

You can:
| Name | Urotensin-2 receptor |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | UTS2R |
| Synonym | urotensin II receptor UT receptor UR-II-R UR-2-R UII-R1 [ Show all ] |
| Disease | Asthma Diabetic nephropathy Renal failure |
| Length | 389 |
| Amino acid sequence | MALTPESPSSFPGLAATGSSVPEPPGGPNATLNSSWASPTEPSSLEDLVATGTIGTLLSAMGVVGVVGNAYTLVVTCRSLRAVASMYVYVVNLALADLLYLLSIPFIVATYVTKEWHFGDVGCRVLFGLDFLTMHASIFTLTVMSSERYAAVLRPLDTVQRPKGYRKLLALGTWLLALLLTLPVMLAMRLVRRGPKSLCLPAWGPRAHRAYLTLLFATSIAGPGLLIGLLYARLARAYRRSQRASFKRARRPGARALRLVLGIVLLFWACFLPFWLWQLLAQYHQAPLAPRTARIVNYLTTCLTYGNSCANPFLYTLLTRNYRDHLRGRVRGPGSGGGRGPVPSLQPRARFQRCSGRSLSSCSPQPTDSLVLAPAAPARPAPEGPRAPA |
| UniProt | Q9UKP6 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | Q9UKP6 |
| 3D structure model | This predicted structure model is from GPCR-EXP Q9UKP6. |
| BioLiP | N/A |
| Therapeutic Target Database | T49072 |
| ChEMBL | CHEMBL3764 |
| IUPHAR | 365 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 547902 | 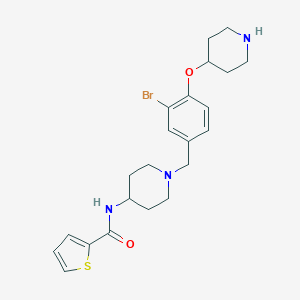 CHEMBL3979822 CHEMBL3979822 | C22H28BrN3O2S | 478.449 | 5 / 2 | 4.1 | Yes |
| 822 | 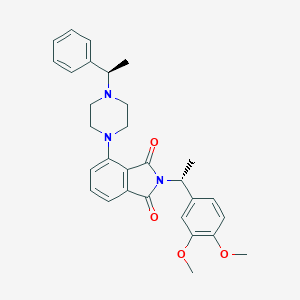 CHEMBL571683 CHEMBL571683 | C30H33N3O4 | 499.611 | 6 / 0 | 4.6 | Yes |
| 825 | 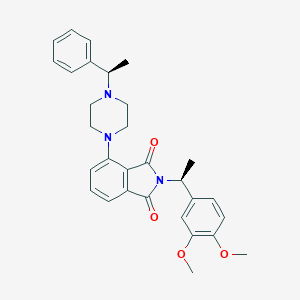 CHEMBL577518 CHEMBL577518 | C30H33N3O4 | 499.611 | 6 / 0 | 4.6 | Yes |
| 827 | 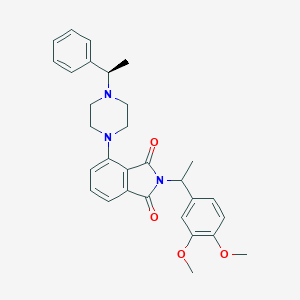 CHEMBL566389 CHEMBL566389 | C30H33N3O4 | 499.611 | 6 / 0 | 4.6 | Yes |
| 1106 | 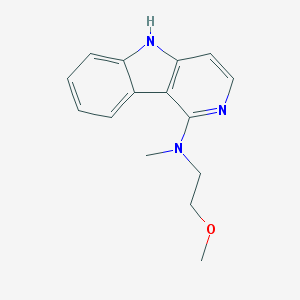 CHEMBL451255 CHEMBL451255 | C15H17N3O | 255.321 | 3 / 1 | 2.6 | Yes |
| 1434 | 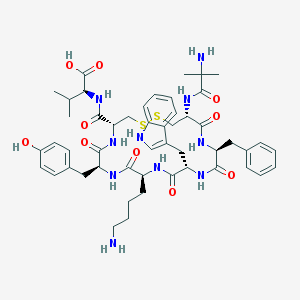 CHEMBL218825 CHEMBL218825 | C50H66N10O10S2 | 1031.26 | 14 / 12 | 0.4 | No |
| 1489 | 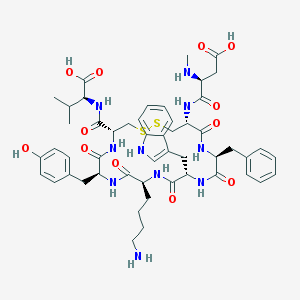 CHEMBL374468 CHEMBL374468 | C51H66N10O12S2 | 1075.27 | 16 / 13 | -2.4 | No |
| 1758 | 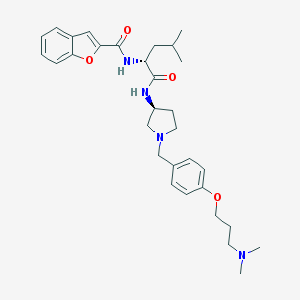 CHEMBL190975 CHEMBL190975 | C31H42N4O4 | 534.701 | 6 / 2 | 5.1 | No |
| 1759 | 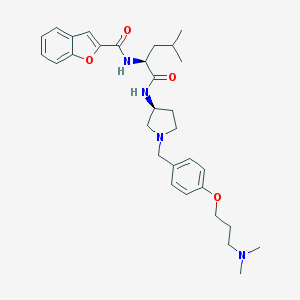 CHEMBL361664 CHEMBL361664 | C31H42N4O4 | 534.701 | 6 / 2 | 5.1 | No |
| 1760 | 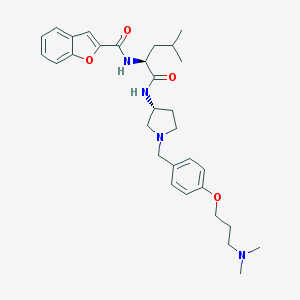 CHEMBL192295 CHEMBL192295 | C31H42N4O4 | 534.701 | 6 / 2 | 5.1 | No |
| 2949 | 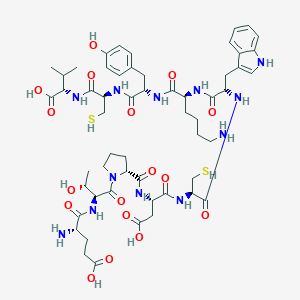 CHEMBL412753 CHEMBL412753 | C55H78N12O17S2 | 1243.42 | 21 / 18 | -5.7 | No |
| 3086 | 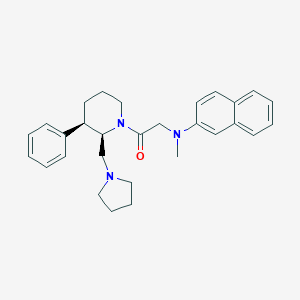 CHEMBL437216 CHEMBL437216 | C29H35N3O | 441.619 | 3 / 0 | 5.7 | No |
| 3529 | 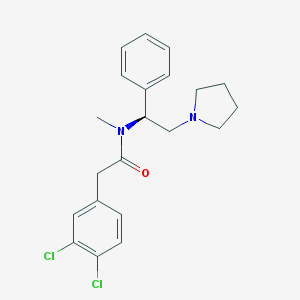 CHEMBL38576 CHEMBL38576 | C21H24Cl2N2O | 391.336 | 2 / 0 | 4.6 | Yes |
| 5661 | 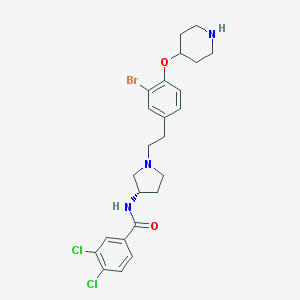 CHEMBL479994 CHEMBL479994 | C24H28BrCl2N3O2 | 541.311 | 4 / 2 | 5.4 | No |
| 7099 | 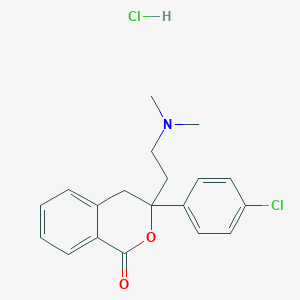 477313-09-0 477313-09-0 | C19H21Cl2NO2 | 366.282 | 3 / 1 | N/A | N/A |
| 8301 | 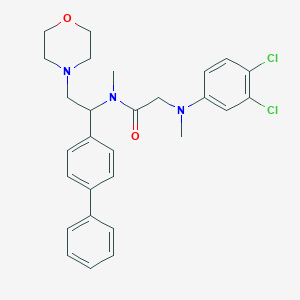 CHEMBL452298 CHEMBL452298 | C28H31Cl2N3O2 | 512.475 | 4 / 0 | 5.6 | No |
| 9208 | 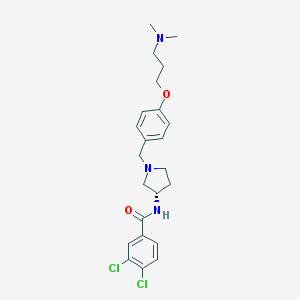 SB-436811 SB-436811 | C23H29Cl2N3O2 | 450.404 | 4 / 1 | 4.5 | Yes |
| 10458 | 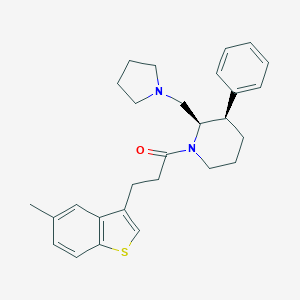 CHEMBL412746 CHEMBL412746 | C28H34N2OS | 446.653 | 3 / 0 | 5.8 | No |
| 521741 | 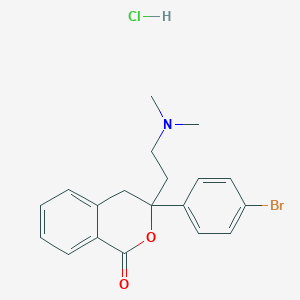 CHEMBL1172605 CHEMBL1172605 | C19H21BrClNO2 | 410.736 | 3 / 1 | N/A | N/A |
| 11836 | 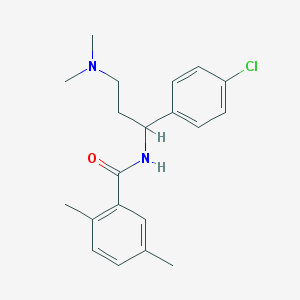 CHEMBL382401 CHEMBL382401 | C20H25ClN2O | 344.883 | 2 / 1 | 4.5 | Yes |
| 12824 | 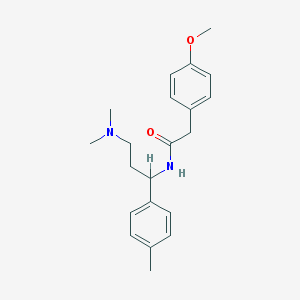 CHEMBL228526 CHEMBL228526 | C21H28N2O2 | 340.467 | 3 / 1 | 3.4 | Yes |
| 13471 | 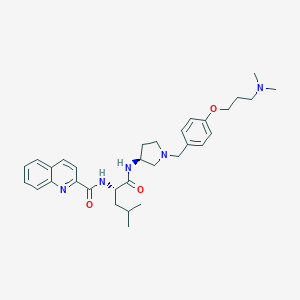 CHEMBL193033 CHEMBL193033 | C32H43N5O3 | 545.728 | 6 / 2 | 5.0 | No |
| 15250 | 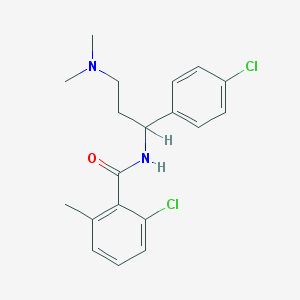 CHEMBL382688 CHEMBL382688 | C19H22Cl2N2O | 365.298 | 2 / 1 | 4.7 | Yes |
| 521931 | 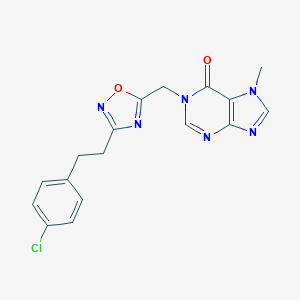 AM-0902 AM-0902 | C17H15ClN6O2 | 370.797 | 6 / 0 | 2.3 | Yes |
| 16681 | 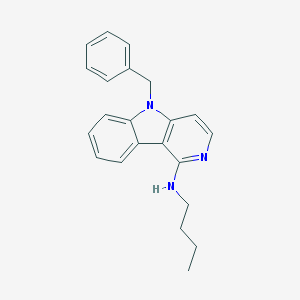 CHEMBL495919 CHEMBL495919 | C22H23N3 | 329.447 | 2 / 1 | 5.4 | No |
| 16813 | 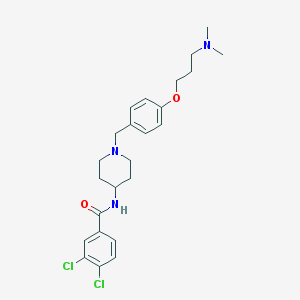 CHEMBL481129 CHEMBL481129 | C24H31Cl2N3O2 | 464.431 | 4 / 1 | 4.9 | Yes |
| 18656 | 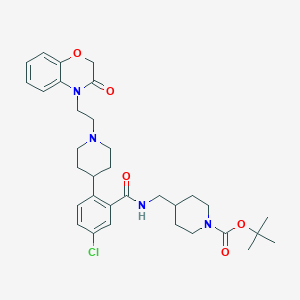 CHEMBL238066 CHEMBL238066 | C33H43ClN4O5 | 611.18 | 6 / 1 | 4.9 | No |
| 18747 | 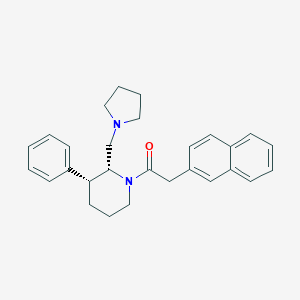 CHEMBL257187 CHEMBL257187 | C28H32N2O | 412.577 | 2 / 0 | 5.4 | No |
| 19268 | 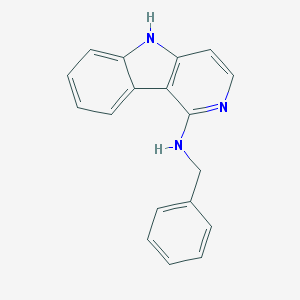 CHEMBL500880 CHEMBL500880 | C18H15N3 | 273.339 | 2 / 2 | 4.1 | Yes |
| 19492 | 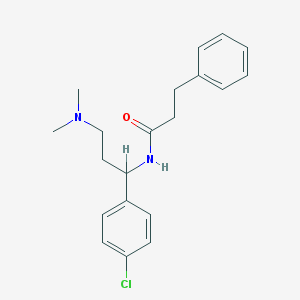 CHEMBL228412 CHEMBL228412 | C20H25ClN2O | 344.883 | 2 / 1 | 4.0 | Yes |
| 21648 | 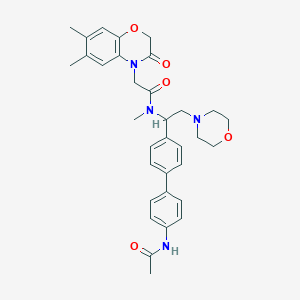 CHEMBL449192 CHEMBL449192 | C33H38N4O5 | 570.69 | 6 / 1 | 3.3 | No |
| 22521 | 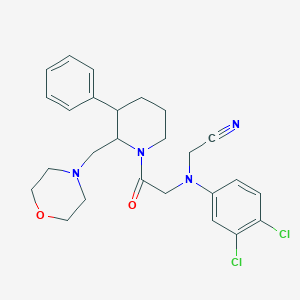 CHEMBL258251 CHEMBL258251 | C26H30Cl2N4O2 | 501.452 | 5 / 0 | 4.6 | No |
| 23476 | 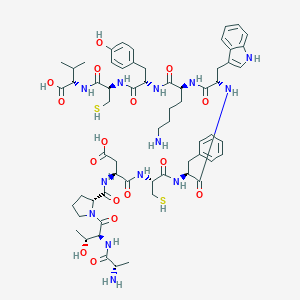 CHEMBL2372899 CHEMBL2372899 | C62H85N13O16S2 | 1332.56 | 20 / 18 | -3.9 | No |
| 23477 | 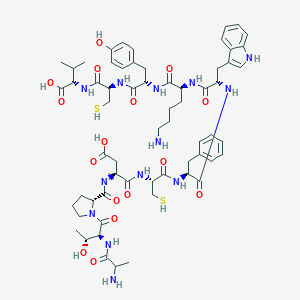 CHEMBL414728 CHEMBL414728 | C62H85N13O16S2 | 1332.56 | 20 / 18 | -3.9 | No |
| 24914 | 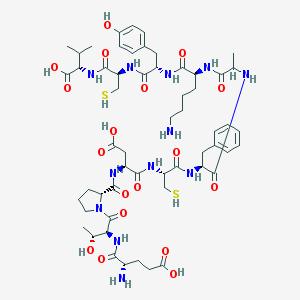 CHEMBL441391 CHEMBL441391 | C56H82N12O18S2 | 1275.46 | 22 / 18 | -6.1 | No |
| 24915 | 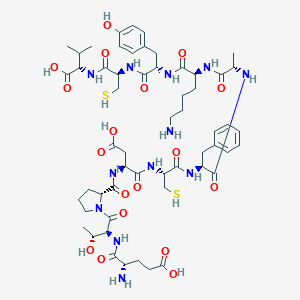 CHEMBL2372905 CHEMBL2372905 | C56H82N12O18S2 | 1275.46 | 22 / 18 | -6.1 | No |
| 25631 | 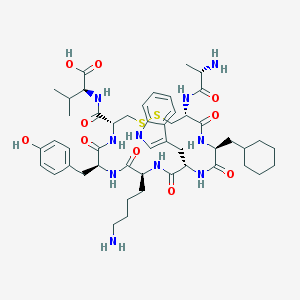 urotensin-related peptide urotensin-related peptide | C49H70N10O10S2 | 1023.28 | 14 / 12 | 1.4 | No |
| 25750 | 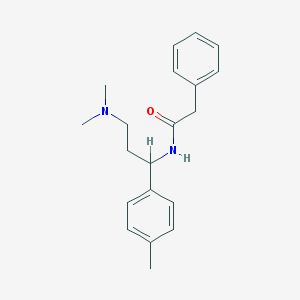 CHEMBL228472 CHEMBL228472 | C20H26N2O | 310.441 | 2 / 1 | 3.4 | Yes |
| 548182 | 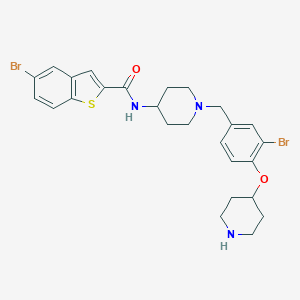 CHEMBL3915670 CHEMBL3915670 | C26H29Br2N3O2S | 607.405 | 5 / 2 | 6.1 | No |
| 27265 | 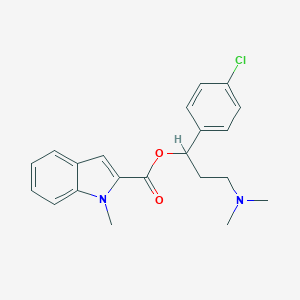 CHEMBL377946 CHEMBL377946 | C21H23ClN2O2 | 370.877 | 3 / 0 | 4.9 | Yes |
| 27632 | 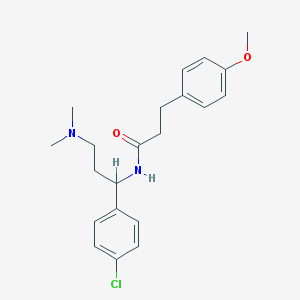 CHEMBL228461 CHEMBL228461 | C21H27ClN2O2 | 374.909 | 3 / 1 | 4.0 | Yes |
| 28115 | 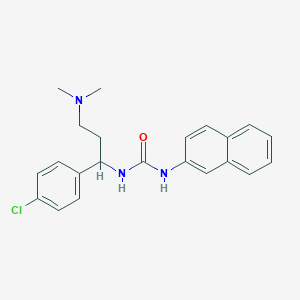 CHEMBL206551 CHEMBL206551 | C22H24ClN3O | 381.904 | 2 / 2 | 4.7 | Yes |
| 522402 | 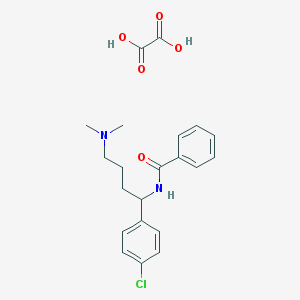 CHEMBL573836 CHEMBL573836 | C21H25ClN2O5 | 420.89 | 6 / 3 | N/A | N/A |
| 29748 | 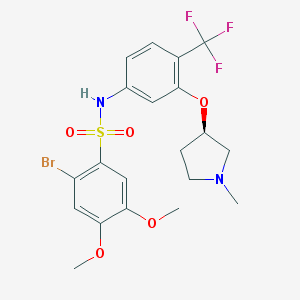 SB 706375 SB 706375 | C20H22BrF3N2O5S | 539.364 | 10 / 1 | 4.2 | No |
| 522476 | 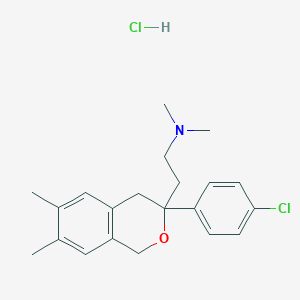 CHEMBL1173492 CHEMBL1173492 | C21H27Cl2NO | 380.353 | 2 / 1 | N/A | N/A |
| 30747 | 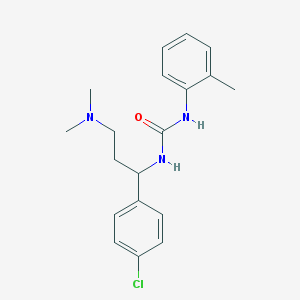 CHEMBL205390 CHEMBL205390 | C19H24ClN3O | 345.871 | 2 / 2 | 3.8 | Yes |
| 548276 | 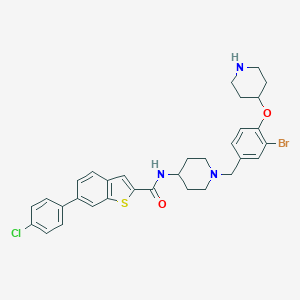 CHEMBL3907266 CHEMBL3907266 | C32H33BrClN3O2S | 639.049 | 5 / 2 | 7.7 | No |
| 33735 | 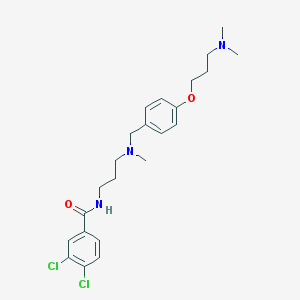 CHEMBL481130 CHEMBL481130 | C23H31Cl2N3O2 | 452.42 | 4 / 1 | 4.7 | Yes |
| 34945 | 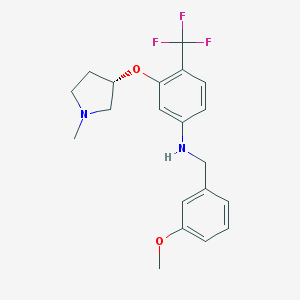 CHEMBL2348517 CHEMBL2348517 | C20H23F3N2O2 | 380.411 | 7 / 1 | 4.5 | Yes |
| 36168 | 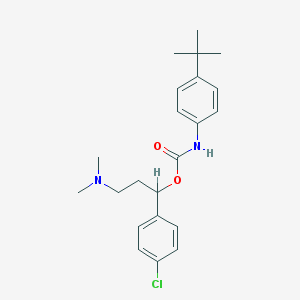 CHEMBL377973 CHEMBL377973 | C22H29ClN2O2 | 388.936 | 3 / 1 | 5.7 | No |
| 36741 | 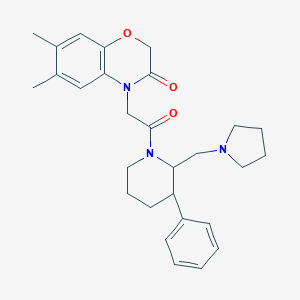 CHEMBL257171 CHEMBL257171 | C28H35N3O3 | 461.606 | 4 / 0 | 4.1 | Yes |
| 38530 | 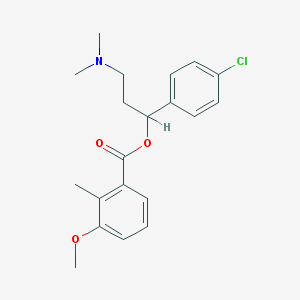 CHEMBL378235 CHEMBL378235 | C20H24ClNO3 | 361.866 | 4 / 0 | 4.7 | Yes |
| 39986 | 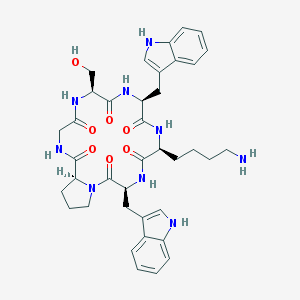 CHEMBL608099 CHEMBL608099 | C38H47N9O7 | 741.85 | 8 / 9 | 1.1 | No |
| 42185 | 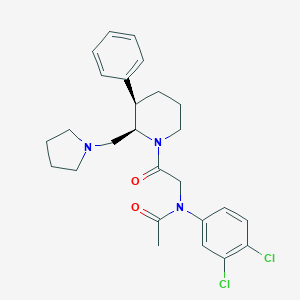 CHEMBL256159 CHEMBL256159 | C26H31Cl2N3O2 | 488.453 | 3 / 0 | 4.9 | Yes |
| 44795 | 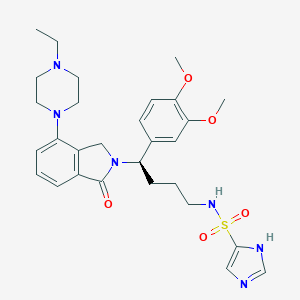 CHEMBL578207 CHEMBL578207 | C29H38N6O5S | 582.72 | 9 / 2 | 2.6 | No |
| 45369 | 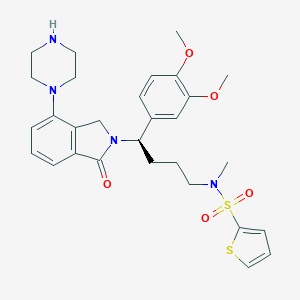 CHEMBL565416 CHEMBL565416 | C29H36N4O5S2 | 584.75 | 9 / 1 | 3.5 | No |
| 45842 | 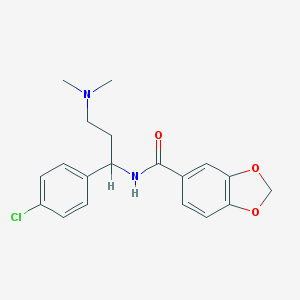 CHEMBL205628 CHEMBL205628 | C19H21ClN2O3 | 360.838 | 4 / 1 | 3.6 | Yes |
| 46366 | 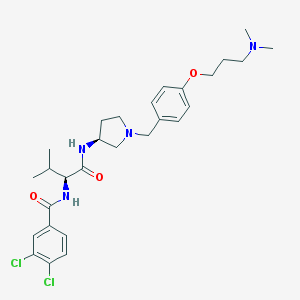 CHEMBL189525 CHEMBL189525 | C28H38Cl2N4O3 | 549.537 | 5 / 2 | 5.3 | No |
| 47513 | 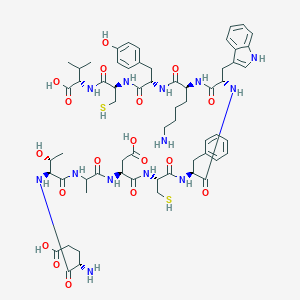 CHEMBL436891 CHEMBL436891 | C62H85N13O18S2 | 1364.56 | 22 / 20 | -4.7 | No |
| 47514 | 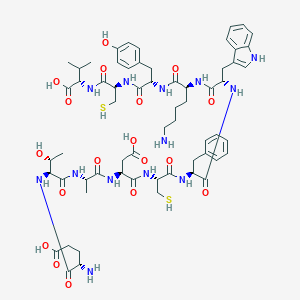 CHEMBL2372897 CHEMBL2372897 | C62H85N13O18S2 | 1364.56 | 22 / 20 | -4.7 | No |
| 49191 | 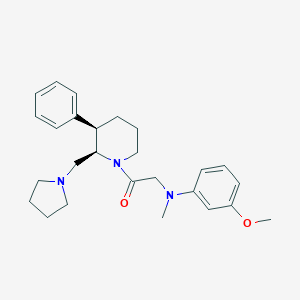 CHEMBL253747 CHEMBL253747 | C26H35N3O2 | 421.585 | 4 / 0 | 4.4 | Yes |
| 50161 | 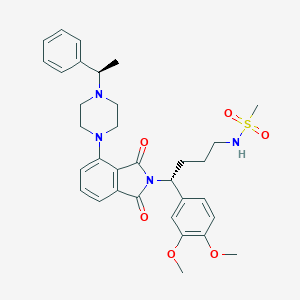 CHEMBL569689 CHEMBL569689 | C33H40N4O6S | 620.765 | 9 / 1 | 4.0 | No |
| 443690 | 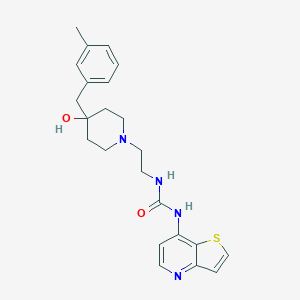 CHEMBL3358683 CHEMBL3358683 | C23H28N4O2S | 424.563 | 5 / 3 | 3.0 | Yes |
| 50415 | 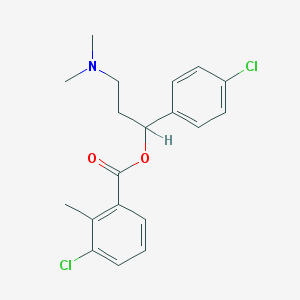 CHEMBL274508 CHEMBL274508 | C19H21Cl2NO2 | 366.282 | 3 / 0 | 5.3 | No |
| 50620 | 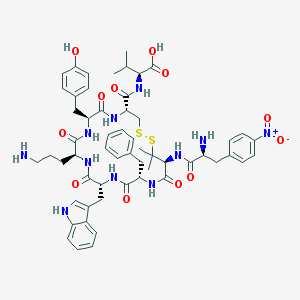 CHEMBL500949 CHEMBL500949 | C56H69N11O12S2 | 1152.35 | 16 / 12 | 1.9 | No |
| 523056 | 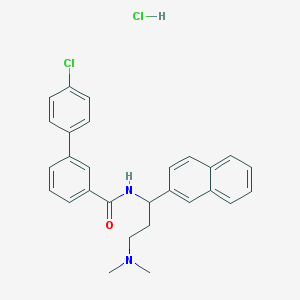 CHEMBL574726 CHEMBL574726 | C28H28Cl2N2O | 479.445 | 2 / 2 | N/A | N/A |
| 52184 | 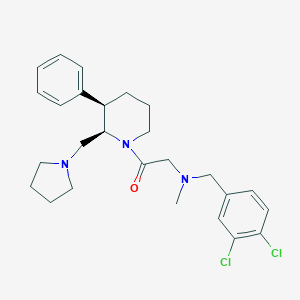 CHEMBL253540 CHEMBL253540 | C26H33Cl2N3O | 474.47 | 3 / 0 | 5.4 | No |
| 52456 | 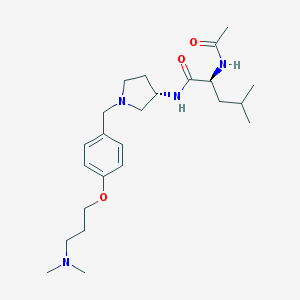 CHEMBL410825 CHEMBL410825 | C24H40N4O3 | 432.609 | 5 / 2 | 2.7 | Yes |
| 52685 | 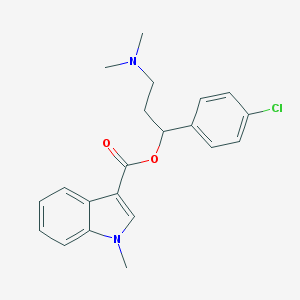 CHEMBL204516 CHEMBL204516 | C21H23ClN2O2 | 370.877 | 3 / 0 | 4.4 | Yes |
| 52693 | 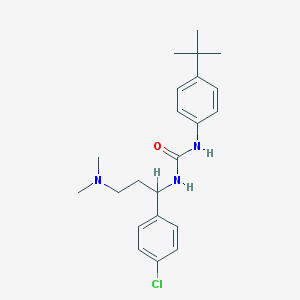 CHEMBL205681 CHEMBL205681 | C22H30ClN3O | 387.952 | 2 / 2 | 5.2 | No |
| 53886 | 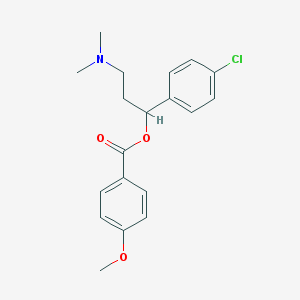 CHEMBL208886 CHEMBL208886 | C19H22ClNO3 | 347.839 | 4 / 0 | 4.3 | Yes |
| 54142 | 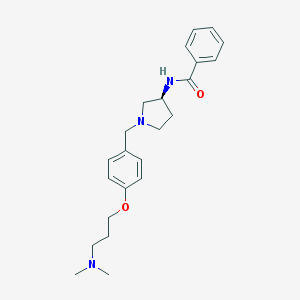 CHEMBL372911 CHEMBL372911 | C23H31N3O2 | 381.52 | 4 / 1 | 3.3 | Yes |
| 54724 | 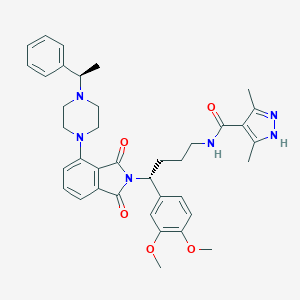 CHEMBL565388 CHEMBL565388 | C38H44N6O5 | 664.807 | 8 / 2 | 5.0 | No |
| 56103 | 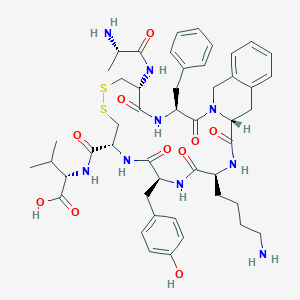 CHEMBL3104467 CHEMBL3104467 | C48H63N9O10S2 | 990.205 | 14 / 10 | -0.2 | No |
| 56105 | 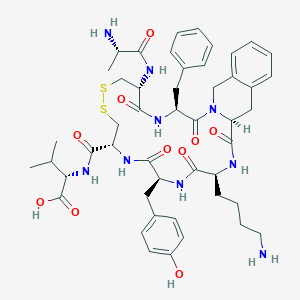 CHEMBL3104466 CHEMBL3104466 | C48H63N9O10S2 | 990.205 | 14 / 10 | -0.2 | No |
| 56384 | 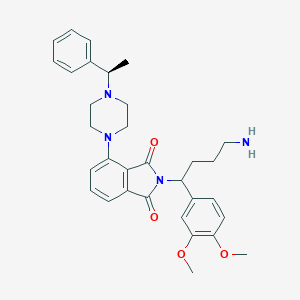 CHEMBL584545 CHEMBL584545 | C32H38N4O4 | 542.68 | 7 / 1 | 4.0 | No |
| 56796 | 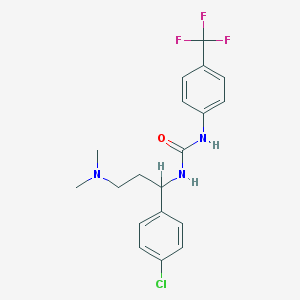 CHEMBL425592 CHEMBL425592 | C19H21ClF3N3O | 399.842 | 5 / 2 | 4.4 | Yes |
| 444060 | 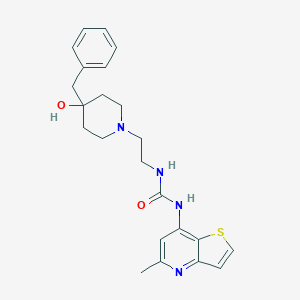 CHEMBL3358679 CHEMBL3358679 | C23H28N4O2S | 424.563 | 5 / 3 | 3.0 | Yes |
| 61530 | 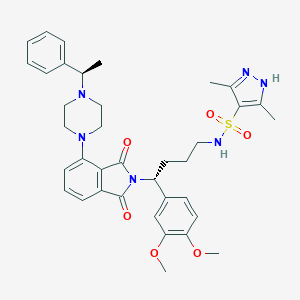 CHEMBL567759 CHEMBL567759 | C37H44N6O6S | 700.855 | 10 / 2 | 4.7 | No |
| 62736 | 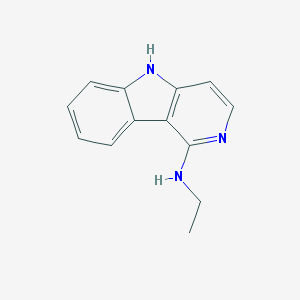 CHEMBL447747 CHEMBL447747 | C13H13N3 | 211.268 | 2 / 2 | 3.0 | Yes |
| 63479 | 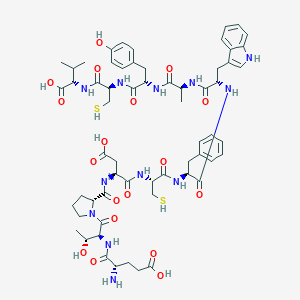 CHEMBL2372904 CHEMBL2372904 | C61H80N12O18S2 | 1333.5 | 21 / 18 | -1.9 | No |
| 63480 | 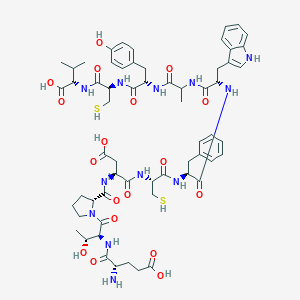 CHEMBL429558 CHEMBL429558 | C61H80N12O18S2 | 1333.5 | 21 / 18 | -1.9 | No |
| 523386 | 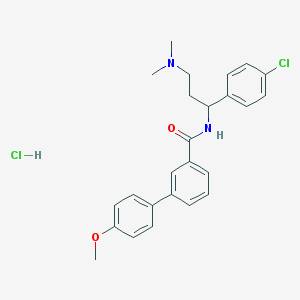 CHEMBL574730 CHEMBL574730 | C25H28Cl2N2O2 | 459.411 | 3 / 2 | N/A | N/A |
| 64373 | 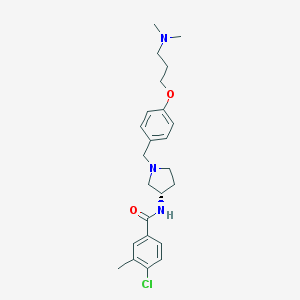 CHEMBL193266 CHEMBL193266 | C24H32ClN3O2 | 429.989 | 4 / 1 | 4.3 | Yes |
| 64570 | 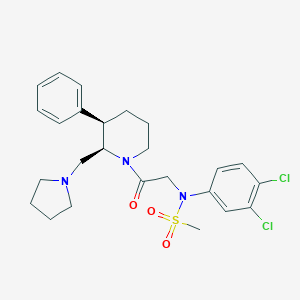 CHEMBL256300 CHEMBL256300 | C25H31Cl2N3O3S | 524.501 | 5 / 0 | 4.7 | No |
| 64790 | 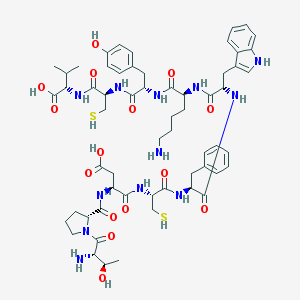 CHEMBL427632 CHEMBL427632 | C59H80N12O15S2 | 1261.48 | 19 / 17 | -3.6 | No |
| 65778 | 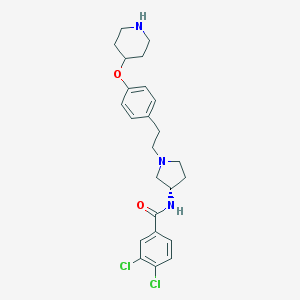 CHEMBL480165 CHEMBL480165 | C24H29Cl2N3O2 | 462.415 | 4 / 2 | 4.7 | Yes |
| 65825 | 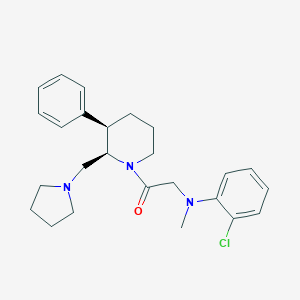 CHEMBL404244 CHEMBL404244 | C25H32ClN3O | 426.001 | 3 / 0 | 5.0 | Yes |
| 444308 | 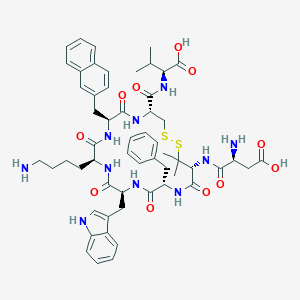 CHEMBL3315140 CHEMBL3315140 | C56H70N10O11S2 | 1123.35 | 15 / 12 | -0.7 | No |
| 67401 | 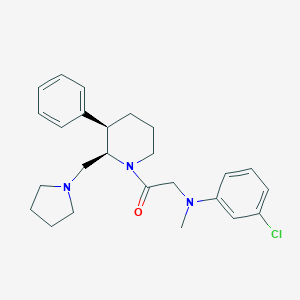 CHEMBL256299 CHEMBL256299 | C25H32ClN3O | 426.001 | 3 / 0 | 5.0 | Yes |
| 68334 | 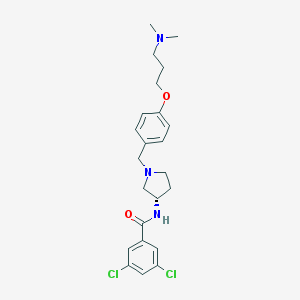 CHEMBL190137 CHEMBL190137 | C23H29Cl2N3O2 | 450.404 | 4 / 1 | 4.5 | Yes |
| 68752 | 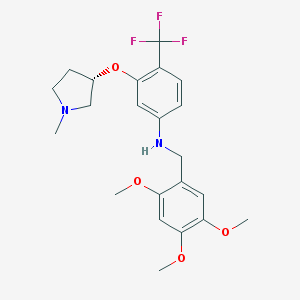 CHEMBL2348520 CHEMBL2348520 | C22H27F3N2O4 | 440.463 | 9 / 1 | 4.4 | Yes |
| 68835 | 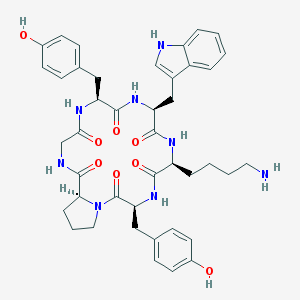 CHEMBL607810 CHEMBL607810 | C42H50N8O8 | 794.91 | 9 / 9 | 2.3 | No |
| 471465 | 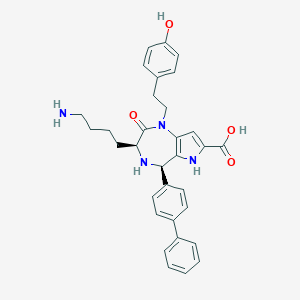 CHEMBL3577310 CHEMBL3577310 | C32H34N4O4 | 538.648 | 6 / 5 | 1.9 | No |
| 69796 | 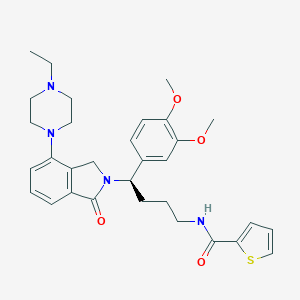 CHEMBL567075 CHEMBL567075 | C31H38N4O4S | 562.729 | 7 / 1 | 4.4 | No |
| 70101 | 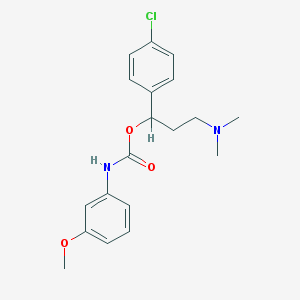 CHEMBL208536 CHEMBL208536 | C19H23ClN2O3 | 362.854 | 4 / 1 | 4.0 | Yes |
| 70421 | 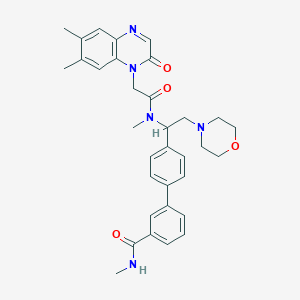 CHEMBL499582 CHEMBL499582 | C33H37N5O4 | 567.69 | 6 / 1 | 3.2 | No |
| 72236 | 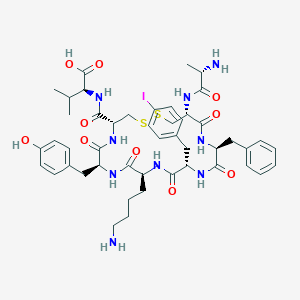 CHEMBL3104639 CHEMBL3104639 | C47H62IN9O10S2 | 1104.09 | 14 / 11 | 0.7 | No |
| 73299 | 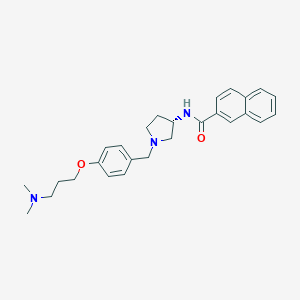 CHEMBL193462 CHEMBL193462 | C27H33N3O2 | 431.58 | 4 / 1 | 4.5 | Yes |
| 444613 | 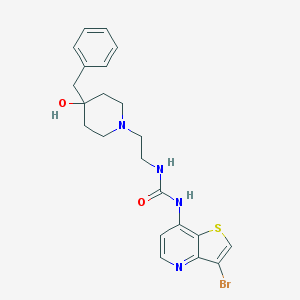 CHEMBL3358676 CHEMBL3358676 | C22H25BrN4O2S | 489.432 | 5 / 3 | 3.3 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218