

You can:
| Name | CHEMBL593435 |
|---|---|
| Molecular formula | C15H16ClN3O |
| IUPAC name | (2S)-2-(4-chlorophenyl)-3-methyl-N-pyridazin-3-ylbutanamide |
| Molecular weight | 289.763 |
| Hydrogen bond acceptor | 3 |
| Hydrogen bond donor | 1 |
| XlogP | 2.9 |
| Synonyms | BDBM50305954 (S)-2-(4-chlorophenyl)-3-methyl-N-(pyridazin-3-yl)butanamide |
| Inchi Key | DYKQXHLDYNKKKR-AWEZNQCLSA-N |
| Inchi ID | InChI=1S/C15H16ClN3O/c1-10(2)14(11-5-7-12(16)8-6-11)15(20)18-13-4-3-9-17-19-13/h3-10,14H,1-2H3,(H,18,19,20)/t14-/m0/s1 |
| PubChem CID | 46226161 |
| ChEMBL | CHEMBL593435 |
| IUPHAR | N/A |
| BindingDB | 50305954 |
| DrugBank | N/A |
Structure | 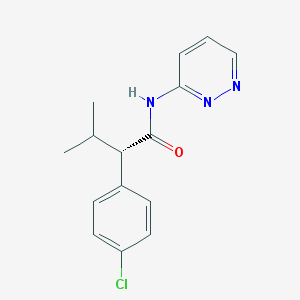 |
| Lipinski's druglikeness | This ligand satisfies Lipinski's rule of five. |
You can:
| GLASS ID | Name | UniProt | Gene | Species | Length |
|---|---|---|---|---|---|
| 72046 | Free fatty acid receptor 2 | O15552 | FFAR2 | Homo sapiens (Human) | 330 |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218