

| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| MEEAILLNQTSLVTYFRLRGLSVNHKARIAMFSMFLIFYVLTLIGNVLIVITIIYDHRLHTPMYFFLSNLSFIDVCHSTVTVPKMLRDVWSEEKLISFDACVTQMFFLHLFACTEIFLLTVMAYDRYVAICKPLQYMIVMNWKVCVLLAVALWTGGTIHSIALTSLTIKLPYCGPDEIDNFFCDVPQVIKLACIDTPYVLEILIVSNSGLISVVCFVVLVVSYAVILVSLRQQISKGKWKALSTCAAHLTVVTLFLGHCIFIYSRPSTSLPEDKAVSVFFTAVTPLLNPIIYTLRNEEMKSALNKLVGRKERKEEK | |
| CCHHHCCCCCCSSSSSSSSCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHSSSSSCCCCCCCCHHHHHHHHHHHHHHHHCCCHHHHHHHHHCCCCSSCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCHHCCCHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCCCCCSSCCCHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCHHHHHHHHHHHHHHHHHHCCSSSSSCCCCCCCCCCCSSHHHHHHHHHCCCCHHHCCCCHHHHHHHHHHHHCCCCCCCC | |
| 9223305578602377881699883168999999999999999878873552330799999789998759988664433461999998616996785899999999999999999999999986508873622026112687599999999999999999999999531899898837786428288788862681514564255675599999999999999999999513573123199888887999321552065468868899997631332456433430114743356199999999999702676679 |
| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| MEEAILLNQTSLVTYFRLRGLSVNHKARIAMFSMFLIFYVLTLIGNVLIVITIIYDHRLHTPMYFFLSNLSFIDVCHSTVTVPKMLRDVWSEEKLISFDACVTQMFFLHLFACTEIFLLTVMAYDRYVAICKPLQYMIVMNWKVCVLLAVALWTGGTIHSIALTSLTIKLPYCGPDEIDNFFCDVPQVIKLACIDTPYVLEILIVSNSGLISVVCFVVLVVSYAVILVSLRQQISKGKWKALSTCAAHLTVVTLFLGHCIFIYSRPSTSLPEDKAVSVFFTAVTPLLNPIIYTLRNEEMKSALNKLVGRKERKEEK | |
| 4754353644120000000000324400100013133323323332330000020033030000100230012001100000020000002532100050000000110331131020001003000000022110100013310000012013102301300131023030000122000100221002000120313010300230131023103313311210020013214612220010030021003023210100000033313301000031023103310300001044023003301343143688 |
| Rank | PDB Hit | Iden1 | Iden2 | Cov. | Norm. Z-score | Download Align. | 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | | | | | | | | | | | | | | | | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sec.Str Seq | CCHHHCCCCCCSSSSSSSSCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHSSSSSCCCCCCCCHHHHHHHHHHHHHHHHCCCHHHHHHHHHCCCCSSCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCHHCCCHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCCCCCSSCCCHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCHHHHHHHHHHHHHHHHHHCCSSSSSCCCCCCCCCCCSSHHHHHHHHHCCCCHHHCCCCHHHHHHHHHHHHCCCCCCCC MEEAILLNQTSLVTYFRLRGLSVNHKARIAMFSMFLIFYVLTLIGNVLIVITIIYDHRLHTPMYFFLSNLSFIDVCHSTVTVPKMLRDVWSEEKLISFDACVTQMFFLHLFACTEIFLLTVMAYDRYVAICKPLQYMIVMNWKVCVLLAVALWTGGTIHSIALTSLTIKLPYCGPDEIDNFFCDVPQVIKLACIDTPYVLEILIVSNSGLISVVCFVVLVVSYAVILVSLRQQISKGKWKALSTCAAHLTVVTLFLGHCIFIYSRPSTSLPEDKAVSVFFTAVTPLLNPIIYTLRNEEMKSALNKLVGRKERKEEK | |||||||||||||||||||||||||
| 1 | 3emlA | 0.19 | 0.21 | 0.87 | 3.29 | Download | ------------------------IMGSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAITIS---GFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGGCGEGQVACLFEDVVP------------MNYMVYFNFFACVLVPLLLMLGVYLRIFLAARQLVHAAKSLAIIVGLFALCWLPLH-IINCFTFFCPDCSHAWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVLRQ-- | |||||||||||||||||||
| 2 | 5tgzA | 0.19 | 0.23 | 0.88 | 2.31 | Download | -------ENFMDIECF----MVLNPSQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRCPSYHFIGSLAVADLLGSVIFVYSFIDFHVFHRK-DSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIAVL--------PLLGWNCEKL---------QSVCSDIFPHIDKTYLMFWIGVVSVLLLFIVYAYMYILWKAHSHAPDQARELAKTLVLILVVLIICWGPLLAIMVYDVFGKMNKFAFCSMLCLLNSTVNPIIYALRSKDLRHAFRSM---------- | |||||||||||||||||||
| 3 | 5tgzA | 0.18 | 0.23 | 0.85 | 2.11 | Download | --------------GGRGENFMDIPSQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRCPSYHFIGSLAVADLLGSVIFVYSFIDFHVFHRK-DSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIAVLPLLCEKLQSVCSD-----------------------IFPHIDKTYLMFWIGVVSVLLLFIVYAYMYILWKAHSHADIELAKTLVLILVVLIICWGPLLAIMVGKMNLIKTVFAFCSMLCLLNSTVNPIIYALRSKDLRHAFRSMF--------- | |||||||||||||||||||
| 4 | 4ib4 | 0.17 | 0.22 | 0.87 | 1.55 | Download | ---------------------EEQGNKLHWAALLILMVIIPTIGGNTLVILAVSLEKKLQYATNYFLMSLAVADLLVGLFVMPIALLTIMEAMWPLPLVLCPAWLFLDVLFSTASIWHLCAISVDRYIAIKKPIQANQYNSRATAFIKITVVWLISIGIAIPVPIKGIE------TNPNNITCVLTK----------ERFGDFMLFGSLAAFFTPLAIMIVTYFLTIHALQKKAADNEQRASKVLGIVFFLFLLMWCPFFITNITLVLCQQMLLEIFVWIGYVSSGVNPLVYTLFNKTFRDAFGRYITCNYR---- | |||||||||||||||||||
| 5 | 4yay | 0.16 | 0.21 | 0.90 | 1.23 | Download | AEQLKTT-RNAYIQKYLILNSSDCNYIFVMIPTLYSIIFVVGIFGNSLVVIVIYFYMKLKTVASVFLLNLALADLCFLLTLPLWAVYTAMEYRWPFGNYLCKIASASVSFNLYASVFLLTCLSIDRYLAIVHPMKSRLRRTMLVAKVTCIIIWLLAGLASLPAIIHRNVFF-------IITVCAFHYE------TL---PIGLGLTKNILGFLFPFLIILTSYTLIWKALK-----KNDDIFKIIMAIVLFFFFSWIPHQIFTFLDVLIVDTAMPITICIAYFNNCLNPLFYGFLGKKFKRYFLQLL--------- | |||||||||||||||||||
| 6 | 3emlA | 0.19 | 0.21 | 0.87 | 3.44 | Download | -------------------------MGSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAIT--ISTGFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGGCGEGQVACLFEDVVP------------MNYMVYFNFFACVLVPLLLMLGVYLRIFLAARQLVHAAKSLAIIVGLFALCWLPLHII-NCFTFFCPDCAPLWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVLRQ-- | |||||||||||||||||||
| 7 | 4iaq | 0.21 | 0.22 | 0.83 | 1.74 | Download | ------------------------LPWKVLLVMLLALITLATTLSNAFVIATVYRTRKLHTPANYLIASLAVTDLLVSILVMPISTMYTVTGRWTLGQVVCDFWLSSDITCCTASIWHLCVIALDRYWAITDAVEYSAKRTPKRAAVMIALVWVFSISISLPPFFW-RQAS----------ECVVNT-------D-H---ILYTVYSTVGAFYFPTLLLIALYGRIYVEARSRIADLERKATKTLGIILGAFIVCWLPFFIISLVMPIH-LAIFDFFTWLGYLNSLINPIIYTMSNEDFKQAFHKLIRFK------ | |||||||||||||||||||
| 8 | 4ea3A | 0.22 | 0.22 | 0.82 | 2.92 | Download | -----------------------PLGLKVTIVGLYLAVCVGGLLGNCLVMYVILRHTKMKTATNIYIFNLALADTLVLL-TLPFQGTDILLGFWPFGNALCKTVIAIDYYNMFTSTFTLTAMSVDRYVAICHPTSSKAQAVNVAIWALASVVGVPVAIMGSAQV----------EDEEIECLVEIP-------TPQDYWGPVFAICIFLFSFIVPVLVISVCYSLMIRRLRGVRLLSGSAVFVGCWTPVQVFVLAQGLG----VQPSSTAVAILRFCTALGYVNSCLNPILYAFLDENFKACFR------------ | |||||||||||||||||||
| 9 | 5tgzA | 0.18 | 0.23 | 0.89 | 4.64 | Download | ---GGRGENFMDIECFMVL----NPSQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRCPSYHFIGSLAVADLLGSVIFVYSFIDFHVFH-RKDSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIAVLPLLG---------NCEK---------LQSVCSDIFPHIDKTYLMFWIGVVSVLLLFIVYAYMYILWKAHSHAPDQAIELAKTLVLILVVLIICWGPLLAIMVYDVFGKMN-FAFCSMLCLLNSTVNPIIYALRSKDLRHAFRSMF--------- | |||||||||||||||||||
| 10 | 2ydoA | 0.17 | 0.20 | 0.91 | 5.31 | Download | ---------------------------SSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSAAAADILVGVLAIPFAIA--ISTGFCAACHGCLFIACFVLVLTASSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGQPKEGKAHSQGCGEGQVACLFEDVVP-MNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRLKQMESTLQKEVHAAKSLAIIVGLFALCCFTFFCPDCSHAWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVLRQQE | |||||||||||||||||||
| ||||||||||||||||||||||||||
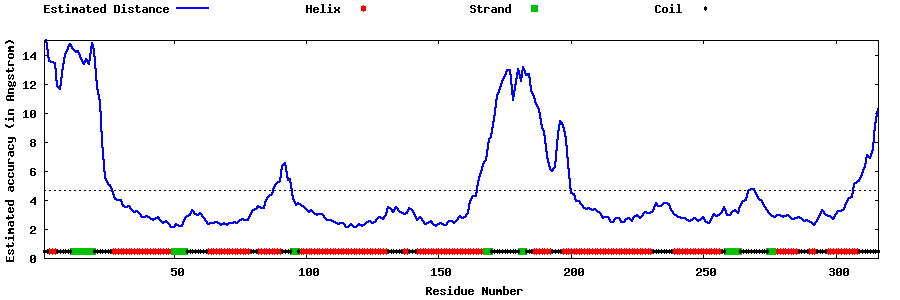
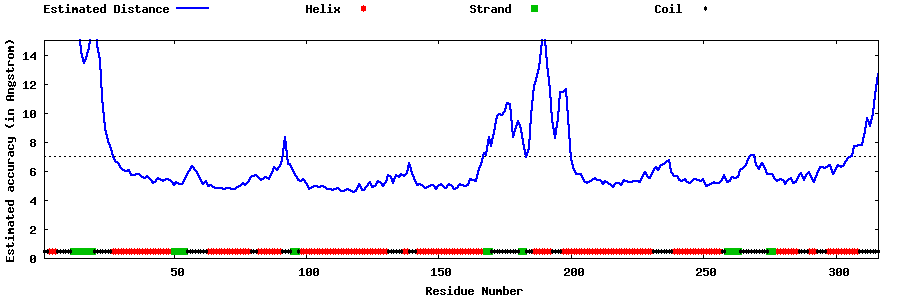
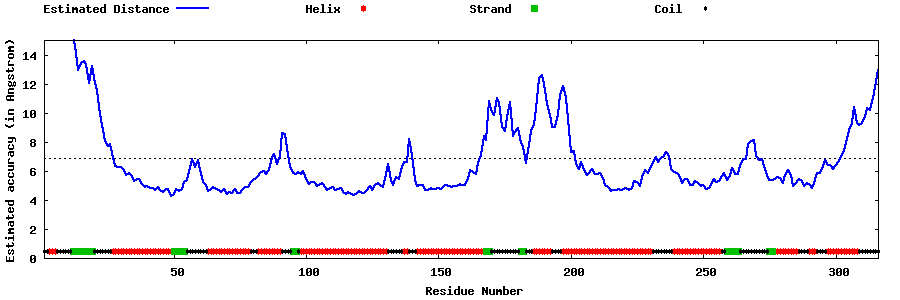
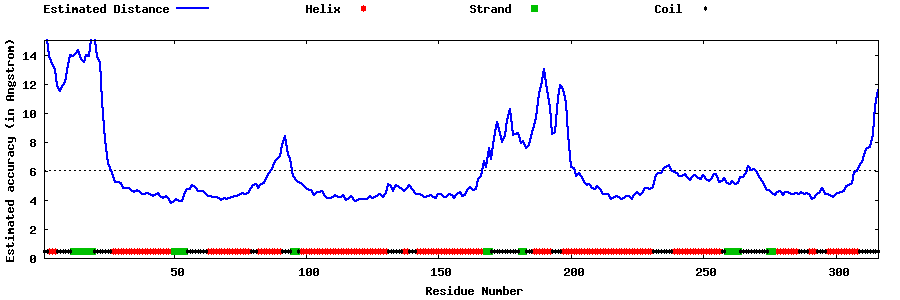
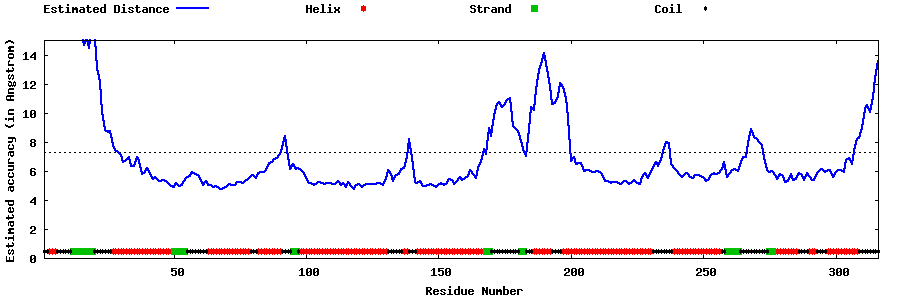
| Generated 3D models | Estimated local accuracy of models | ||
 | |||
|
|
|||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||