

You can:
| Name | P2Y purinoceptor 12 |
|---|---|
| Species | Rattus norvegicus (Rat) |
| Gene | P2ry12 |
| Synonym | purinergic receptor P2Y P2YADP P2Y12 receptor P2Y12 platelet ADP receptor P2Y12 [ Show all ] |
| Disease | N/A for non-human GPCRs |
| Length | 343 |
| Amino acid sequence | MEVPGANATSANTTSIPGTSTLCSRDYKITQVLFPLLYTVLFFAGLITNSLAMRIFFQIRSKSNFIIFLKNTVISDLLMILTFPFKILSDAKLGAGHLRTLVCQVTSVTFYFTMYISISFLGLITIDRYLKTTRPFKTSSPSNLLGAKILSVAIWAFMFLLSLPNMILTNRRPKDKDITKCSFLKSEFGLVWHEIVNYICQVIFWINFLIVIVCYSLITKELYRSYVRTRGSAKAPKKRVNIKVFIIIAVFFICFVPFHFARIPYTLSQTRAVFDCNAENTLFYVKESTLWLTSLNACLDPFIYFFLCKSFRNSLMSMLRCSTSGANKKKGQEGGDPSEETPM |
| UniProt | Q9EPX4 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL2188 |
| IUPHAR | N/A |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 441829 | 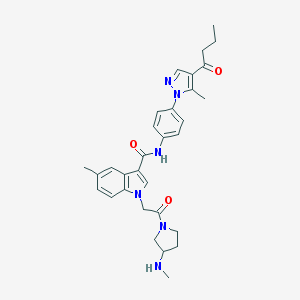 CHEMBL3325646 CHEMBL3325646 | C31H36N6O3 | 540.668 | 5 / 2 | 3.5 | No |
| 3057 | 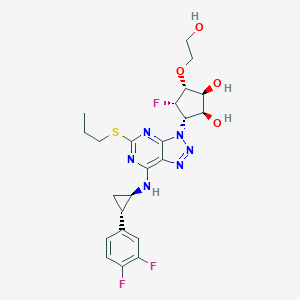 CHEMBL3098242 CHEMBL3098242 | C23H27F3N6O4S | 540.562 | 13 / 4 | 2.1 | No |
| 442389 | 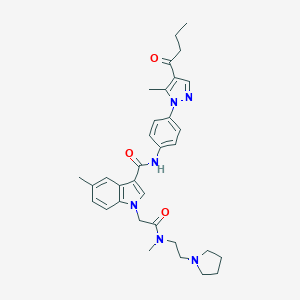 CHEMBL3325636 CHEMBL3325636 | C33H40N6O3 | 568.722 | 5 / 1 | 4.2 | No |
| 442437 | 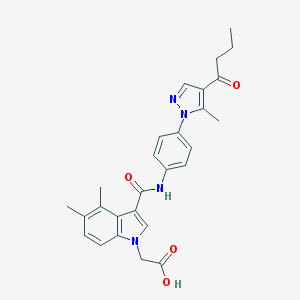 CHEMBL3325626 CHEMBL3325626 | C27H28N4O4 | 472.545 | 5 / 2 | 4.1 | Yes |
| 442551 | 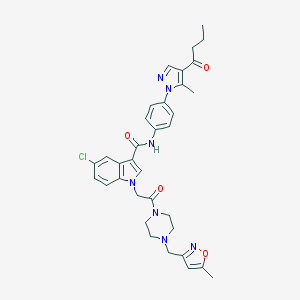 CHEMBL3325804 CHEMBL3325804 | C34H36ClN7O4 | 642.157 | 7 / 1 | 4.2 | No |
| 442649 | 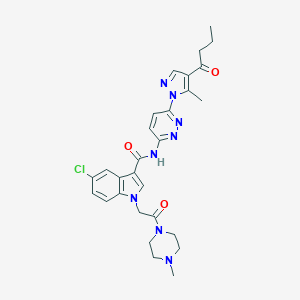 CHEMBL3325891 CHEMBL3325891 | C28H31ClN8O3 | 563.059 | 7 / 1 | 2.4 | No |
| 442777 | 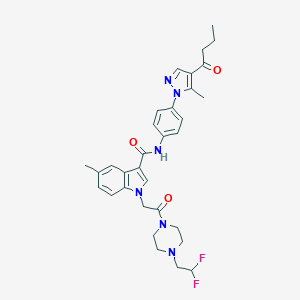 CHEMBL3325783 CHEMBL3325783 | C32H36F2N6O3 | 590.676 | 7 / 1 | 4.4 | No |
| 442782 | 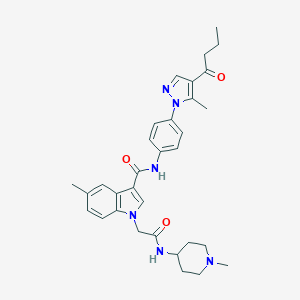 CHEMBL3325639 CHEMBL3325639 | C32H38N6O3 | 554.695 | 5 / 2 | 4.1 | No |
| 442917 | 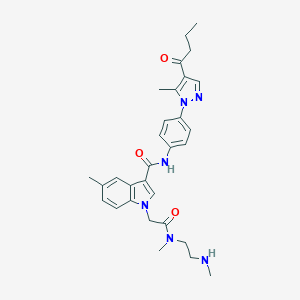 CHEMBL3325633 CHEMBL3325633 | C30H36N6O3 | 528.657 | 5 / 2 | 3.3 | No |
| 443055 | 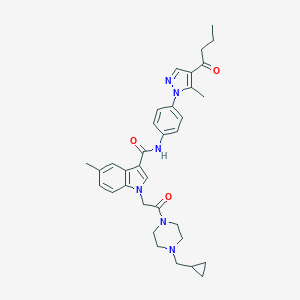 CHEMBL3325661 CHEMBL3325661 | C34H40N6O3 | 580.733 | 5 / 1 | 4.3 | No |
| 443070 | 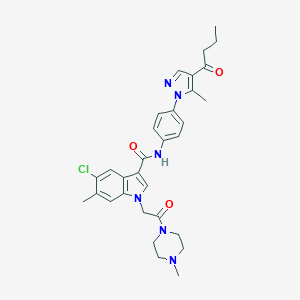 CHEMBL3325793 CHEMBL3325793 | C31H35ClN6O3 | 575.11 | 5 / 1 | 4.1 | No |
| 37598 | 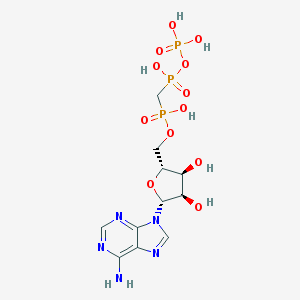 alpha,beta-Methylene ATP alpha,beta-Methylene ATP | C11H18N5O12P3 | 505.209 | 16 / 7 | -5.9 | No |
| 443207 | 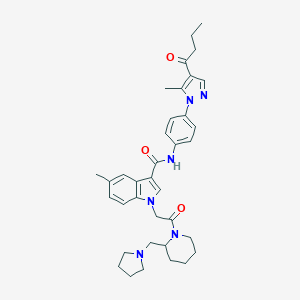 CHEMBL3325652 CHEMBL3325652 | C36H44N6O3 | 608.787 | 5 / 1 | 5.1 | No |
| 38815 | 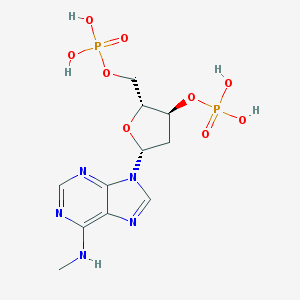 CHEMBL129841 CHEMBL129841 | C11H17N5O9P2 | 425.231 | 13 / 5 | -2.9 | No |
| 443481 | 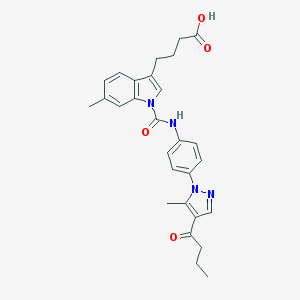 CHEMBL3326904 CHEMBL3326904 | C28H30N4O4 | 486.572 | 5 / 2 | 4.6 | Yes |
| 443559 | 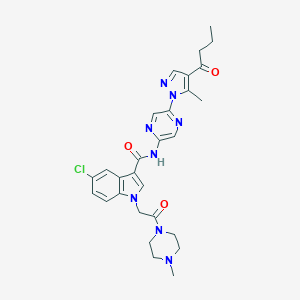 CHEMBL3325892 CHEMBL3325892 | C28H31ClN8O3 | 563.059 | 7 / 1 | 2.3 | No |
| 443636 | 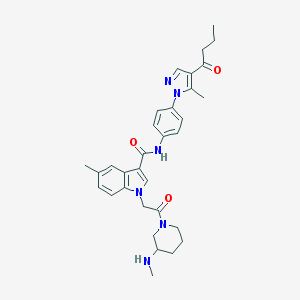 CHEMBL3325649 CHEMBL3325649 | C32H38N6O3 | 554.695 | 5 / 2 | 3.8 | No |
| 443682 | 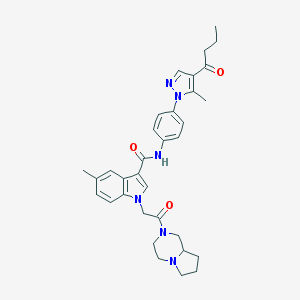 CHEMBL3325663 CHEMBL3325663 | C33H38N6O3 | 566.706 | 5 / 1 | 4.1 | No |
| 50882 | 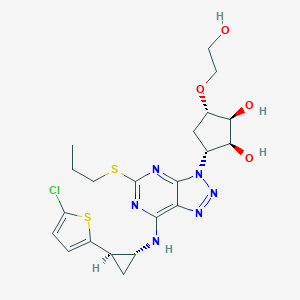 CHEMBL3098234 CHEMBL3098234 | C21H27ClN6O4S2 | 527.055 | 11 / 4 | 2.5 | No |
| 443845 | 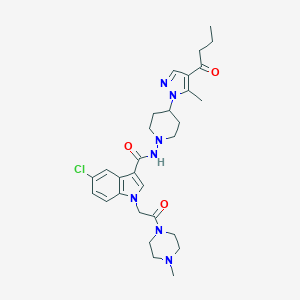 CHEMBL3325894 CHEMBL3325894 | C29H38ClN7O3 | 568.119 | 6 / 1 | 3.0 | No |
| 444170 | 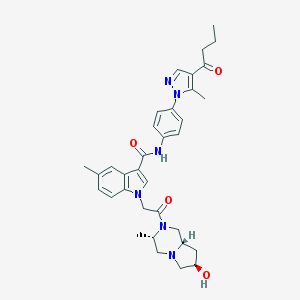 CHEMBL3325785 CHEMBL3325785 | C34H40N6O4 | 596.732 | 6 / 2 | 3.5 | No |
| 444337 | 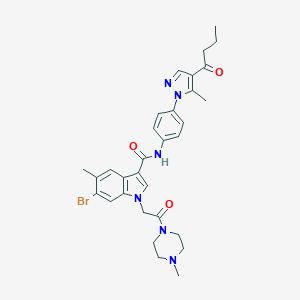 CHEMBL3325795 CHEMBL3325795 | C31H35BrN6O3 | 619.564 | 5 / 1 | 4.2 | No |
| 444572 | 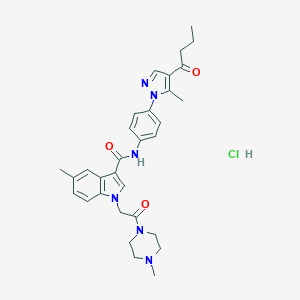 CHEMBL3325631 CHEMBL3325631 | C31H37ClN6O3 | 577.126 | 5 / 2 | N/A | No |
| 444628 | 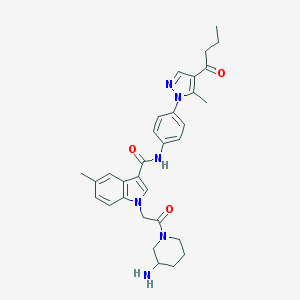 CHEMBL3325648 CHEMBL3325648 | C31H36N6O3 | 540.668 | 5 / 2 | 3.3 | No |
| 444706 | 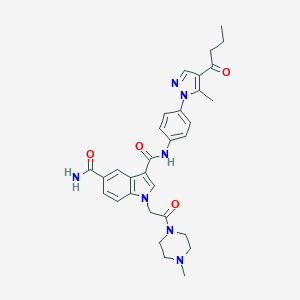 CHEMBL3325800 CHEMBL3325800 | C31H35N7O4 | 569.666 | 6 / 2 | 2.0 | No |
| 78872 | 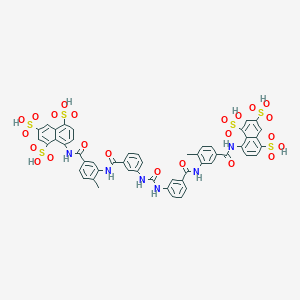 suramin suramin | C51H40N6O23S6 | 1297.26 | 23 / 12 | 1.5 | No |
| 444864 | 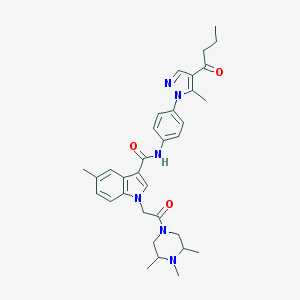 CHEMBL3325780 CHEMBL3325780 | C33H40N6O3 | 568.722 | 5 / 1 | 4.4 | No |
| 445217 | 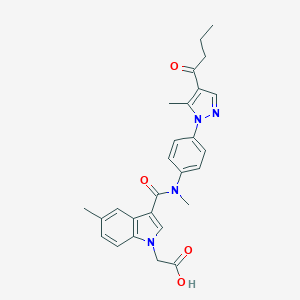 CHEMBL3326908 CHEMBL3326908 | C27H28N4O4 | 472.545 | 5 / 1 | 4.0 | Yes |
| 445222 | 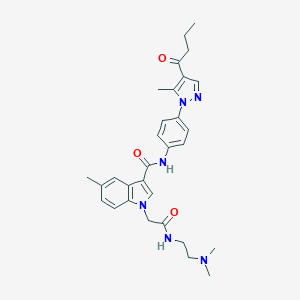 CHEMBL3325634 CHEMBL3325634 | C30H36N6O3 | 528.657 | 5 / 2 | 3.6 | No |
| 445464 | 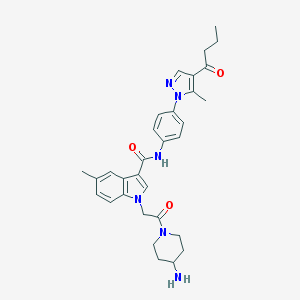 CHEMBL3325644 CHEMBL3325644 | C31H36N6O3 | 540.668 | 5 / 2 | 3.3 | No |
| 445696 | 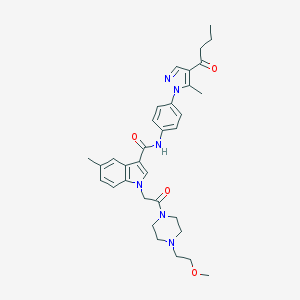 CHEMBL3325660 CHEMBL3325660 | C33H40N6O4 | 584.721 | 6 / 1 | 3.4 | No |
| 445741 | 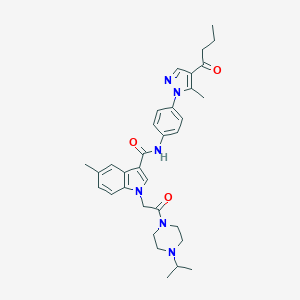 CHEMBL3325664 CHEMBL3325664 | C33H40N6O3 | 568.722 | 5 / 1 | 4.3 | No |
| 111331 | 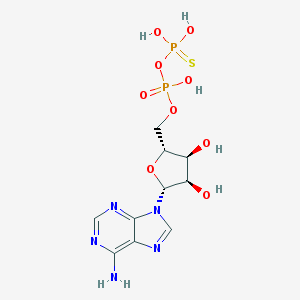 Adpbetas Adpbetas | C10H15N5O9P2S | 443.264 | 14 / 6 | -2.9 | No |
| 115225 | 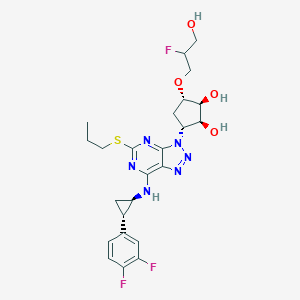 CHEMBL3098250 CHEMBL3098250 | C24H29F3N6O4S | 554.589 | 13 / 4 | 2.4 | No |
| 446744 | 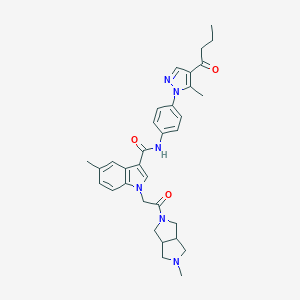 CHEMBL3325656 CHEMBL3325656 | C33H38N6O3 | 566.706 | 5 / 1 | 3.8 | No |
| 447016 | 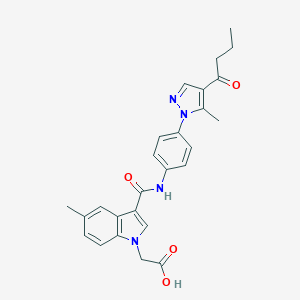 CHEMBL3326907 CHEMBL3326907 | C26H26N4O4 | 458.518 | 5 / 2 | 3.8 | Yes |
| 154132 | 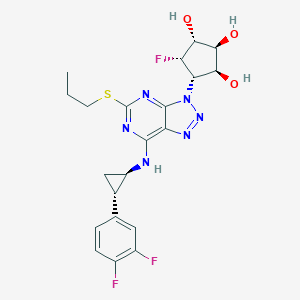 CHEMBL3098241 CHEMBL3098241 | C21H23F3N6O3S | 496.509 | 12 / 4 | 2.2 | No |
| 154530 | 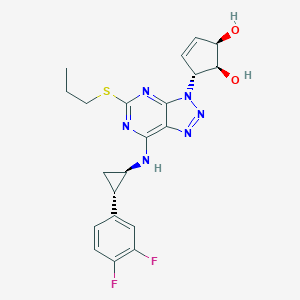 CHEMBL3098246 CHEMBL3098246 | C21H22F2N6O2S | 460.504 | 10 / 3 | 3.0 | Yes |
| 157775 | 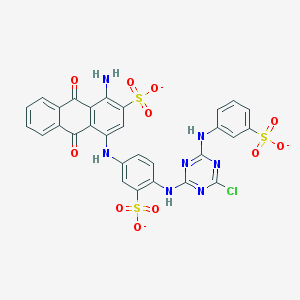 Basilen Blue Basilen Blue | C29H17ClN7O11S3-3 | 771.123 | 18 / 4 | 4.1 | No |
| 157836 | 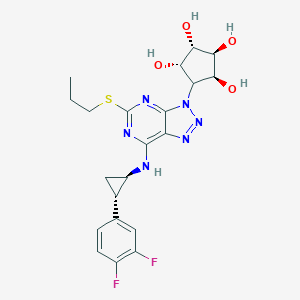 CHEMBL3098240 CHEMBL3098240 | C21H24F2N6O4S | 494.518 | 12 / 5 | 1.2 | No |
| 447958 | 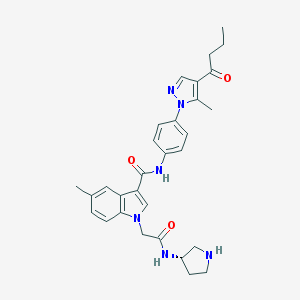 CHEMBL3325632 CHEMBL3325632 | C30H34N6O3 | 526.641 | 5 / 3 | 3.3 | No |
| 448414 | 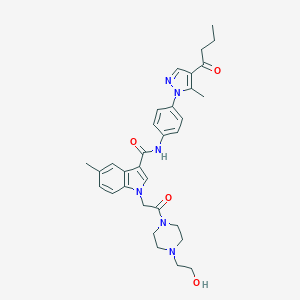 CHEMBL3325659 CHEMBL3325659 | C32H38N6O4 | 570.694 | 6 / 2 | 2.8 | No |
| 448455 | 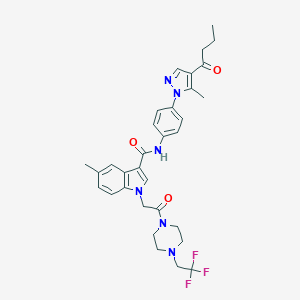 CHEMBL3325784 CHEMBL3325784 | C32H35F3N6O3 | 608.666 | 8 / 1 | 4.6 | No |
| 180672 | 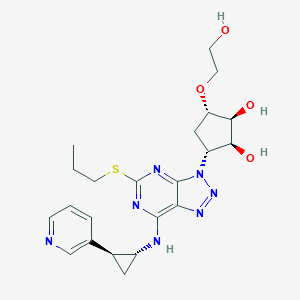 CHEMBL3098236 CHEMBL3098236 | C22H29N7O4S | 487.579 | 11 / 4 | 0.8 | No |
| 448873 | 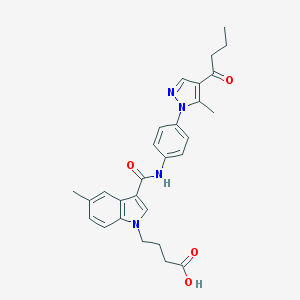 CHEMBL3326905 CHEMBL3326905 | C28H30N4O4 | 486.572 | 5 / 2 | 3.9 | Yes |
| 448899 | 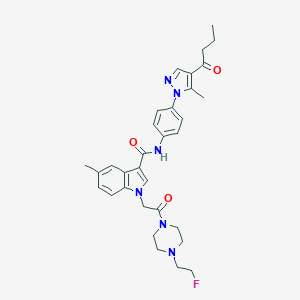 CHEMBL3325782 CHEMBL3325782 | C32H37FN6O3 | 572.685 | 6 / 1 | 3.9 | No |
| 448997 | 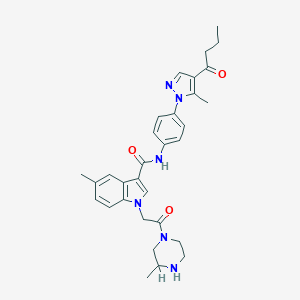 CHEMBL3325665 CHEMBL3325665 | C31H36N6O3 | 540.668 | 5 / 2 | 3.5 | No |
| 449330 | 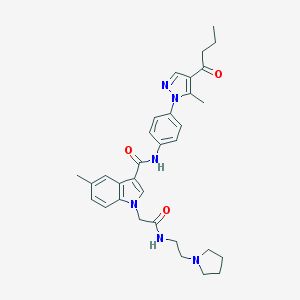 CHEMBL3325635 CHEMBL3325635 | C32H38N6O3 | 554.695 | 5 / 2 | 4.1 | No |
| 449410 | 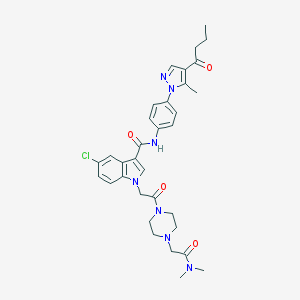 CHEMBL3325803 CHEMBL3325803 | C33H38ClN7O4 | 632.162 | 6 / 1 | 3.3 | No |
| 449583 | 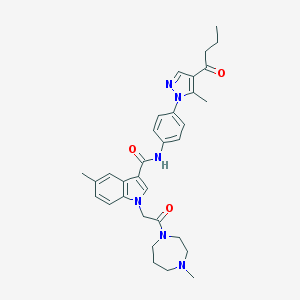 CHEMBL3325643 CHEMBL3325643 | C32H38N6O3 | 554.695 | 5 / 1 | 3.9 | No |
| 449665 | 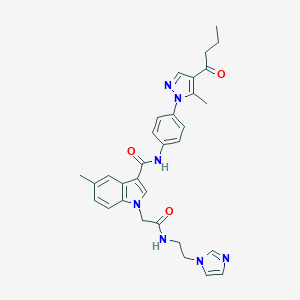 CHEMBL3325637 CHEMBL3325637 | C31H33N7O3 | 551.651 | 5 / 2 | 3.2 | No |
| 449735 | 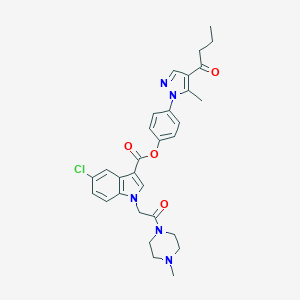 CHEMBL3325895 CHEMBL3325895 | C30H32ClN5O4 | 562.067 | 6 / 0 | 4.4 | No |
| 449773 | 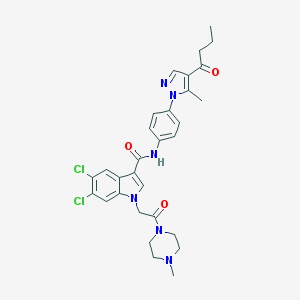 CHEMBL3325797 CHEMBL3325797 | C30H32Cl2N6O3 | 595.525 | 5 / 1 | 4.4 | No |
| 206294 | 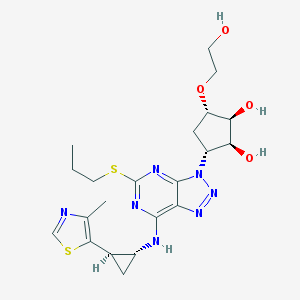 CHEMBL3098235 CHEMBL3098235 | C21H29N7O4S2 | 507.628 | 12 / 4 | 1.3 | No |
| 450021 | 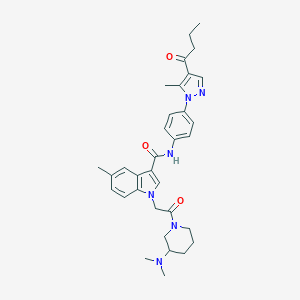 CHEMBL3325650 CHEMBL3325650 | C33H40N6O3 | 568.722 | 5 / 1 | 4.3 | No |
| 220431 | 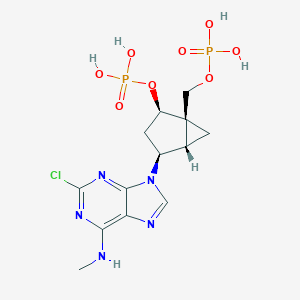 CHEMBL3085531 CHEMBL3085531 | C13H18ClN5O8P2 | 469.712 | 12 / 5 | -1.6 | No |
| 450455 | 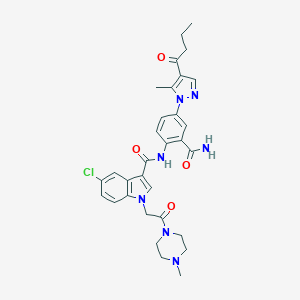 CHEMBL3325898 CHEMBL3325898 | C31H34ClN7O4 | 604.108 | 6 / 2 | 3.2 | No |
| 226528 | 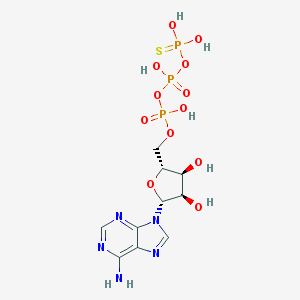 gamma-Thio-ATP gamma-Thio-ATP | C10H16N5O12P3S | 523.242 | 17 / 7 | -4.0 | No |
| 450753 | 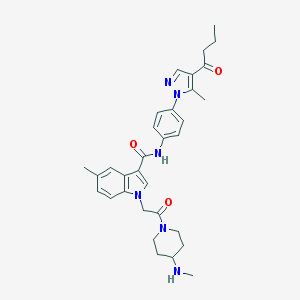 CHEMBL3325645 CHEMBL3325645 | C32H38N6O3 | 554.695 | 5 / 2 | 3.8 | No |
| 450870 | 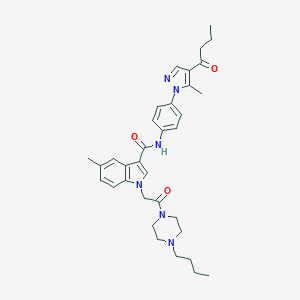 CHEMBL3325658 CHEMBL3325658 | C34H42N6O3 | 582.749 | 5 / 1 | 4.8 | No |
| 239227 | 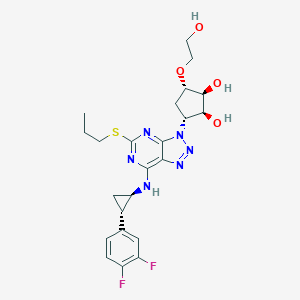 Ticagrelor Ticagrelor | C23H28F2N6O4S | 522.572 | 12 / 4 | 2.0 | No |
| 451109 | 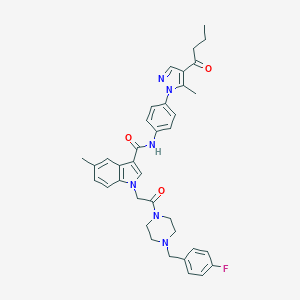 CHEMBL3325788 CHEMBL3325788 | C37H39FN6O3 | 634.756 | 6 / 1 | 5.1 | No |
| 451348 | 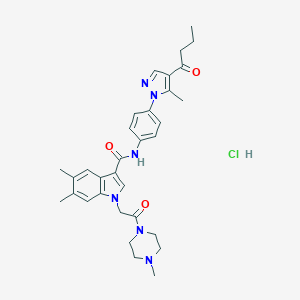 CHEMBL3325630 CHEMBL3325630 | C32H39ClN6O3 | 591.153 | 5 / 2 | N/A | No |
| 451359 | 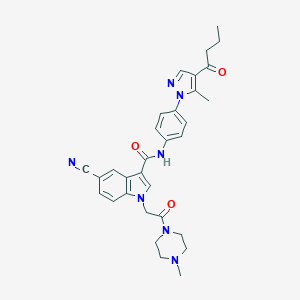 CHEMBL3325794 CHEMBL3325794 | C31H33N7O3 | 551.651 | 6 / 1 | 2.9 | No |
| 451444 | 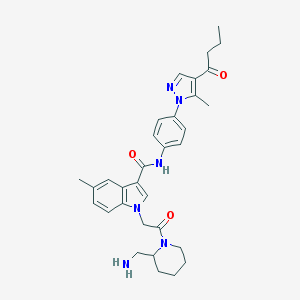 CHEMBL3325651 CHEMBL3325651 | C32H38N6O3 | 554.695 | 5 / 2 | 3.7 | No |
| 451499 | 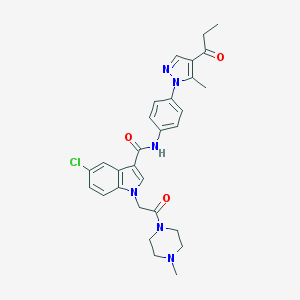 CHEMBL3325806 CHEMBL3325806 | C29H31ClN6O3 | 547.056 | 5 / 1 | 3.4 | No |
| 451539 | 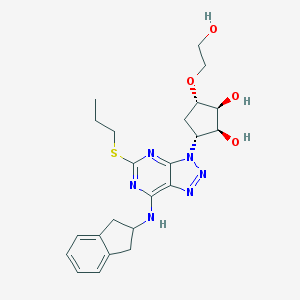 CHEMBL3098237 CHEMBL3098237 | C23H30N6O4S | 486.591 | 10 / 4 | 1.9 | Yes |
| 451611 | 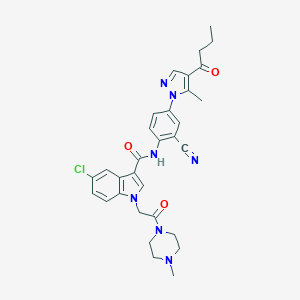 CHEMBL3325896 CHEMBL3325896 | C31H32ClN7O3 | 586.093 | 6 / 1 | 4.1 | No |
| 451617 | 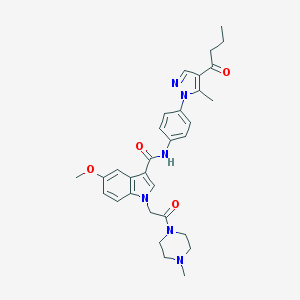 CHEMBL3325789 CHEMBL3325789 | C31H36N6O4 | 556.667 | 6 / 1 | 3.1 | No |
| 451642 | 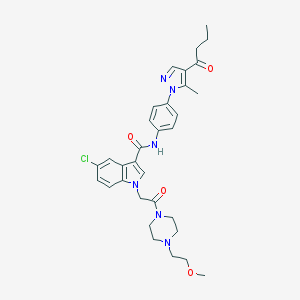 CHEMBL3325801 CHEMBL3325801 | C32H37ClN6O4 | 605.136 | 6 / 1 | 3.6 | No |
| 257000 | 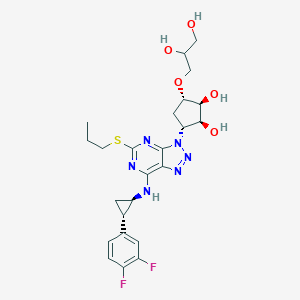 CHEMBL3098249 CHEMBL3098249 | C24H30F2N6O5S | 552.598 | 13 / 5 | 1.4 | No |
| 451912 | 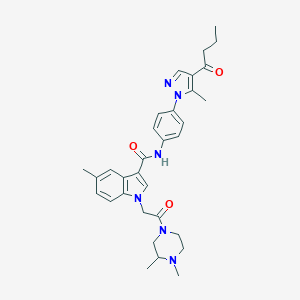 CHEMBL3325666 CHEMBL3325666 | C32H38N6O3 | 554.695 | 5 / 1 | 4.0 | No |
| 565476 | 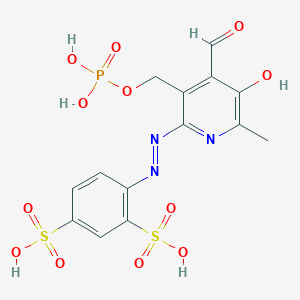 ppads ppads | C14H14N3O12PS2 | 511.369 | 15 / 5 | -1.5 | No |
| 452345 | 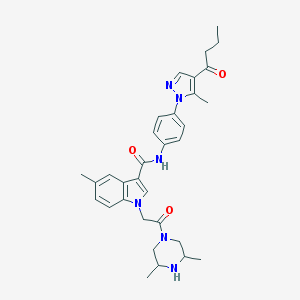 CHEMBL3325779 CHEMBL3325779 | C32H38N6O3 | 554.695 | 5 / 2 | 3.9 | No |
| 452428 | 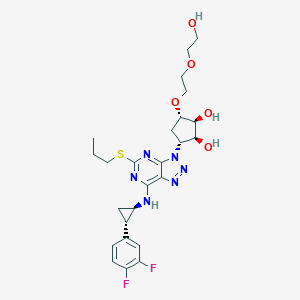 CHEMBL3098248 CHEMBL3098248 | C25H32F2N6O5S | 566.625 | 13 / 4 | 1.9 | No |
| 272373 | 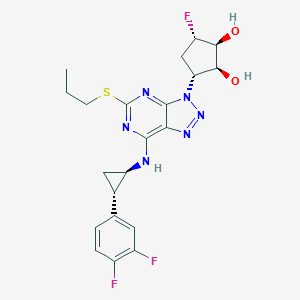 CHEMBL3098247 CHEMBL3098247 | C21H23F3N6O2S | 480.51 | 11 / 3 | 3.2 | No |
| 452497 | 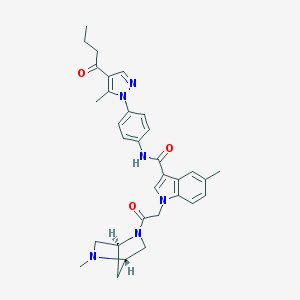 CHEMBL3325787 CHEMBL3325787 | C32H36N6O3 | 552.679 | 5 / 1 | 3.8 | No |
| 452640 | 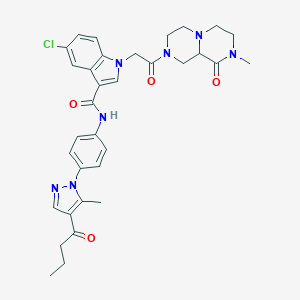 CHEMBL3325805 CHEMBL3325805 | C33H36ClN7O4 | 630.146 | 6 / 1 | 3.1 | No |
| 452711 | 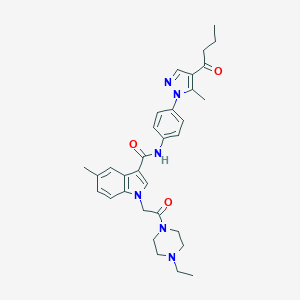 CHEMBL3325657 CHEMBL3325657 | C32H38N6O3 | 554.695 | 5 / 1 | 3.9 | No |
| 453761 | 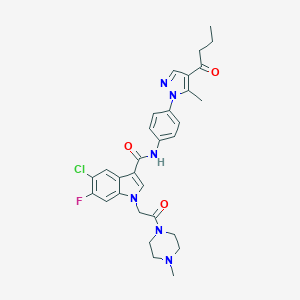 CHEMBL3325790 CHEMBL3325790 | C30H32ClFN6O3 | 579.073 | 6 / 1 | 3.9 | No |
| 453947 | 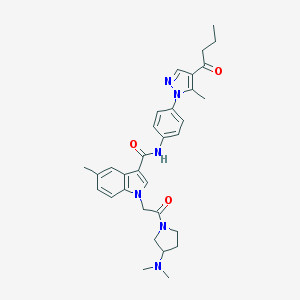 CHEMBL3325647 CHEMBL3325647 | C32H38N6O3 | 554.695 | 5 / 1 | 3.9 | No |
| 453950 | 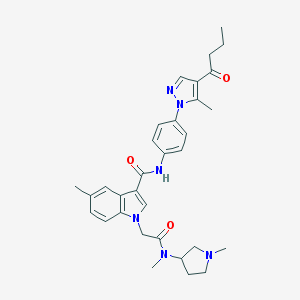 CHEMBL3325640 CHEMBL3325640 | C32H38N6O3 | 554.695 | 5 / 1 | 3.9 | No |
| 454013 | 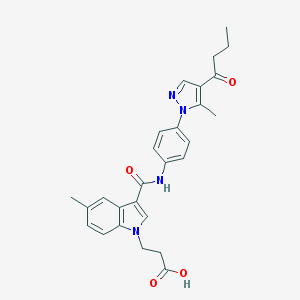 CHEMBL3326906 CHEMBL3326906 | C27H28N4O4 | 472.545 | 5 / 2 | 3.5 | Yes |
| 454166 | 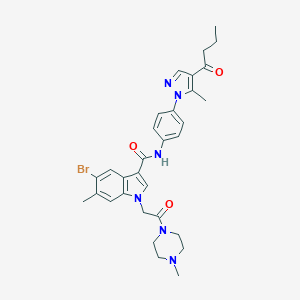 CHEMBL3325791 CHEMBL3325791 | C31H35BrN6O3 | 619.564 | 5 / 1 | 4.2 | No |
| 455024 | 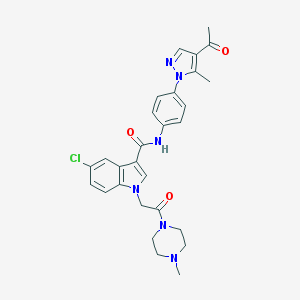 CHEMBL3325807 CHEMBL3325807 | C28H29ClN6O3 | 533.029 | 5 / 1 | 3.0 | No |
| 455030 | 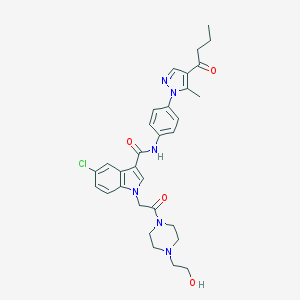 CHEMBL3325802 CHEMBL3325802 | C31H35ClN6O4 | 591.109 | 6 / 2 | 3.1 | No |
| 336022 | 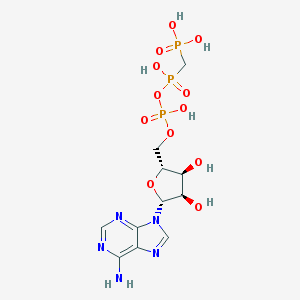 AMP-PCP AMP-PCP | C11H18N5O12P3 | 505.209 | 16 / 7 | -5.9 | No |
| 455071 | 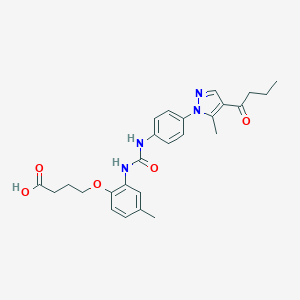 CHEMBL3326903 CHEMBL3326903 | C26H30N4O5 | 478.549 | 6 / 3 | 3.6 | Yes |
| 455209 | 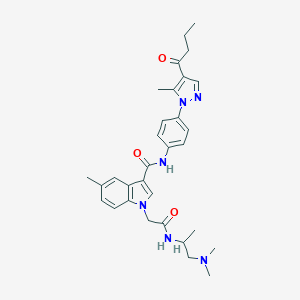 CHEMBL3325638 CHEMBL3325638 | C31H38N6O3 | 542.684 | 5 / 2 | 4.0 | No |
| 455470 | 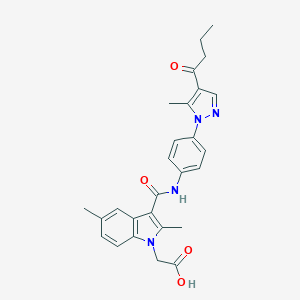 CHEMBL3325625 CHEMBL3325625 | C27H28N4O4 | 472.545 | 5 / 2 | 4.2 | Yes |
| 455760 | 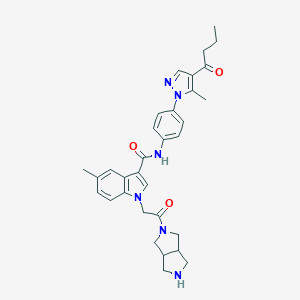 CHEMBL3325655 CHEMBL3325655 | C32H36N6O3 | 552.679 | 5 / 2 | 3.3 | No |
| 455765 | 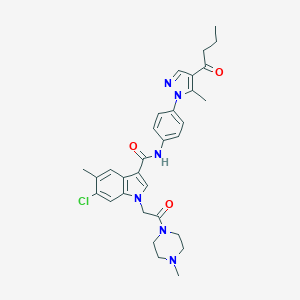 CHEMBL3325796 CHEMBL3325796 | C31H35ClN6O3 | 575.11 | 5 / 1 | 4.1 | No |
| 455810 | 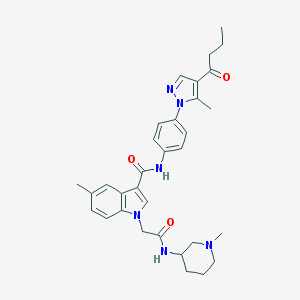 CHEMBL3325641 CHEMBL3325641 | C32H38N6O3 | 554.695 | 5 / 2 | 4.1 | No |
| 455824 | 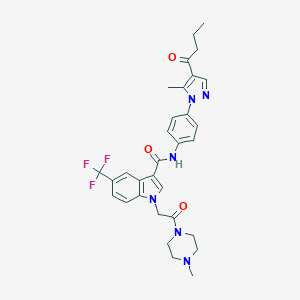 CHEMBL3325792 CHEMBL3325792 | C31H33F3N6O3 | 594.639 | 8 / 1 | 4.0 | No |
| 455985 | 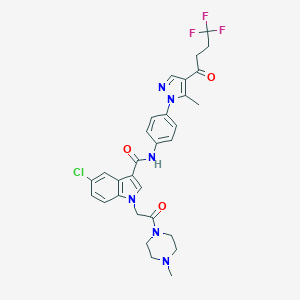 CHEMBL3325808 CHEMBL3325808 | C30H30ClF3N6O3 | 615.054 | 8 / 1 | 4.4 | No |
| 363335 | 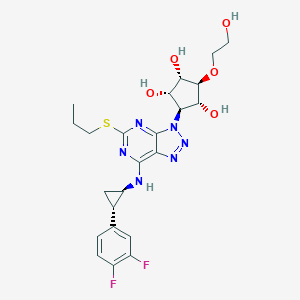 CHEMBL3098239 CHEMBL3098239 | C23H28F2N6O5S | 538.571 | 13 / 5 | 1.1 | No |
| 456130 | 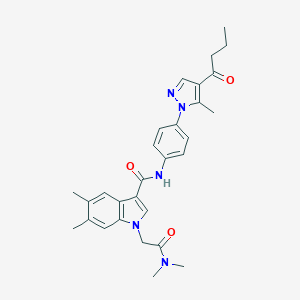 CHEMBL3325629 CHEMBL3325629 | C29H33N5O3 | 499.615 | 4 / 1 | 4.1 | Yes |
| 368121 | 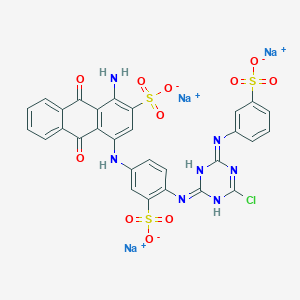 CID 136908877 CID 136908877 | C29H17ClN7Na3O11S3 | 840.092 | 15 / 4 | N/A | No |
| 456453 | 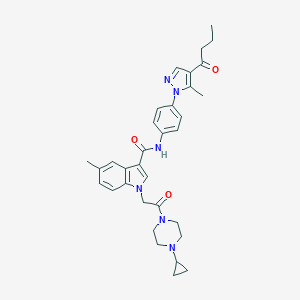 CHEMBL3325662 CHEMBL3325662 | C33H38N6O3 | 566.706 | 5 / 1 | 4.1 | No |
| 456490 | 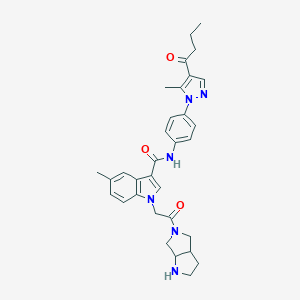 CHEMBL3325653 CHEMBL3325653 | C32H36N6O3 | 552.679 | 5 / 2 | 3.5 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218