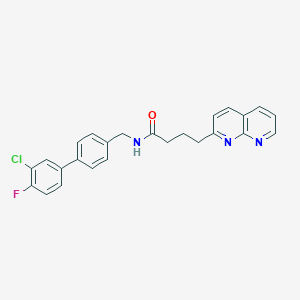 CHEMBL2153442 CHEMBL2153442 | C25H21ClFN3O | 433.911 | 4 / 1 | 5.2 | No |
| 353651 | 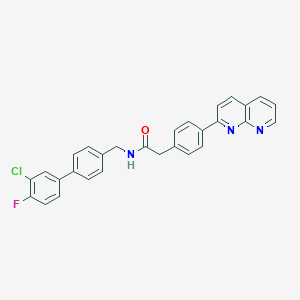 CHEMBL2153459 CHEMBL2153459 | C29H21ClFN3O | 481.955 | 4 / 1 | 6.2 | No |
| 555129 | 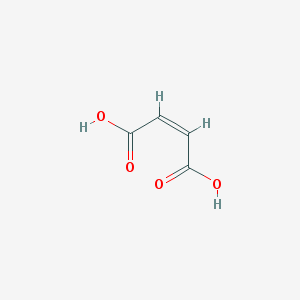 maleic acid maleic acid | C4H4O4 | 116.072 | 4 / 2 | -0.3 | Yes |
| 376635 | 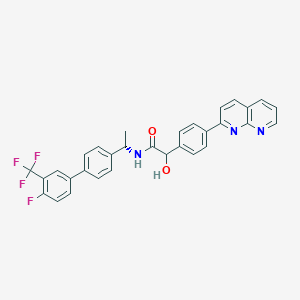 CHEMBL2153467 CHEMBL2153467 | C31H23F4N3O2 | 545.538 | 8 / 2 | 6.3 | No |
| 379902 | 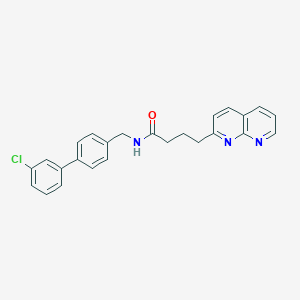 CHEMBL2153440 CHEMBL2153440 | C25H22ClN3O | 415.921 | 3 / 1 | 5.1 | No |
| 427096 | 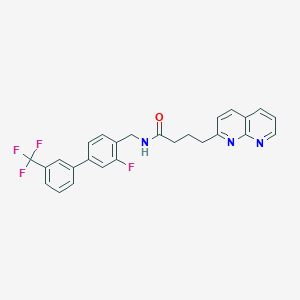 CHEMBL2153443 CHEMBL2153443 | C26H21F4N3O | 467.468 | 7 / 1 | 5.5 | No |
| 555422 | 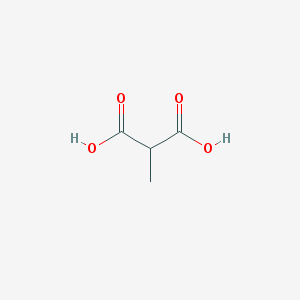 Methylmalonic acid Methylmalonic acid | C4H6O4 | 118.088 | 4 / 2 | 0.1 | Yes |
| 432115 | 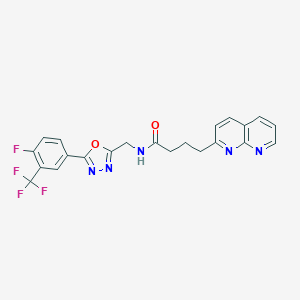 CHEMBL2153584 CHEMBL2153584 | C22H17F4N5O2 | 459.405 | 10 / 1 | 3.4 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218