

| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| MAEENKILVTHFVLTGLTDHPGLQAPLFLVFLVIYLITLVGNLGLMALIWKDPHLHTPIYLFLGSLAFADACTSSSVTSKMLINFLSKNHMLSMAKCATQFYFFGSNATTECFLLVVMAYDRYVAICNPLLYPVVMSNSLCTQFIGISYFIGFLHSAIHVGLLFRLTFCRSNIIHYFYCEILQLFKISCTNPTVNILLIFIFSAFIQVFTFMTLIVSYSYILSAILKKKSEKGRSKAFSTCSAHLLSVSLFYGTLFFMYVSSRSGSAADQAKMYSLFYTIIIPLLNPFIYSLRNKEVIDALRRIMKK | |
| CCCCCCCCCSSSSSSCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCHHHHHHHHHHHHCCCCCCCCHHHHHHHHCCCCSSCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCCCCCCCCHHHHHHHCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCHHHCHHHHHHHHHHHHHHHHSSSSCCCCCCCCCCCCSSSSSSCCHHHCCCCHHHCCCCHHHHHHHHHHHCC | |
| 9998873220246865899935189999999999999999997999999508887674989986899984580024828898987248978848999999999999999999999999865145206210188016777999999999999999999999998453088999038441682888877157862767888889999999999999999999999980376647775521055889998999985451237378999878889889987210252233574505649899999998649 |
| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| MAEENKILVTHFVLTGLTDHPGLQAPLFLVFLVIYLITLVGNLGLMALIWKDPHLHTPIYLFLGSLAFADACTSSSVTSKMLINFLSKNHMLSMAKCATQFYFFGSNATTECFLLVVMAYDRYVAICNPLLYPVVMSNSLCTQFIGISYFIGFLHSAIHVGLLFRLTFCRSNIIHYFYCEILQLFKISCTNPTVNILLIFIFSAFIQVFTFMTLIVSYSYILSAILKKKSEKGRSKAFSTCSAHLLSVSLFYGTLFFMYVSSRSGSAADQAKMYSLFYTIIIPLLNPFIYSLRNKEVIDALRRIMKK | |
| 8466230200100010004326010000130333133123233200200100130000002102000000010000000200010014433020300000010000000000100000000000000211202030033000000210111012002000000010201442201000001210030002233110100011013113313300330331001000103046123101000000210010232011000000324423344200002103200330030000213302400331277 |
| Rank | PDB Hit | Iden1 | Iden2 | Cov. | Norm. Z-score | Download Align. | 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | | | | | | | | | | | | | | | | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sec.Str Seq | CCCCCCCCCSSSSSSCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCHHHHHHHHHHHHCCCCCCCCHHHHHHHHCCCCSSCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCCCCCCCCHHHHHHHCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCHHHCHHHHHHHHHHHHHHHHSSSSCCCCCCCCCCCCSSSSSSCCHHHCCCCHHHCCCCHHHHHHHHHHHCC MAEENKILVTHFVLTGLTDHPGLQAPLFLVFLVIYLITLVGNLGLMALIWKDPHLHTPIYLFLGSLAFADACTSSSVTSKMLINFLSKNHMLSMAKCATQFYFFGSNATTECFLLVVMAYDRYVAICNPLLYPVVMSNSLCTQFIGISYFIGFLHSAIHVGLLFRLTFCRSNIIHYFYCEILQLFKISCTNPTVNILLIFIFSAFIQVFTFMTLIVSYSYILSAILKKKSEKGRSKAFSTCSAHLLSVSLFYGTLFFMYVSSRSGSAADQAKMYSLFYTIIIPLLNPFIYSLRNKEVIDALRRIMKK | |||||||||||||||||||||||||
| 1 | 3emlA | 0.17 | 0.22 | 0.89 | 3.61 | Download | --------------------IMGSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAITIS---GFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGGCGEGQVACLFEDVVP-----------MNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAI-IVGLFALCWLPLHIINCFTFFCPDCSHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRS | |||||||||||||||||||
| 2 | 5tgzA | 0.21 | 0.23 | 0.89 | 2.03 | Download | -RGENFMDIECF----MVLNPSQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRCPSYHFIGSLAVADLLGSVIFVYSFIDFHVFHRK-DSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIAV------------------------LPLLGWNCEKLQSVDKTYLMFWIGVVSVLLLFIVYAYMYILWKAHSHAPDQARMELAKTLVLILVVLIICWGPLLAIMVYDVFGKMNKLIKTVFSMLCLLNSTVNPIIYALRSKDLRHAFRSM--- | |||||||||||||||||||
| 3 | 4iaqA | 0.18 | 0.22 | 0.87 | 2.10 | Download | ------------YIYQDSISLPWKVLLVMLLALITLATTLSNAFVIATVYRTRKLHTPANYLIASLAVTDLLVSILVMPISTMYTVTGRWTLGQVVCDFWLSSDITCCTASIWHLCVIALDRYWAITDAVEYSAKRTPKRAAVMIALVWVFSISISL---------PPCVV------------------NTDHILYTVYSTVGAF--YFPTLLLIALYGRIYVEARSQKYLLMAARERKATKTLGIILGAFIVCWLPFFIISLVMPIHLAIFDFFTWLGYLNSLINPIIYTMSNEDFKQAFHKLIRF | |||||||||||||||||||
| 4 | 4djh | 0.16 | 0.20 | 0.92 | 1.56 | Download | -------------------SPAIPVIITAVYSVVFVVGLVGNSLVMFVIIRYTKMKTATNIYIFNLALADALVTT-TMPFQSTVYLMNSWPFGDVLCKIVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKALDFRTPLKAKIINICIWLLSSSVGISAIVLGGTKVR---EDVDVIECSLQFPDD---DYSWWDLFMKICVFIFAFVIPVLIIIVCYTLMILRLKSVRLLRNLRRITRLVLVVVAVFVVCWTPIHIFILVEALGSAALSSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCFP | |||||||||||||||||||
| 5 | 4yay | 0.17 | 0.23 | 0.93 | 1.26 | Download | LKTTRNAYIQKYLILNSSDCNYIFVMIPTLYSIIFVVGIFGNSLVVIVIYFYMKLKTVASVFLLNLALADLCFLLTLPLWAVYTAMEYRWPFGNYLCKIASASVSFNLYASVFLLTCLSIDRYLAIVHPMKSRLRRTMLVAKVTCIIIWLLAGLASLPAIIHR-NVFF------IITVCAFHYE--------TLPIGLGLTKNILGFLFPFLIILTSYTLIWKALK------KNDDIFKIIMAIVLFFFFSWIPHQIFTFLDLGIDIVDTAMPITICIAYFNNCLNPLFYGFLGKKFKRYFLQLL-- | |||||||||||||||||||
| 6 | 3emlA | 0.18 | 0.22 | 0.89 | 3.79 | Download | ---------------------MGSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAIT--ISTGFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGGCGEGQVACLFEDVVP-----------MNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAIIVGLFALCWLPLHIINCF-TFFCPDCSHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRS | |||||||||||||||||||
| 7 | 4iaq | 0.20 | 0.22 | 0.85 | 1.71 | Download | ---------------------PWKVLLVMLLALITLATTLSNAFVIATVYRTRKLHTPANYLIASLAVTDLLVSILVMPISTMYTVTGRWTLGQVVCDFWLSSDITCCTASIWHLCVIALDRYWAITDAVEYSAKRTPKRAAVMIALVWVFSISISLPPFFW-RQASECVVN--------------------TDHILYTVYSTVGAFYFPTLLLIALYGRIYVEARSRIADARERKATKTLGIILGAFIVCWLPFFIISLVMPIH--L-AIFDFFTWLGYLNSLINPIIYTMSNEDFKQAFHKLIR- | |||||||||||||||||||
| 8 | 2z73A | 0.18 | 0.23 | 0.94 | 2.92 | Download | --EYNPSIVVHPHWREFDQVPAVYYSLGIFIGICGIIGCGGNGIVIYLFTKTKSLQTPANMFIINLAFSDFTFSLVNGPLMTISCFLKKWIFGFAACKVYGFIGGIFGFMSIMTMAMISIDRYNVIGRPMAASKKMSHRRAFIMIIFVWLWSVLWAIGPIFGWGAYTLEGVL----------------CSRDSTTRSNILCMFILGFFGPILIIFFCYFNIVMAQAGANAEMRLAKISIVIVSQFLLSWSPYAVVALLAQFGPLEWVTPYAAQLPVMFAKASAIHNPMIYSVSHPKFREAISQTFPW | |||||||||||||||||||
| 9 | 3emlA | 0.17 | 0.22 | 0.89 | 5.17 | Download | --------------------IMGSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAITIS---GFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLNCGQSQGCGEGQVACLFEDVVP-----------MNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAI-IVGLFALCWLPLHIINCFTFFCPDCSHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRS | |||||||||||||||||||
| 10 | 2ydoA | 0.19 | 0.21 | 0.92 | 5.91 | Download | -----------------------SSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSAAAADILVGVLAIPFAIA--ISTGFCAACHGCLFIACFVLVLTASSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGQPKEGKAHSQGCGEGQVACLFEDVVPMNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLKQMESTLQKEVHAAKSLAIIVGLFALCWLTFFCPDCSHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRS | |||||||||||||||||||
| ||||||||||||||||||||||||||
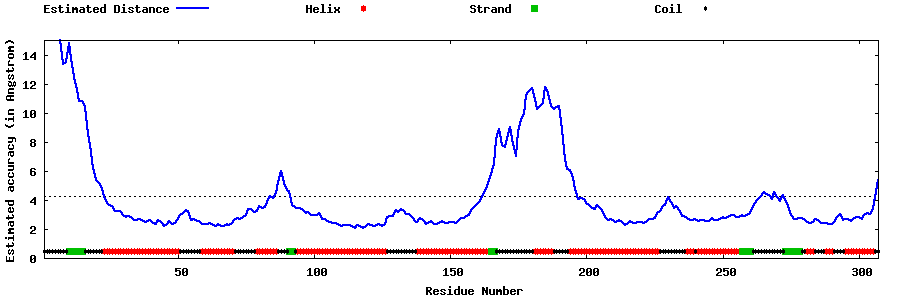
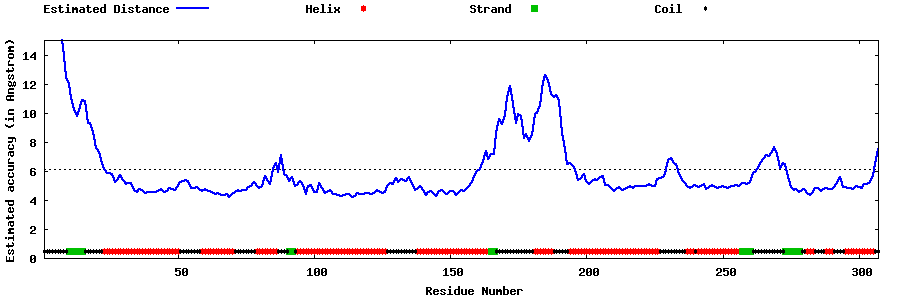
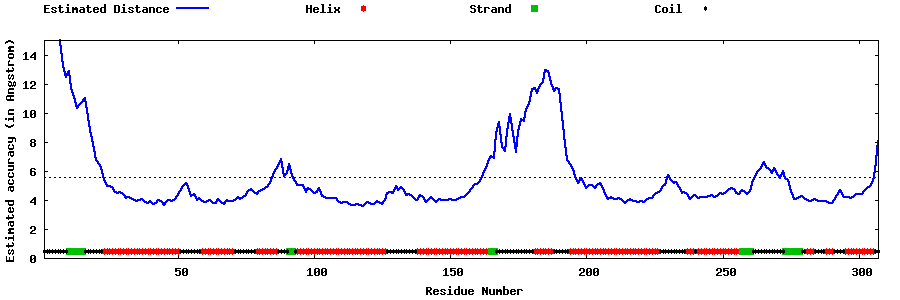
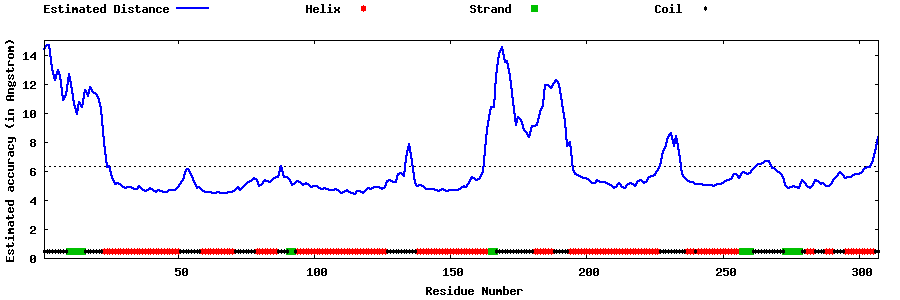
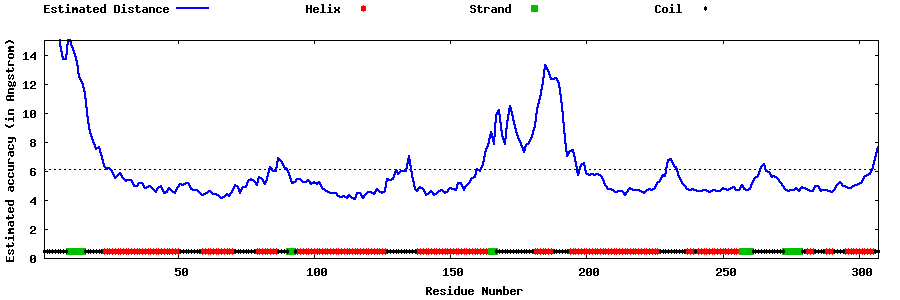
| Generated 3D models | Estimated local accuracy of models | ||
 | |||
|
|
|||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||