

You can:
| Name | P2Y purinoceptor 12 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | P2RY12 |
| Synonym | P2Y12 platelet ADP receptor P2Y12 receptor P2YADP purinergic receptor P2Y P2Y12 [ Show all ] |
| Disease | N/A |
| Length | 342 |
| Amino acid sequence | MQAVDNLTSAPGNTSLCTRDYKITQVLFPLLYTVLFFVGLITNGLAMRIFFQIRSKSNFIIFLKNTVISDLLMILTFPFKILSDAKLGTGPLRTFVCQVTSVIFYFTMYISISFLGLITIDRYQKTTRPFKTSNPKNLLGAKILSVVIWAFMFLLSLPNMILTNRQPRDKNVKKCSFLKSEFGLVWHEIVNYICQVIFWINFLIVIVCYTLITKELYRSYVRTRGVGKVPRKKVNVKVFIIIAVFFICFVPFHFARIPYTLSQTRDVFDCTAENTLFYVKESTLWLTSLNACLDPFIYFFLCKSFRNSLISMLKCPNSATSLSQDNRKKEQDGGDPNEETPM |
| UniProt | Q9H244 |
| Protein Data Bank | 4py0, 4pxz, 4ntj |
| GPCR-HGmod model | Q9H244 |
| 3D structure model | This structure is from PDB ID 4py0. |
| BioLiP | BL0272414,BL0272415, BL0276068, BL0276067, BL0276066, BL0272413 |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL2001 |
| IUPHAR | 328 |
| DrugBank | BE0000110 |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 432 | 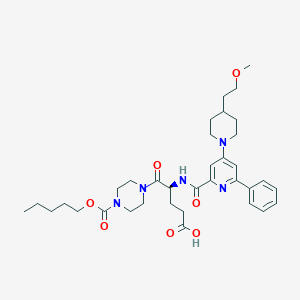 CHEMBL590001 CHEMBL590001 | C35H49N5O7 | 651.805 | 9 / 2 | 4.1 | No |
| 564 | 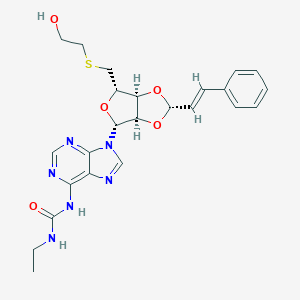 CHEMBL402633 CHEMBL402633 | C24H28N6O5S | 512.585 | 9 / 3 | 1.4 | No |
| 441721 | 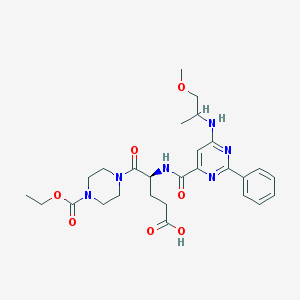 CHEMBL3322638 CHEMBL3322638 | C27H36N6O7 | 556.62 | 10 / 3 | 1.5 | No |
| 2338 | 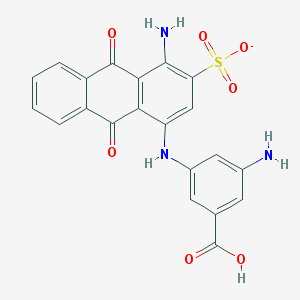 CHEMBL498038 CHEMBL498038 | C21H14N3O7S- | 452.417 | 10 / 4 | 2.3 | Yes |
| 441830 | 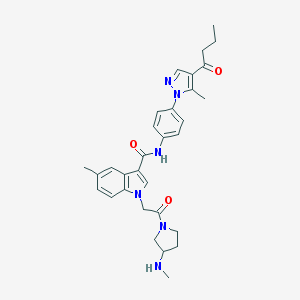 CHEMBL3325646 CHEMBL3325646 | C31H36N6O3 | 540.668 | 5 / 2 | 3.5 | No |
| 3055 | 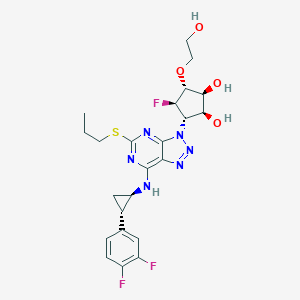 CHEMBL3098244 CHEMBL3098244 | C23H27F3N6O4S | 540.562 | 13 / 4 | 2.1 | No |
| 3056 | 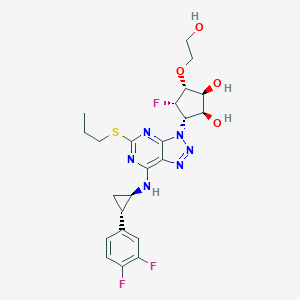 CHEMBL3098242 CHEMBL3098242 | C23H27F3N6O4S | 540.562 | 13 / 4 | 2.1 | No |
| 3757 | 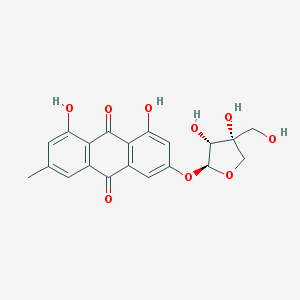 Frangulin B Frangulin B | C20H18O9 | 402.355 | 9 / 5 | 1.3 | Yes |
| 3761 | 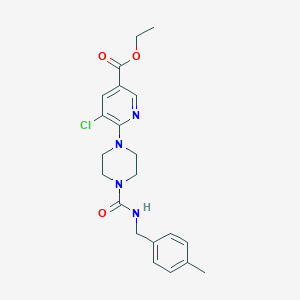 CHEMBL1784230 CHEMBL1784230 | C21H25ClN4O3 | 416.906 | 5 / 1 | 3.1 | Yes |
| 4158 | 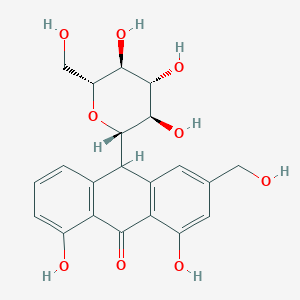 aloin aloin | C21H22O9 | 418.398 | 9 / 7 | -0.1 | No |
| 5200 | 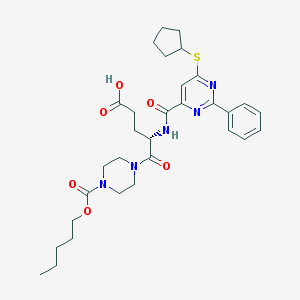 CHEMBL569912 CHEMBL569912 | C31H41N5O6S | 611.758 | 9 / 2 | 4.5 | No |
| 5466 | 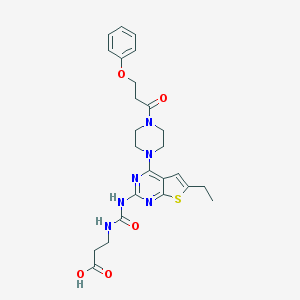 CHEMBL585306 CHEMBL585306 | C25H30N6O5S | 526.612 | 9 / 3 | 2.6 | No |
| 5496 | 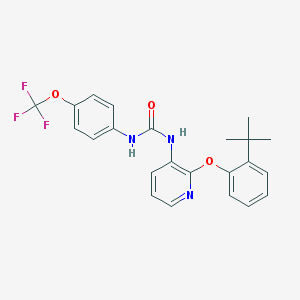 BPTU BPTU | C23H22F3N3O3 | 445.442 | 7 / 2 | 6.1 | No |
| 5706 | 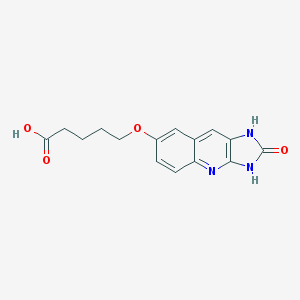 CHEMBL91224 CHEMBL91224 | C15H15N3O4 | 301.302 | 5 / 3 | 1.3 | Yes |
| 5797 | 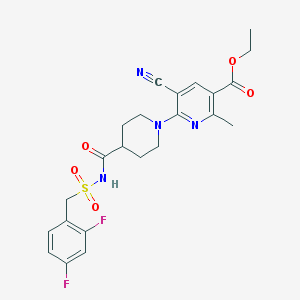 CHEMBL2419495 CHEMBL2419495 | C23H24F2N4O5S | 506.525 | 10 / 1 | 2.7 | No |
| 6156 | 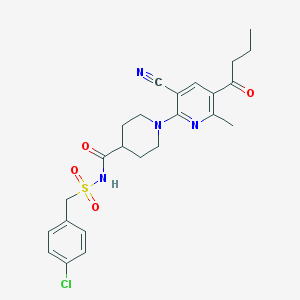 CHEMBL3288129 CHEMBL3288129 | C24H27ClN4O4S | 503.014 | 7 / 1 | 3.4 | No |
| 6157 | 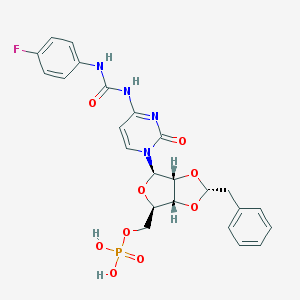 CHEMBL1162205 CHEMBL1162205 | C24H24FN4O9P | 562.447 | 10 / 4 | 0.4 | No |
| 442042 | 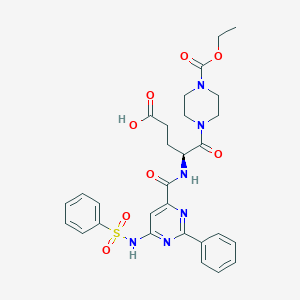 CHEMBL3322598 CHEMBL3322598 | C29H32N6O8S | 624.669 | 11 / 3 | 1.8 | No |
| 8364 | 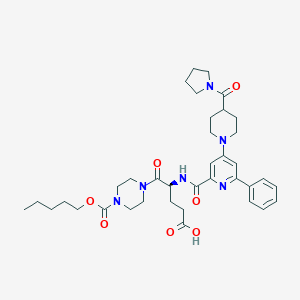 CHEMBL601907 CHEMBL601907 | C37H50N6O7 | 690.842 | 9 / 2 | 3.6 | No |
| 8390 | 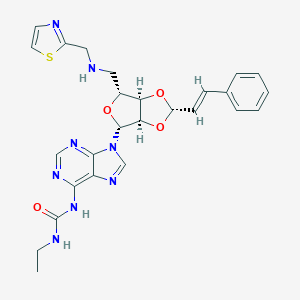 CHEMBL402423 CHEMBL402423 | C26H28N8O4S | 548.622 | 10 / 3 | 1.5 | No |
| 8532 | 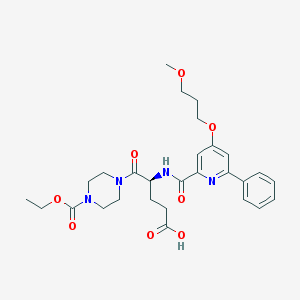 CHEMBL596939 CHEMBL596939 | C28H36N4O8 | 556.616 | 9 / 2 | 1.8 | No |
| 8838 | 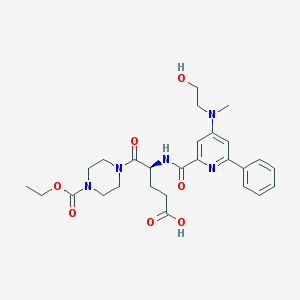 CHEMBL597980 CHEMBL597980 | C27H35N5O7 | 541.605 | 9 / 3 | 1.0 | No |
| 8989 | 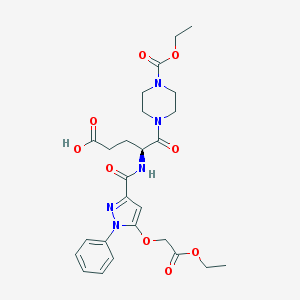 CHEMBL2172278 CHEMBL2172278 | C26H33N5O9 | 559.576 | 10 / 2 | 1.6 | No |
| 9273 | 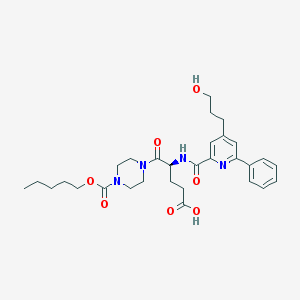 CHEMBL597150 CHEMBL597150 | C30H40N4O7 | 568.671 | 8 / 3 | 2.9 | No |
| 9414 | 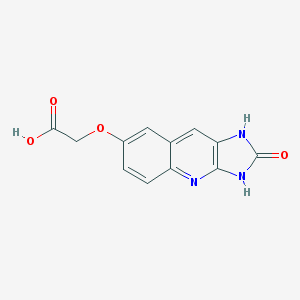 CHEMBL88525 CHEMBL88525 | C12H9N3O4 | 259.221 | 5 / 3 | 0.7 | Yes |
| 9631 | 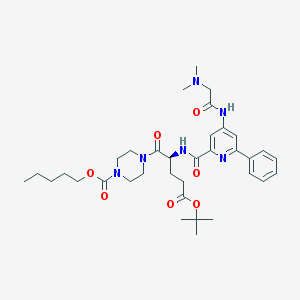 CHEMBL578482 CHEMBL578482 | C35H50N6O7 | 666.82 | 9 / 2 | 3.6 | No |
| 10079 | 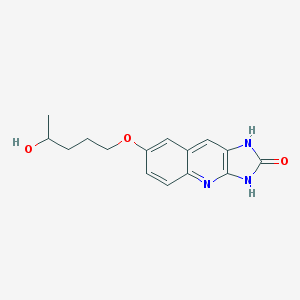 CHEMBL91359 CHEMBL91359 | C15H17N3O3 | 287.319 | 4 / 3 | 1.6 | Yes |
| 10289 | 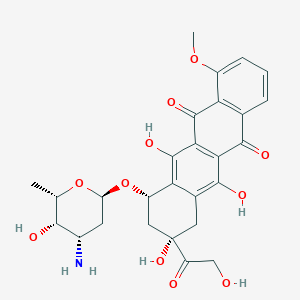 doxorubicin doxorubicin | C27H29NO11 | 543.525 | 12 / 6 | 1.3 | No |
| 10459 | 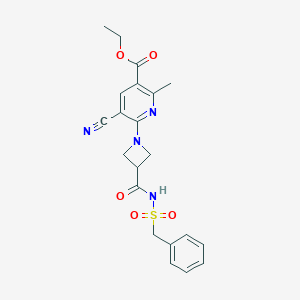 CHEMBL2419501 CHEMBL2419501 | C21H22N4O5S | 442.49 | 8 / 1 | 1.8 | Yes |
| 10775 | 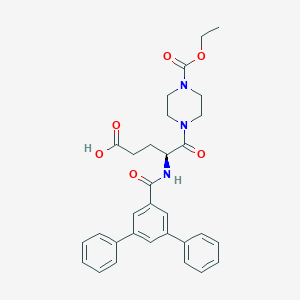 CHEMBL599606 CHEMBL599606 | C31H33N3O6 | 543.62 | 6 / 2 | 3.9 | No |
| 11005 | 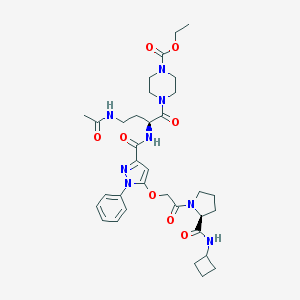 CHEMBL2172131 CHEMBL2172131 | C34H46N8O8 | 694.79 | 9 / 3 | 1.4 | No |
| 11964 | 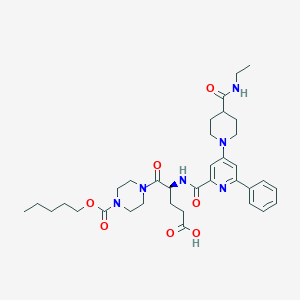 CHEMBL601696 CHEMBL601696 | C35H48N6O7 | 664.804 | 9 / 3 | 3.3 | No |
| 12269 | 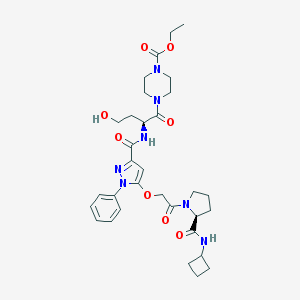 CHEMBL2172132 CHEMBL2172132 | C32H43N7O8 | 653.737 | 9 / 3 | 1.4 | No |
| 12588 | 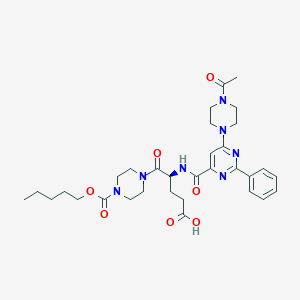 CHEMBL569629 CHEMBL569629 | C32H43N7O7 | 637.738 | 10 / 2 | 2.1 | No |
| 13310 | 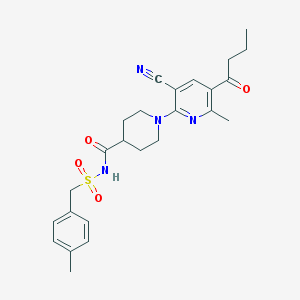 CHEMBL3288128 CHEMBL3288128 | C25H30N4O4S | 482.599 | 7 / 1 | 3.1 | Yes |
| 13484 | 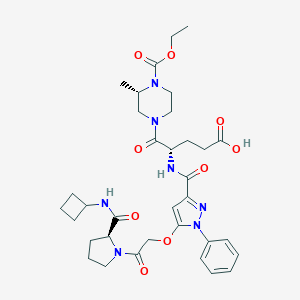 CHEMBL2172153 CHEMBL2172153 | C34H45N7O9 | 695.774 | 10 / 3 | 2.0 | No |
| 13485 | 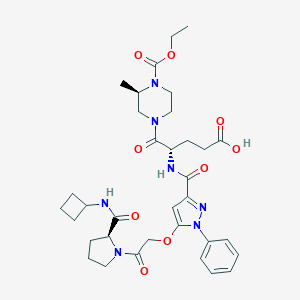 CHEMBL2172154 CHEMBL2172154 | C34H45N7O9 | 695.774 | 10 / 3 | 2.0 | No |
| 14129 | 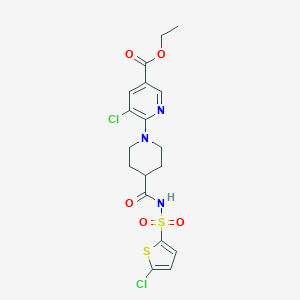 CHEMBL2419498 CHEMBL2419498 | C18H19Cl2N3O5S2 | 492.386 | 8 / 1 | 4.0 | Yes |
| 521884 | 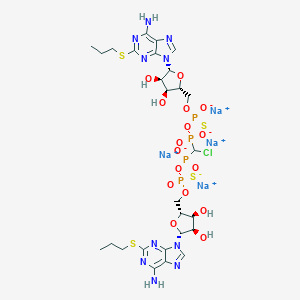 CHEMBL3745863 CHEMBL3745863 | C27H37ClN10Na4O16P4S4 | 1137.19 | 28 / 6 | N/A | No |
| 15063 | 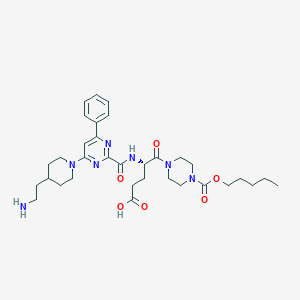 CHEMBL566613 CHEMBL566613 | C33H47N7O6 | 637.782 | 10 / 3 | 0.7 | No |
| 15081 | 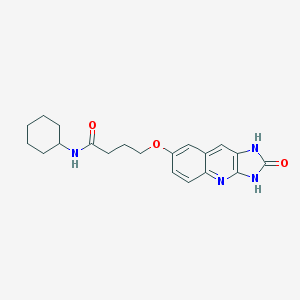 CHEMBL88218 CHEMBL88218 | C20H24N4O3 | 368.437 | 4 / 3 | 2.6 | Yes |
| 15655 | 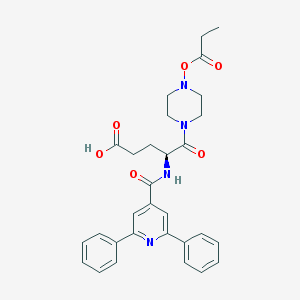 CHEMBL549594 CHEMBL549594 | C30H32N4O6 | 544.608 | 8 / 2 | 1.0 | No |
| 15901 | 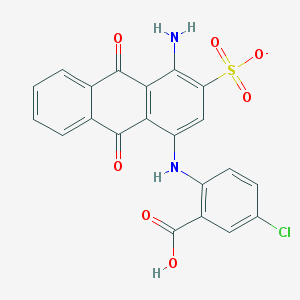 CHEMBL524284 CHEMBL524284 | C21H12ClN2O7S- | 471.844 | 9 / 3 | 3.7 | Yes |
| 16573 | 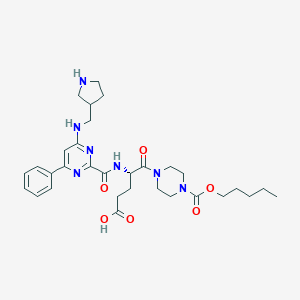 CHEMBL565249 CHEMBL565249 | C31H43N7O6 | 609.728 | 10 / 4 | 0.4 | No |
| 16578 | 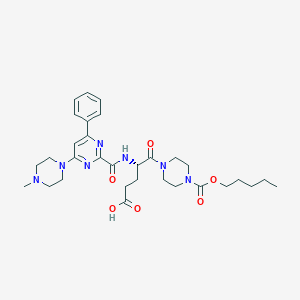 CHEMBL567840 CHEMBL567840 | C31H43N7O6 | 609.728 | 10 / 2 | 0.3 | No |
| 16744 | 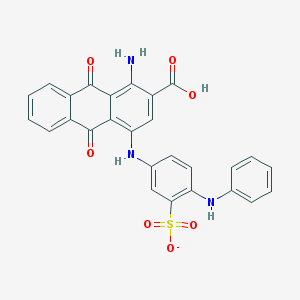 CHEMBL502519 CHEMBL502519 | C27H18N3O7S- | 528.515 | 10 / 4 | 5.1 | No |
| 16926 | 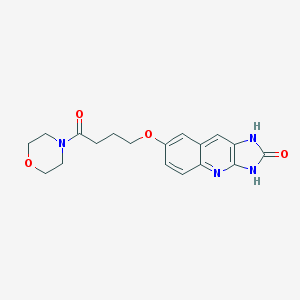 CHEMBL89660 CHEMBL89660 | C18H20N4O4 | 356.382 | 5 / 2 | 0.6 | Yes |
| 17488 | 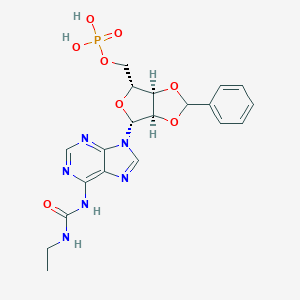 CHEMBL1162179 CHEMBL1162179 | C20H23N6O8P | 506.412 | 11 / 4 | -1.0 | No |
| 17548 | 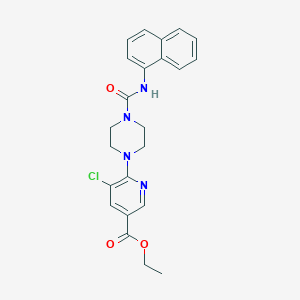 CHEMBL1784212 CHEMBL1784212 | C23H23ClN4O3 | 438.912 | 5 / 1 | 4.1 | Yes |
| 17593 | 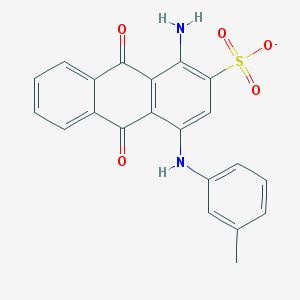 CHEMBL445413 CHEMBL445413 | C21H15N2O5S- | 407.42 | 7 / 2 | 3.9 | Yes |
| 18318 | 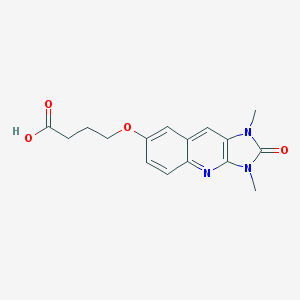 CHEMBL91579 CHEMBL91579 | C16H17N3O4 | 315.329 | 5 / 1 | 1.4 | Yes |
| 442388 | 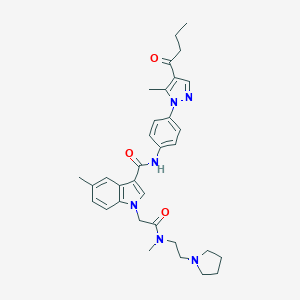 CHEMBL3325636 CHEMBL3325636 | C33H40N6O3 | 568.722 | 5 / 1 | 4.2 | No |
| 522028 | 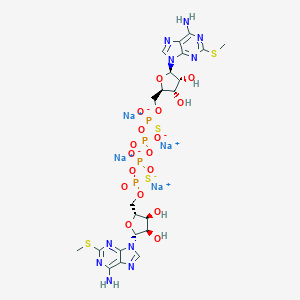 CHEMBL3746558 CHEMBL3746558 | C22H28N10Na4O17P4S4 | 1048.61 | 29 / 6 | N/A | No |
| 19155 | 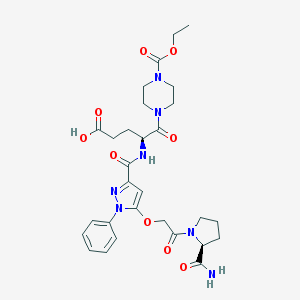 CHEMBL2172264 CHEMBL2172264 | C29H37N7O9 | 627.655 | 10 / 3 | 0.3 | No |
| 19378 | 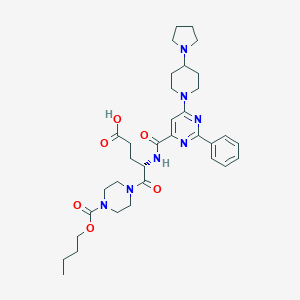 CHEMBL570064 CHEMBL570064 | C34H47N7O6 | 649.793 | 10 / 2 | 1.1 | No |
| 442436 | 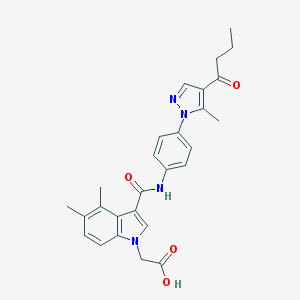 CHEMBL3325626 CHEMBL3325626 | C27H28N4O4 | 472.545 | 5 / 2 | 4.1 | Yes |
| 20379 | 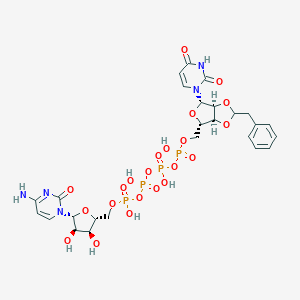 CHEMBL1162180 CHEMBL1162180 | C26H33N5O22P4 | 891.458 | 22 / 8 | -6.5 | No |
| 20695 | 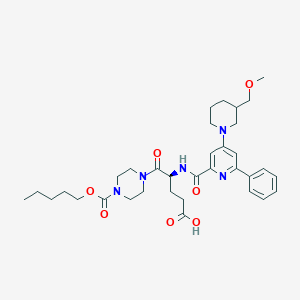 CHEMBL590256 CHEMBL590256 | C34H47N5O7 | 637.778 | 9 / 2 | 3.7 | No |
| 20743 | 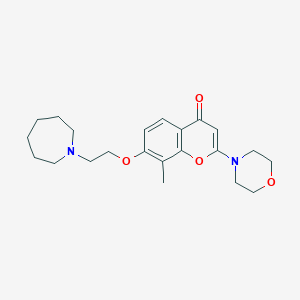 CHEMBL64383 CHEMBL64383 | C22H30N2O4 | 386.492 | 6 / 0 | 3.1 | Yes |
| 21220 | 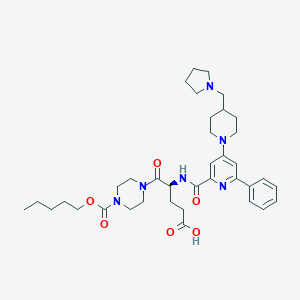 CHEMBL601692 CHEMBL601692 | C37H52N6O6 | 676.859 | 9 / 2 | 2.1 | No |
| 21759 | 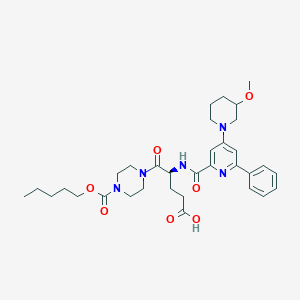 CHEMBL601270 CHEMBL601270 | C33H45N5O7 | 623.751 | 9 / 2 | 3.5 | No |
| 21767 | 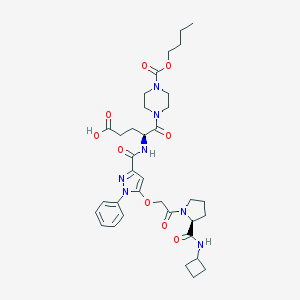 CHEMBL2172149 CHEMBL2172149 | C35H47N7O9 | 709.801 | 10 / 3 | 2.5 | No |
| 21904 | 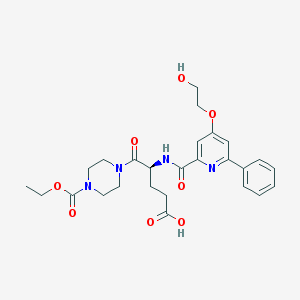 CHEMBL562485 CHEMBL562485 | C26H32N4O8 | 528.562 | 9 / 3 | 0.9 | No |
| 22141 | 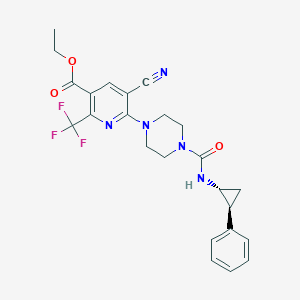 CHEMBL1784235 CHEMBL1784235 | C24H24F3N5O3 | 487.483 | 9 / 1 | 3.2 | Yes |
| 22509 | 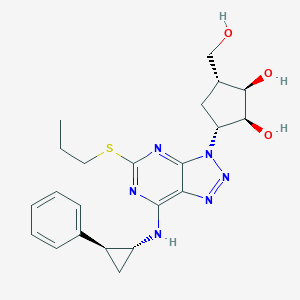 CHEMBL250687 CHEMBL250687 | C22H28N6O3S | 456.565 | 9 / 4 | 2.7 | Yes |
| 442550 | 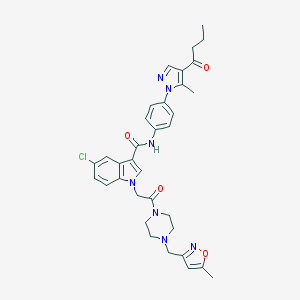 CHEMBL3325804 CHEMBL3325804 | C34H36ClN7O4 | 642.157 | 7 / 1 | 4.2 | No |
| 22747 | 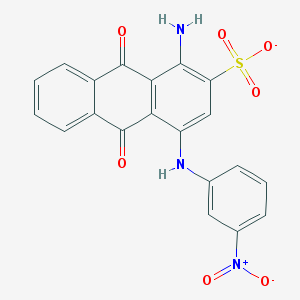 CHEMBL404659 CHEMBL404659 | C20H12N3O7S- | 438.39 | 9 / 2 | 3.3 | Yes |
| 22963 | 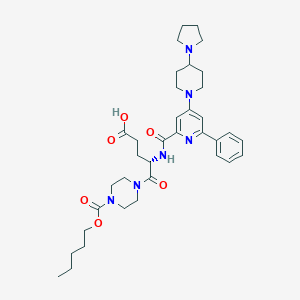 CHEMBL603144 CHEMBL603144 | C36H50N6O6 | 662.832 | 9 / 2 | 1.9 | No |
| 23040 | 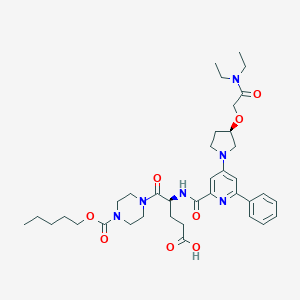 CHEMBL590069 CHEMBL590069 | C37H52N6O8 | 708.857 | 10 / 2 | 3.4 | No |
| 23278 | 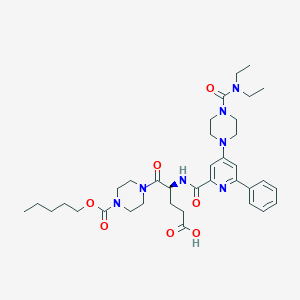 CHEMBL601447 CHEMBL601447 | C36H51N7O7 | 693.846 | 9 / 2 | 3.1 | No |
| 23409 | 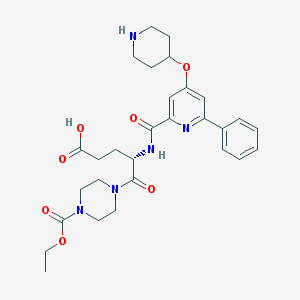 CHEMBL561007 CHEMBL561007 | C29H37N5O7 | 567.643 | 9 / 3 | -0.6 | No |
| 24004 | 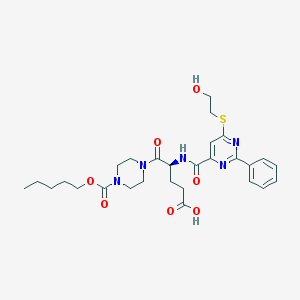 CHEMBL570423 CHEMBL570423 | C28H37N5O7S | 587.692 | 10 / 3 | 2.5 | No |
| 24478 | 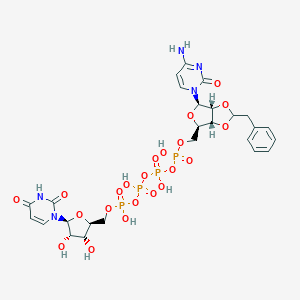 CHEMBL1162183 CHEMBL1162183 | C26H33N5O22P4 | 891.458 | 22 / 8 | -6.5 | No |
| 24573 | 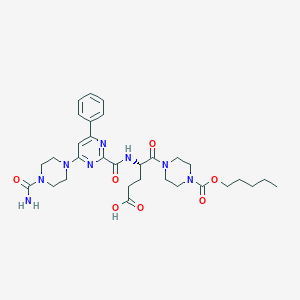 CHEMBL570150 CHEMBL570150 | C31H42N8O7 | 638.726 | 10 / 3 | 1.5 | No |
| 442650 | 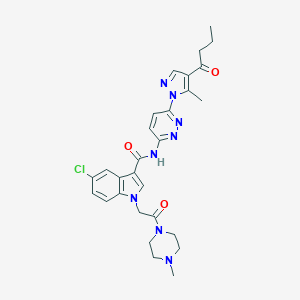 CHEMBL3325891 CHEMBL3325891 | C28H31ClN8O3 | 563.059 | 7 / 1 | 2.4 | No |
| 25106 | 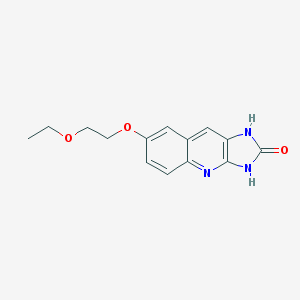 CHEMBL91899 CHEMBL91899 | C14H15N3O3 | 273.292 | 4 / 2 | 1.4 | Yes |
| 25107 | 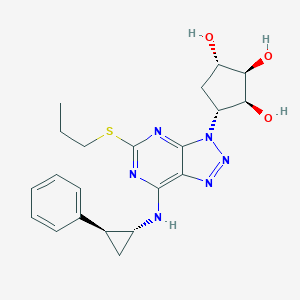 CHEMBL398295 CHEMBL398295 | C21H26N6O3S | 442.538 | 9 / 4 | 2.0 | Yes |
| 25327 | 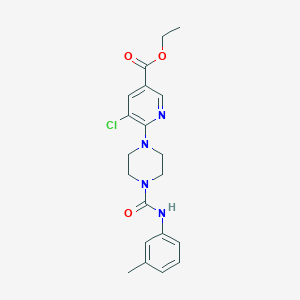 CHEMBL1784205 CHEMBL1784205 | C20H23ClN4O3 | 402.879 | 5 / 1 | 3.2 | Yes |
| 25649 | 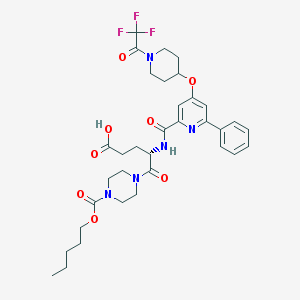 CHEMBL592887 CHEMBL592887 | C34H42F3N5O8 | 705.732 | 12 / 2 | 4.1 | No |
| 25929 | 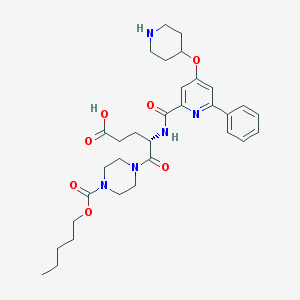 CHEMBL552972 CHEMBL552972 | C32H43N5O7 | 609.724 | 9 / 3 | 0.8 | No |
| 26414 | 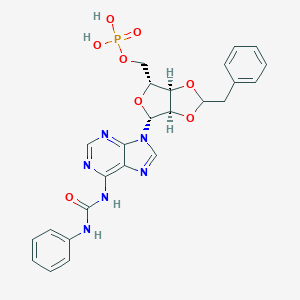 CHEMBL1162188 CHEMBL1162188 | C25H25N6O8P | 568.483 | 11 / 4 | 0.7 | No |
| 26539 | 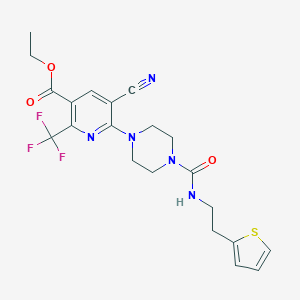 CHEMBL2402156 CHEMBL2402156 | C21H22F3N5O3S | 481.494 | 10 / 1 | 2.9 | Yes |
| 26844 | 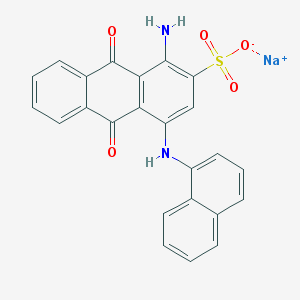 PSB 06126 PSB 06126 | C24H15N2NaO5S | 466.443 | 7 / 2 | N/A | N/A |
| 27762 | 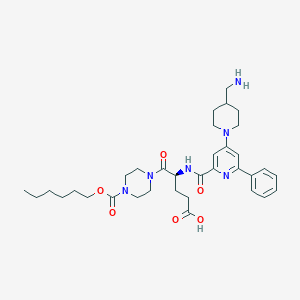 CHEMBL556330 CHEMBL556330 | C34H48N6O6 | 636.794 | 9 / 3 | 1.2 | No |
| 442778 | 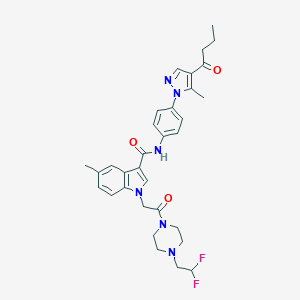 CHEMBL3325783 CHEMBL3325783 | C32H36F2N6O3 | 590.676 | 7 / 1 | 4.4 | No |
| 442781 | 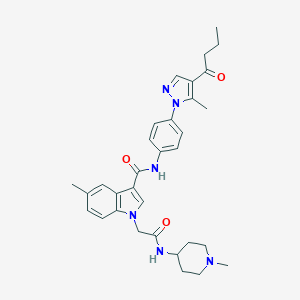 CHEMBL3325639 CHEMBL3325639 | C32H38N6O3 | 554.695 | 5 / 2 | 4.1 | No |
| 28457 | 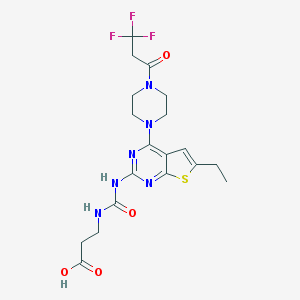 CHEMBL568567 CHEMBL568567 | C19H23F3N6O4S | 488.486 | 11 / 3 | 2.0 | No |
| 28577 | 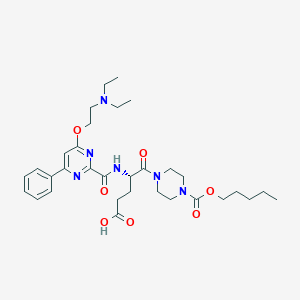 CHEMBL569908 CHEMBL569908 | C32H46N6O7 | 626.755 | 10 / 2 | 1.2 | No |
| 29143 | 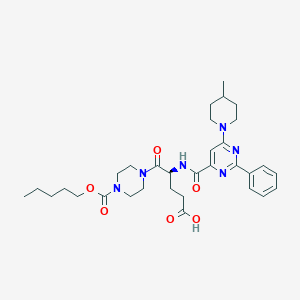 CHEMBL571929 CHEMBL571929 | C32H44N6O6 | 608.74 | 9 / 2 | 4.1 | No |
| 29148 | 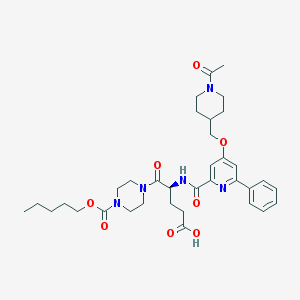 CHEMBL590950 CHEMBL590950 | C35H47N5O8 | 665.788 | 9 / 2 | 3.2 | No |
| 29362 | 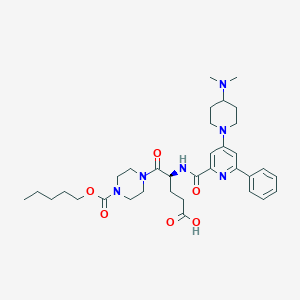 CHEMBL589198 CHEMBL589198 | C34H48N6O6 | 636.794 | 9 / 2 | 1.4 | No |
| 29912 | 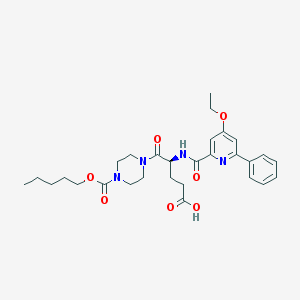 CHEMBL596961 CHEMBL596961 | C29H38N4O7 | 554.644 | 8 / 2 | 3.3 | No |
| 30277 | 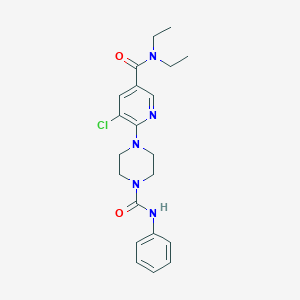 CHEMBL1784181 CHEMBL1784181 | C21H26ClN5O2 | 415.922 | 4 / 1 | 2.8 | Yes |
| 30445 | 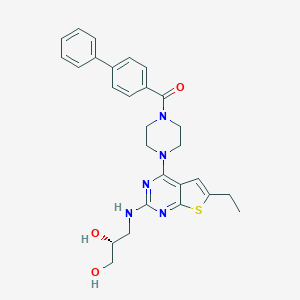 CHEMBL568148 CHEMBL568148 | C28H31N5O3S | 517.648 | 8 / 3 | 5.0 | No |
| 31033 | 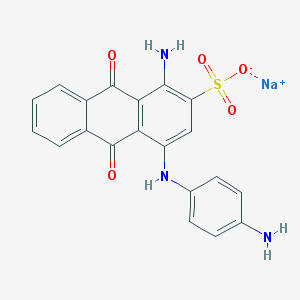 40951-76-6 40951-76-6 | C20H14N3NaO5S | 431.398 | 8 / 3 | N/A | N/A |
| 442918 | 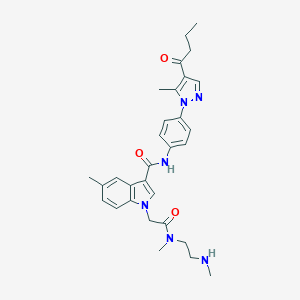 CHEMBL3325633 CHEMBL3325633 | C30H36N6O3 | 528.657 | 5 / 2 | 3.3 | No |
| 32240 | 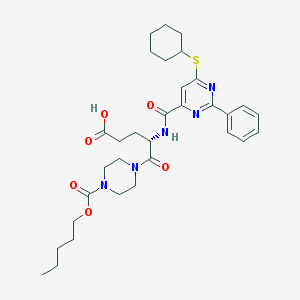 CHEMBL569913 CHEMBL569913 | C32H43N5O6S | 625.785 | 9 / 2 | 5.0 | No |
| 32562 | 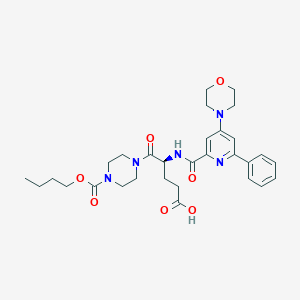 CHEMBL590364 CHEMBL590364 | C30H39N5O7 | 581.67 | 9 / 2 | 2.2 | No |
| 32800 | 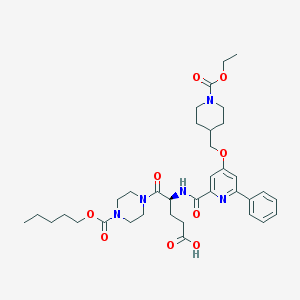 CHEMBL600575 CHEMBL600575 | C36H49N5O9 | 695.814 | 10 / 2 | 4.0 | No |
| 33140 | 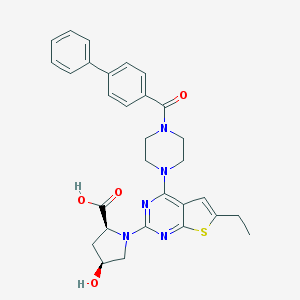 CHEMBL568334 CHEMBL568334 | C30H31N5O4S | 557.669 | 9 / 2 | 5.0 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218