

You can:
| Name | Lysophosphatidic acid receptor 5 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | LPAR5 |
| Synonym | G-protein coupled receptor 92 LPAR5 LPA5 receptor LPA-5 LPA receptor 5 [ Show all ] |
| Disease | N/A |
| Length | 372 |
| Amino acid sequence | MLANSSSTNSSVLPCPDYRPTHRLHLVVYSLVLAAGLPLNALALWVFLRALRVHSVVSVYMCNLAASDLLFTLSLPVRLSYYALHHWPFPDLLCQTTGAIFQMNMYGSCIFLMLINVDRYAAIVHPLRLRHLRRPRVARLLCLGVWALILVFAVPAARVHRPSRCRYRDLEVRLCFESFSDELWKGRLLPLVLLAEALGFLLPLAAVVYSSGRVFWTLARPDATQSQRRRKTVRLLLANLVIFLLCFVPYNSTLAVYGLLRSKLVAASVPARDRVRGVLMVMVLLAGANCVLDPLVYYFSAEGFRNTLRGLGTPHRARTSATNGTRAALAQSERSAVTTDATRPDAASQGLLRPSDSHSLSSFTQCPQDSAL |
| UniProt | Q9H1C0 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | Q9H1C0 |
| 3D structure model | This predicted structure model is from GPCR-EXP Q9H1C0. |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL5700 |
| IUPHAR | 124 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 535973 | 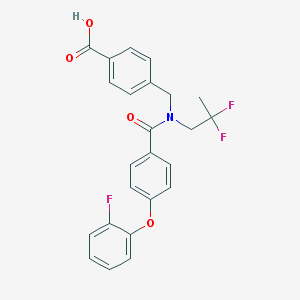 CHEMBL3943469 CHEMBL3943469 | C24H20F3NO4 | 443.422 | 7 / 1 | 5.1 | No |
| 4542 | 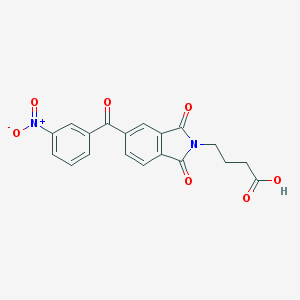 AC1LP5VO AC1LP5VO | C19H14N2O7 | 382.328 | 7 / 1 | 1.9 | Yes |
| 536052 | 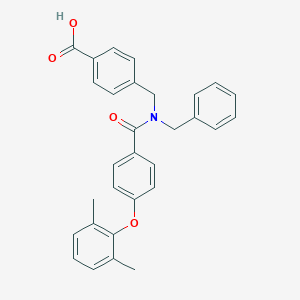 CHEMBL3920547 CHEMBL3920547 | C30H27NO4 | 465.549 | 4 / 1 | 6.2 | No |
| 536167 | 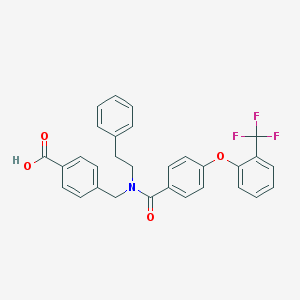 CHEMBL3935519 CHEMBL3935519 | C30H24F3NO4 | 519.52 | 7 / 1 | 6.8 | No |
| 553283 | 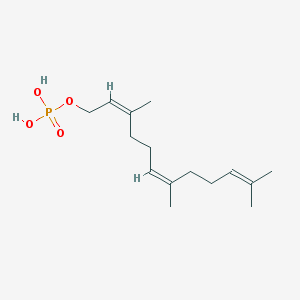 Farnesyl monophosphate Farnesyl monophosphate | C15H27O4P | 302.351 | 4 / 2 | 3.7 | Yes |
| 536194 | 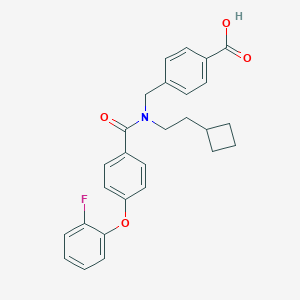 CHEMBL3960586 CHEMBL3960586 | C27H26FNO4 | 447.506 | 5 / 1 | 6.0 | No |
| 14861 | 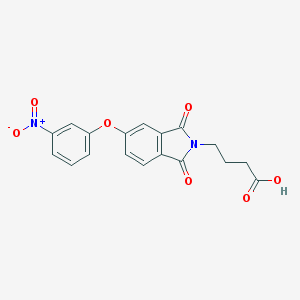 4-[5-(3-nitrophenoxy)-1,3-dioxo-1,3-dihydro-2H-isoindol-2-yl]butanoic acid 4-[5-(3-nitrophenoxy)-1,3-dioxo-1,3-dihydro-2H-isoindol-2-yl]butanoic acid | C18H14N2O7 | 370.317 | 7 / 1 | 2.1 | Yes |
| 536422 | 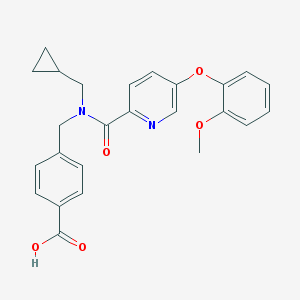 CHEMBL3985179 CHEMBL3985179 | C25H24N2O5 | 432.476 | 6 / 1 | 3.9 | Yes |
| 17533 | 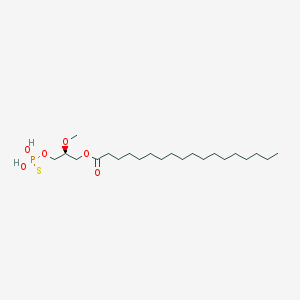 CHEMBL357053 CHEMBL357053 | C22H45O6PS | 468.63 | 7 / 2 | 8.4 | No |
| 17540 | 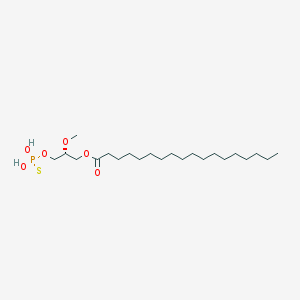 CHEMBL153043 CHEMBL153043 | C22H45O6PS | 468.63 | 7 / 2 | 8.4 | No |
| 17547 | 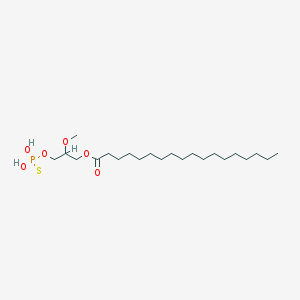 CHEMBL2017139 CHEMBL2017139 | C22H45O6PS | 468.63 | 7 / 2 | 8.4 | No |
| 536534 | 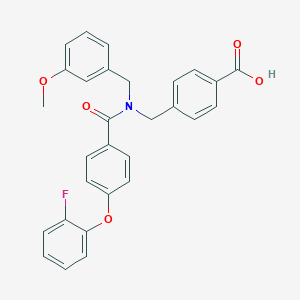 CHEMBL3900559 CHEMBL3900559 | C29H24FNO5 | 485.511 | 6 / 1 | 5.5 | No |
| 536613 | 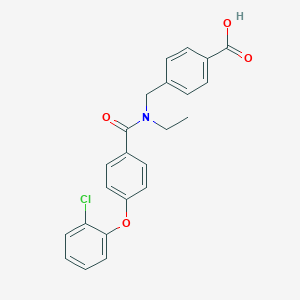 CHEMBL3942927 CHEMBL3942927 | C23H20ClNO4 | 409.866 | 4 / 1 | 4.9 | Yes |
| 536640 | 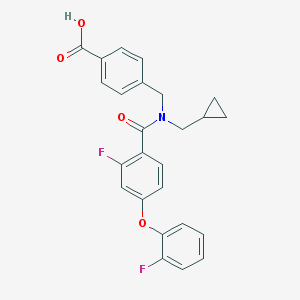 CHEMBL3900867 CHEMBL3900867 | C25H21F2NO4 | 437.443 | 6 / 1 | 4.9 | Yes |
| 536667 | 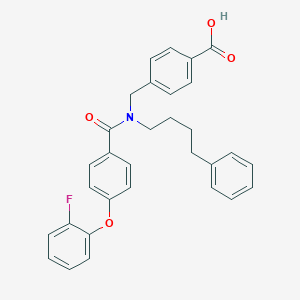 CHEMBL3956413 CHEMBL3956413 | C31H28FNO4 | 497.566 | 5 / 1 | 6.7 | No |
| 27910 | 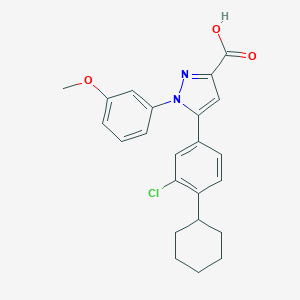 TC LPA5 4 TC LPA5 4 | C23H23ClN2O3 | 410.898 | 4 / 1 | 6.5 | No |
| 29832 | 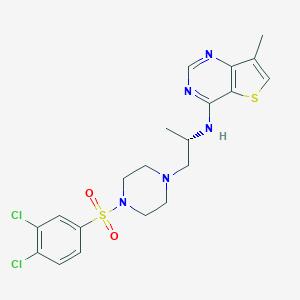 LPA2 antagonist 1 LPA2 antagonist 1 | C20H23Cl2N5O2S2 | 500.457 | 8 / 1 | 4.3 | No |
| 537151 | 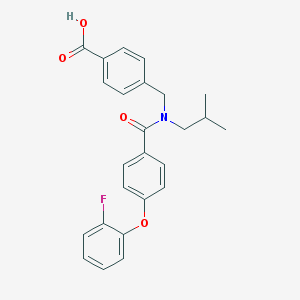 CHEMBL3948456 CHEMBL3948456 | C25H24FNO4 | 421.468 | 5 / 1 | 5.4 | No |
| 537379 | 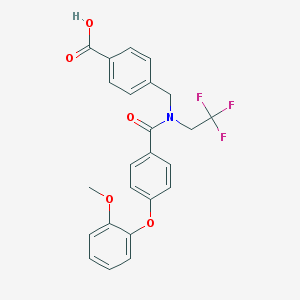 CHEMBL3901720 CHEMBL3901720 | C24H20F3NO5 | 459.421 | 8 / 1 | 5.0 | Yes |
| 537747 | 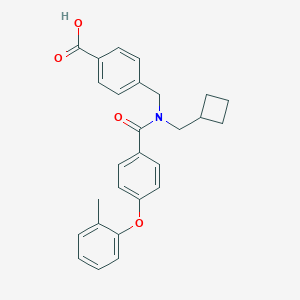 CHEMBL3961807 CHEMBL3961807 | C27H27NO4 | 429.516 | 4 / 1 | 5.6 | No |
| 537837 | 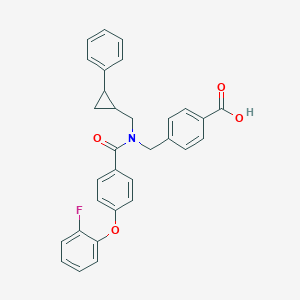 CHEMBL3980736 CHEMBL3980736 | C31H26FNO4 | 495.55 | 5 / 1 | 6.2 | No |
| 537956 | 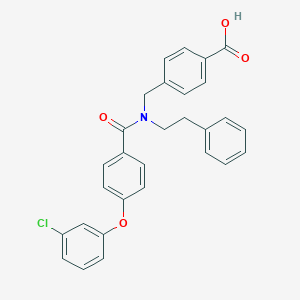 CHEMBL3957378 CHEMBL3957378 | C29H24ClNO4 | 485.964 | 4 / 1 | 6.5 | No |
| 538059 | 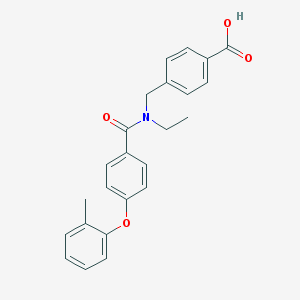 CHEMBL3908939 CHEMBL3908939 | C24H23NO4 | 389.451 | 4 / 1 | 4.7 | Yes |
| 538074 | 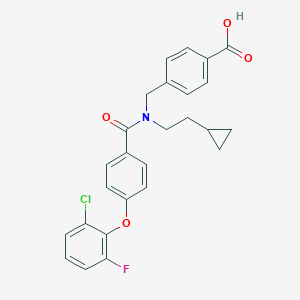 CHEMBL3903990 CHEMBL3903990 | C26H23ClFNO4 | 467.921 | 5 / 1 | 6.1 | No |
| 538115 | 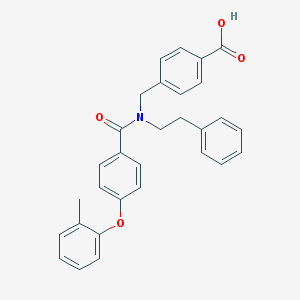 CHEMBL3971670 CHEMBL3971670 | C30H27NO4 | 465.549 | 4 / 1 | 6.3 | No |
| 92819 | 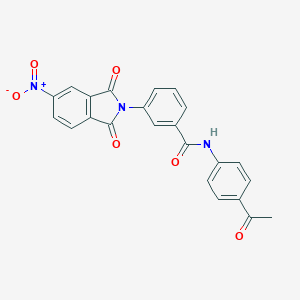 CHEMBL606354 CHEMBL606354 | C23H15N3O6 | 429.388 | 6 / 1 | 2.8 | Yes |
| 538319 | 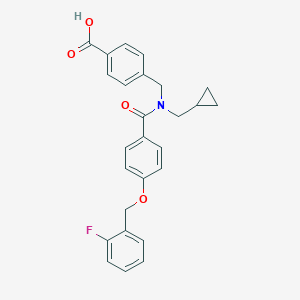 CHEMBL3933338 CHEMBL3933338 | C26H24FNO4 | 433.479 | 5 / 1 | 4.7 | Yes |
| 538361 | 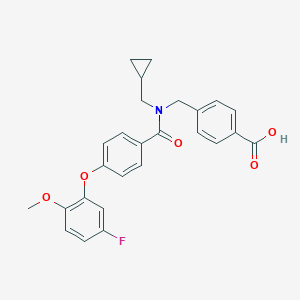 CHEMBL3921376 CHEMBL3921376 | C26H24FNO5 | 449.478 | 6 / 1 | 4.7 | Yes |
| 538580 | 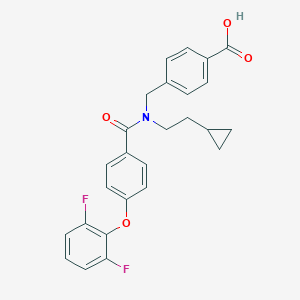 CHEMBL3893992 CHEMBL3893992 | C26H23F2NO4 | 451.47 | 6 / 1 | 5.6 | No |
| 538600 | 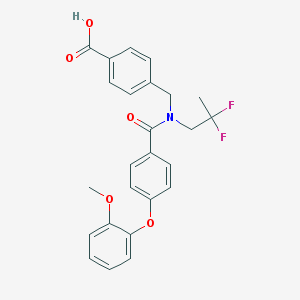 CHEMBL3892760 CHEMBL3892760 | C25H23F2NO5 | 455.458 | 7 / 1 | 4.9 | Yes |
| 538730 | 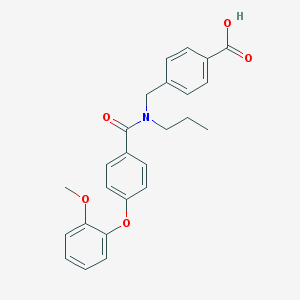 CHEMBL3961765 CHEMBL3961765 | C25H25NO5 | 419.477 | 5 / 1 | 4.8 | Yes |
| 113881 | 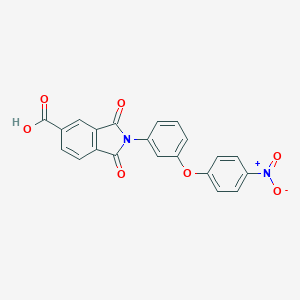 CHEMBL604677 CHEMBL604677 | C21H12N2O7 | 404.334 | 7 / 1 | 3.3 | Yes |
| 538929 | 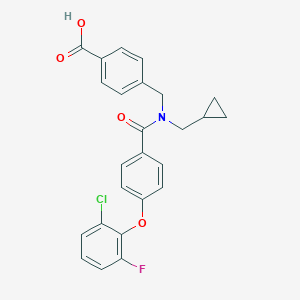 CHEMBL3971635 CHEMBL3971635 | C25H21ClFNO4 | 453.894 | 5 / 1 | 5.4 | No |
| 561205 | 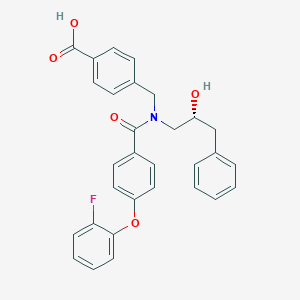 SCHEMBL16506802 SCHEMBL16506802 | C30H26FNO5 | 499.538 | 6 / 2 | 5.4 | No |
| 539128 | 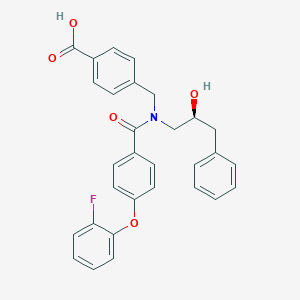 CHEMBL3955284 CHEMBL3955284 | C30H26FNO5 | 499.538 | 6 / 2 | 5.4 | No |
| 539159 | 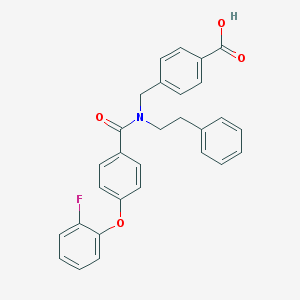 CHEMBL3939578 CHEMBL3939578 | C29H24FNO4 | 469.512 | 5 / 1 | 6.0 | No |
| 539300 | 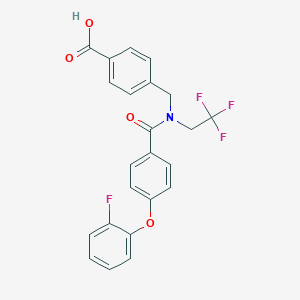 CHEMBL3959098 CHEMBL3959098 | C23H17F4NO4 | 447.386 | 8 / 1 | 5.1 | No |
| 132753 | 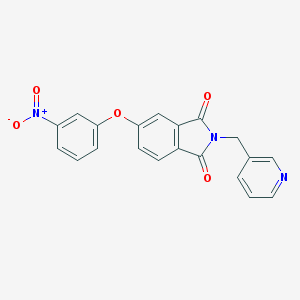 AC1LRAEE AC1LRAEE | C20H13N3O5 | 375.34 | 6 / 0 | 2.7 | Yes |
| 539338 | 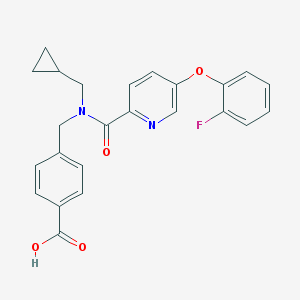 CHEMBL3972525 CHEMBL3972525 | C24H21FN2O4 | 420.44 | 6 / 1 | 4.0 | Yes |
| 539564 | 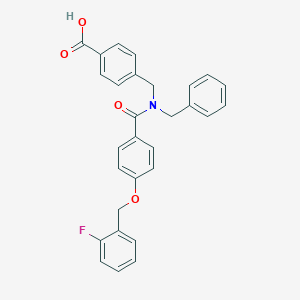 CHEMBL3920905 CHEMBL3920905 | C29H24FNO4 | 469.512 | 5 / 1 | 5.5 | No |
| 539730 | 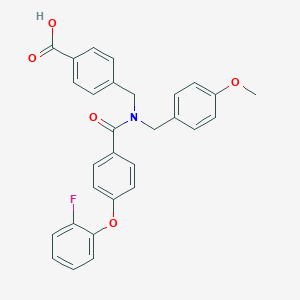 CHEMBL3911605 CHEMBL3911605 | C29H24FNO5 | 485.511 | 6 / 1 | 5.5 | No |
| 540123 | 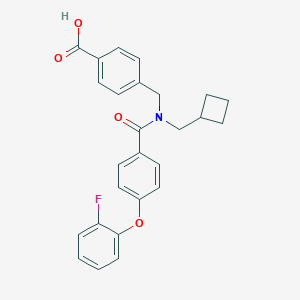 CHEMBL3976680 CHEMBL3976680 | C26H24FNO4 | 433.479 | 5 / 1 | 5.3 | No |
| 162857 | 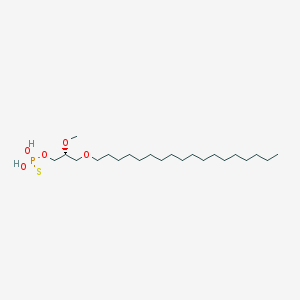 CHEMBL2335048 CHEMBL2335048 | C22H47O5PS | 454.647 | 6 / 2 | 8.8 | No |
| 162864 | 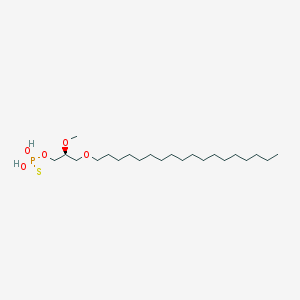 CHEMBL2335047 CHEMBL2335047 | C22H47O5PS | 454.647 | 6 / 2 | 8.8 | No |
| 162869 | 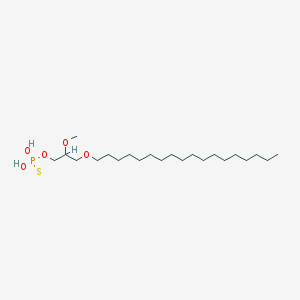 CHEMBL2335052 CHEMBL2335052 | C22H47O5PS | 454.647 | 6 / 2 | 8.8 | No |
| 163621 | 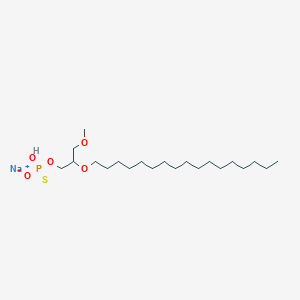 CHEMBL2335050 CHEMBL2335050 | C21H44NaO5PS | 462.602 | 6 / 1 | N/A | N/A |
| 540210 | 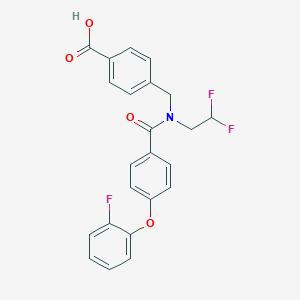 CHEMBL3963121 CHEMBL3963121 | C23H18F3NO4 | 429.395 | 7 / 1 | 4.9 | Yes |
| 540223 | 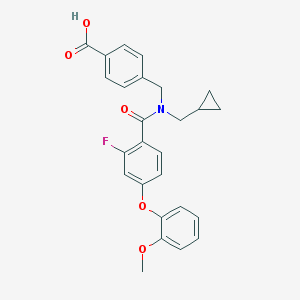 CHEMBL3910049 CHEMBL3910049 | C26H24FNO5 | 449.478 | 6 / 1 | 4.7 | Yes |
| 178096 | 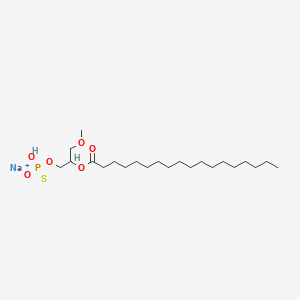 CHEMBL2335051 CHEMBL2335051 | C22H44NaO6PS | 490.612 | 7 / 1 | N/A | N/A |
| 540740 | 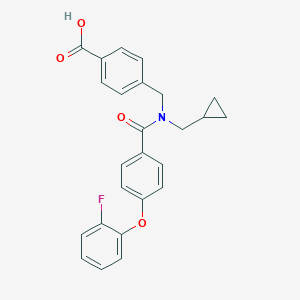 CHEMBL3986752 CHEMBL3986752 | C25H22FNO4 | 419.452 | 5 / 1 | 4.8 | Yes |
| 540921 | 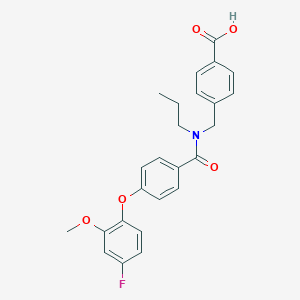 CHEMBL3985727 CHEMBL3985727 | C25H24FNO5 | 437.467 | 6 / 1 | 4.9 | Yes |
| 540925 | 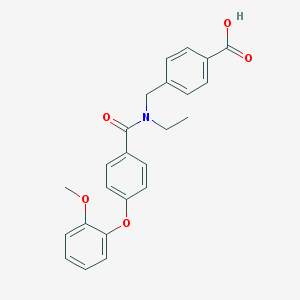 CHEMBL3923068 CHEMBL3923068 | C24H23NO5 | 405.45 | 5 / 1 | 4.3 | Yes |
| 190299 | 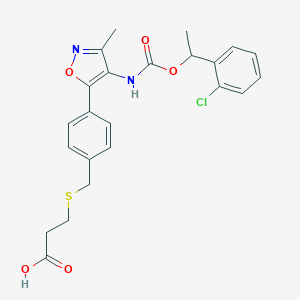 Ki16425 Ki16425 | C23H23ClN2O5S | 474.956 | 7 / 2 | 4.5 | Yes |
| 486635 | 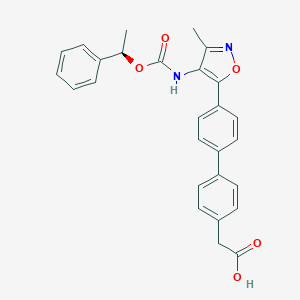 AM095 free acid AM095 free acid | C27H24N2O5 | 456.498 | 6 / 2 | 5.0 | Yes |
| 541282 | 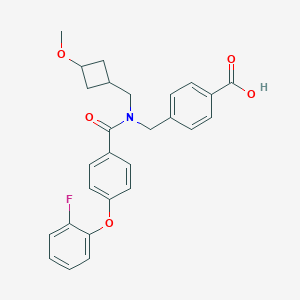 CHEMBL3891142 CHEMBL3891142 | C27H26FNO5 | 463.505 | 6 / 1 | 4.7 | Yes |
| 541364 | 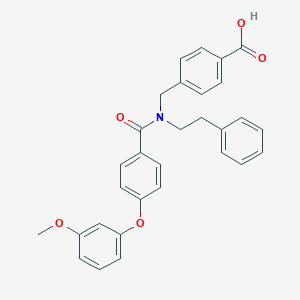 CHEMBL3934403 CHEMBL3934403 | C30H27NO5 | 481.548 | 5 / 1 | 5.9 | No |
| 541617 | 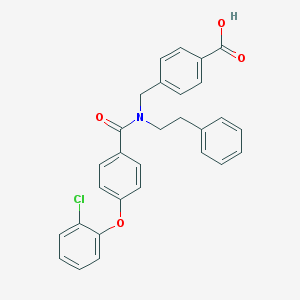 CHEMBL3979729 CHEMBL3979729 | C29H24ClNO4 | 485.964 | 4 / 1 | 6.5 | No |
| 541873 | 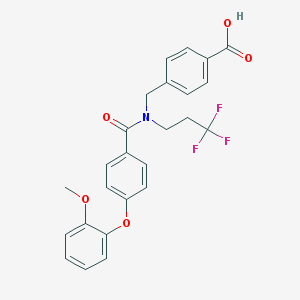 CHEMBL3987061 CHEMBL3987061 | C25H22F3NO5 | 473.448 | 8 / 1 | 5.4 | No |
| 542000 | 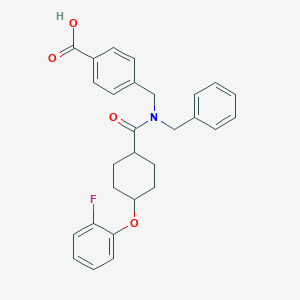 CHEMBL3947618 CHEMBL3947618 | C28H28FNO4 | 461.533 | 5 / 1 | 5.2 | No |
| 542082 | 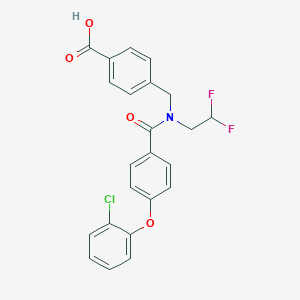 CHEMBL3948271 CHEMBL3948271 | C23H18ClF2NO4 | 445.847 | 6 / 1 | 5.4 | No |
| 542127 | 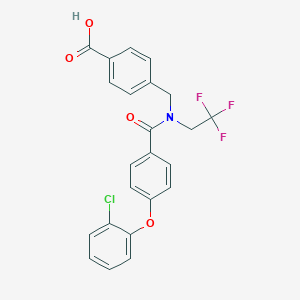 CHEMBL3933337 CHEMBL3933337 | C23H17ClF3NO4 | 463.837 | 7 / 1 | 5.7 | No |
| 233026 | 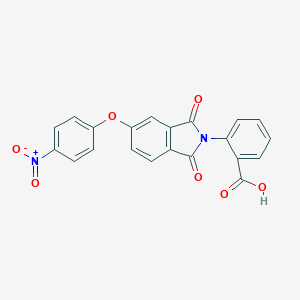 2-[5-(4-nitrophenoxy)-1,3-dioxo-1,3-dihydro-2H-isoindol-2-yl]benzoic acid 2-[5-(4-nitrophenoxy)-1,3-dioxo-1,3-dihydro-2H-isoindol-2-yl]benzoic acid | C21H12N2O7 | 404.334 | 7 / 1 | 3.3 | Yes |
| 542496 | 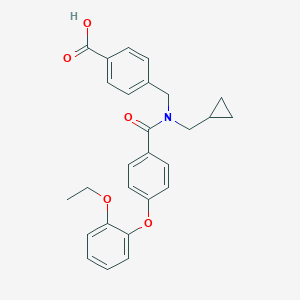 CHEMBL3927774 CHEMBL3927774 | C27H27NO5 | 445.515 | 5 / 1 | 5.0 | Yes |
| 542499 | 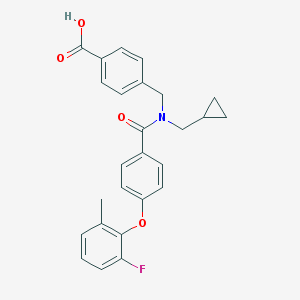 CHEMBL3966386 CHEMBL3966386 | C26H24FNO4 | 433.479 | 5 / 1 | 5.1 | No |
| 542600 | 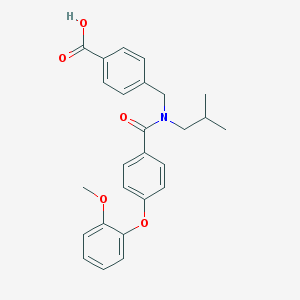 CHEMBL3942192 CHEMBL3942192 | C26H27NO5 | 433.504 | 5 / 1 | 5.2 | No |
| 542640 | 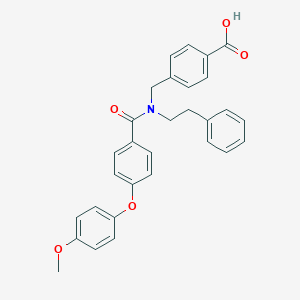 CHEMBL3893864 CHEMBL3893864 | C30H27NO5 | 481.548 | 5 / 1 | 5.9 | No |
| 542717 | 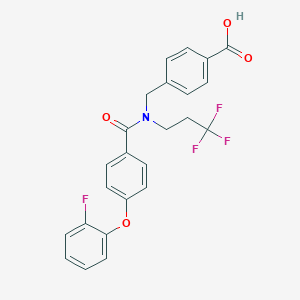 CHEMBL3896384 CHEMBL3896384 | C24H19F4NO4 | 461.413 | 8 / 1 | 5.5 | No |
| 543280 | 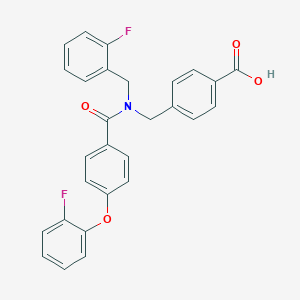 CHEMBL3910281 CHEMBL3910281 | C28H21F2NO4 | 473.476 | 6 / 1 | 5.6 | No |
| 278359 | 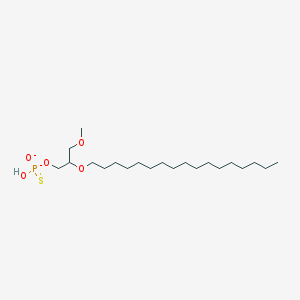 CHEMBL2335050 CHEMBL2335050 | C21H44O5PS- | 439.612 | 6 / 1 | 8.2 | No |
| 543536 | 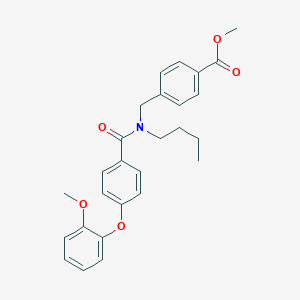 CHEMBL3920849 CHEMBL3920849 | C27H29NO5 | 447.531 | 5 / 0 | 5.5 | No |
| 543708 | 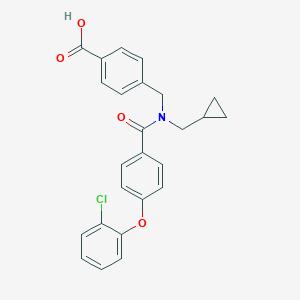 CHEMBL3951978 CHEMBL3951978 | C25H22ClNO4 | 435.904 | 4 / 1 | 5.3 | No |
| 543744 | 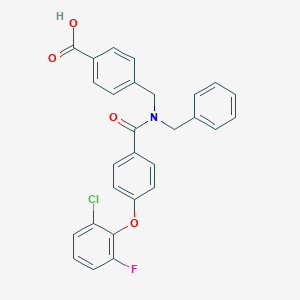 CHEMBL3914963 CHEMBL3914963 | C28H21ClFNO4 | 489.927 | 5 / 1 | 6.2 | No |
| 290483 | 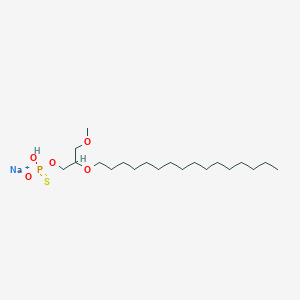 CHEMBL2335049 CHEMBL2335049 | C20H42NaO5PS | 448.575 | 6 / 1 | N/A | N/A |
| 544318 | 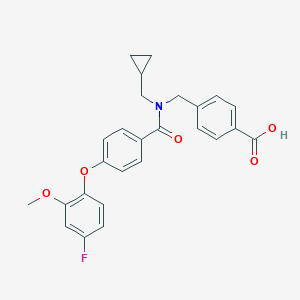 CHEMBL3918970 CHEMBL3918970 | C26H24FNO5 | 449.478 | 6 / 1 | 4.7 | Yes |
| 544356 | 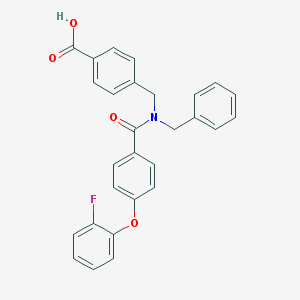 CHEMBL3891550 CHEMBL3891550 | C28H22FNO4 | 455.485 | 5 / 1 | 5.5 | No |
| 310816 | 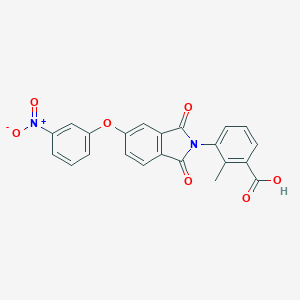 AC1LR8M9 AC1LR8M9 | C22H14N2O7 | 418.361 | 7 / 1 | 3.7 | Yes |
| 544433 | 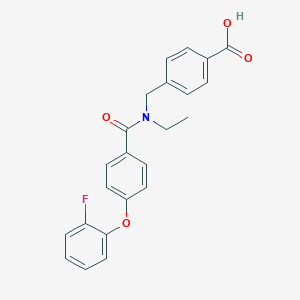 CHEMBL3973570 CHEMBL3973570 | C23H20FNO4 | 393.414 | 5 / 1 | 4.4 | Yes |
| 544449 | 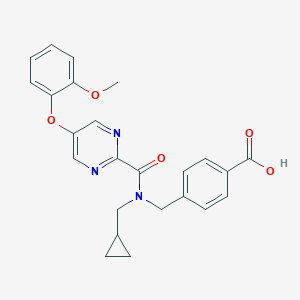 CHEMBL3897650 CHEMBL3897650 | C24H23N3O5 | 433.464 | 7 / 1 | 3.3 | Yes |
| 544591 | 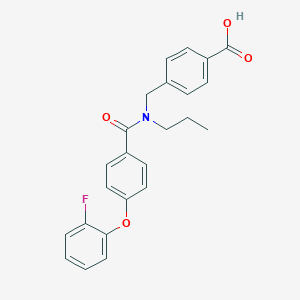 CHEMBL3975980 CHEMBL3975980 | C24H22FNO4 | 407.441 | 5 / 1 | 4.9 | Yes |
| 544628 | 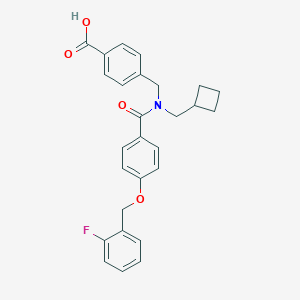 CHEMBL3948718 CHEMBL3948718 | C27H26FNO4 | 447.506 | 5 / 1 | 5.3 | No |
| 544644 | 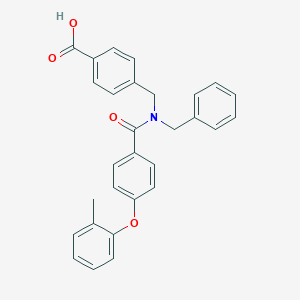 CHEMBL3942894 CHEMBL3942894 | C29H25NO4 | 451.522 | 4 / 1 | 5.8 | No |
| 544761 | 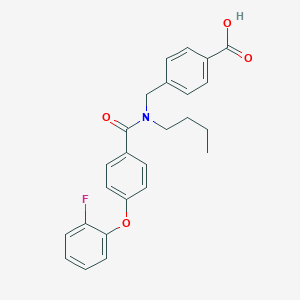 CHEMBL3915509 CHEMBL3915509 | C25H24FNO4 | 421.468 | 5 / 1 | 5.3 | No |
| 544767 | 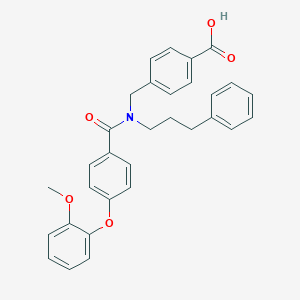 CHEMBL3936755 CHEMBL3936755 | C31H29NO5 | 495.575 | 5 / 1 | 6.2 | No |
| 544897 | 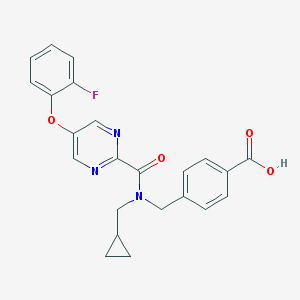 CHEMBL3907299 CHEMBL3907299 | C23H20FN3O4 | 421.428 | 7 / 1 | 3.4 | Yes |
| 544964 | 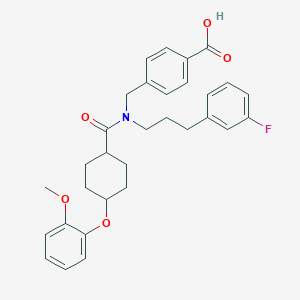 CHEMBL3908908 CHEMBL3908908 | C31H34FNO5 | 519.613 | 6 / 1 | 6.0 | No |
| 544991 | 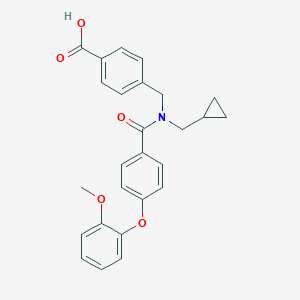 CHEMBL3945517 CHEMBL3945517 | C26H25NO5 | 431.488 | 5 / 1 | 4.6 | Yes |
| 545090 | 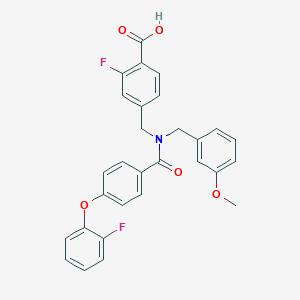 CHEMBL3920145 CHEMBL3920145 | C29H23F2NO5 | 503.502 | 7 / 1 | 5.6 | No |
| 545245 | 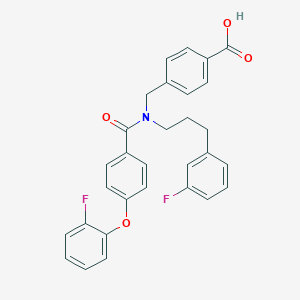 CHEMBL3943133 CHEMBL3943133 | C30H25F2NO4 | 501.53 | 6 / 1 | 6.4 | No |
| 545266 | 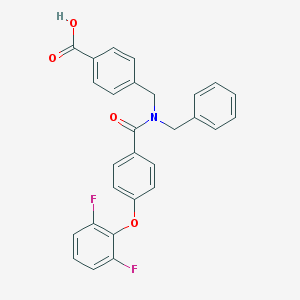 CHEMBL3918468 CHEMBL3918468 | C28H21F2NO4 | 473.476 | 6 / 1 | 5.6 | No |
| 356924 | 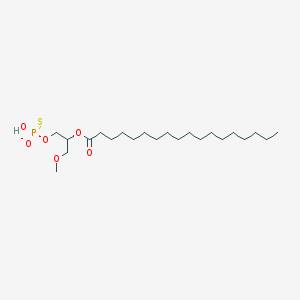 CHEMBL2335051 CHEMBL2335051 | C22H44O6PS- | 467.622 | 7 / 1 | 8.3 | No |
| 555124 | 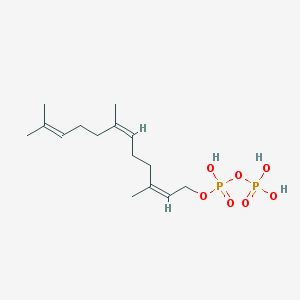 2-cis,6-cis-farnesyl diphosphate 2-cis,6-cis-farnesyl diphosphate | C15H28O7P2 | 382.33 | 7 / 3 | 2.6 | Yes |
| 545913 | 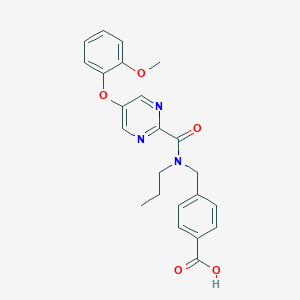 CHEMBL3935056 CHEMBL3935056 | C23H23N3O5 | 421.453 | 7 / 1 | 3.4 | Yes |
| 546222 | 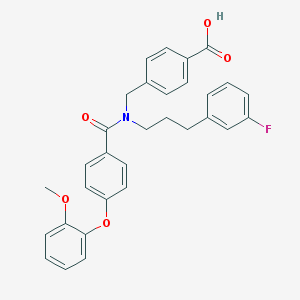 CHEMBL3966526 CHEMBL3966526 | C31H28FNO5 | 513.565 | 6 / 1 | 6.3 | No |
| 380358 | 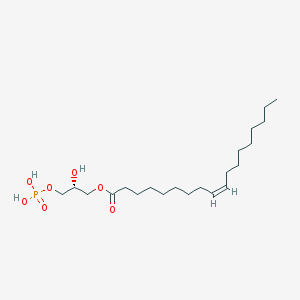 1-Oleoyl Lysophosphatidic Acid 1-Oleoyl Lysophosphatidic Acid | C21H41O7P | 436.526 | 7 / 3 | 5.2 | No |
| 555175 | 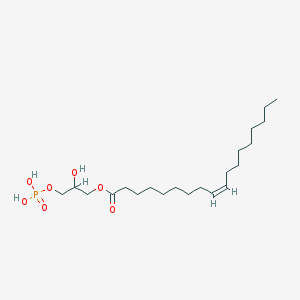 lysophosphatidic acid lysophosphatidic acid | C21H41O7P | 436.526 | 7 / 3 | 5.2 | No |
| 546272 | 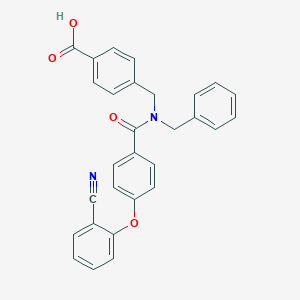 CHEMBL3959515 CHEMBL3959515 | C29H22N2O4 | 462.505 | 5 / 1 | 5.1 | No |
| 385037 | 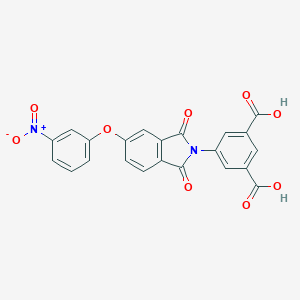 STK010162 STK010162 | C22H12N2O9 | 448.343 | 9 / 2 | 2.9 | Yes |
| 546504 | 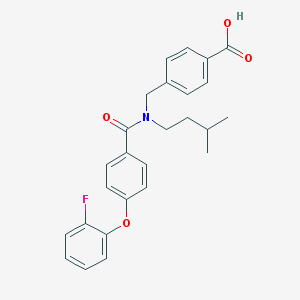 CHEMBL3890476 CHEMBL3890476 | C26H26FNO4 | 435.495 | 5 / 1 | 5.7 | No |
| 395587 | 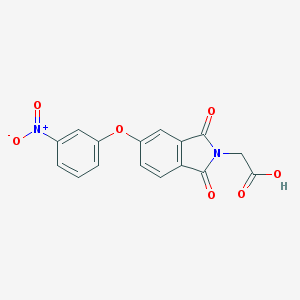 CHEMBL610179 CHEMBL610179 | C16H10N2O7 | 342.263 | 7 / 1 | 1.9 | Yes |
| 546769 | 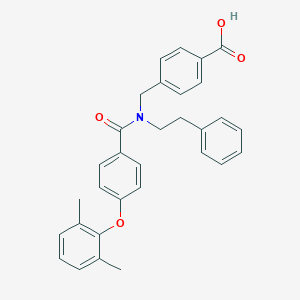 CHEMBL3911962 CHEMBL3911962 | C31H29NO4 | 479.576 | 4 / 1 | 6.6 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218