

| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 340 360 380 | |
| | | | | | | | | | | | | | | | | | | | | |
| MPSKSLSNLSVTTGANESGSVPEGWERDFLPASDGTTTELVIRCVIPSLYLLIITVGLLGNIMLVKIFITNSAMRSVPNIFISNLAAGDLLLLLTCVPVDASRYFFDEWMFGKVGCKLIPVIQLTSVGVSVFTLTALSADRYRAIVNPMDMQTSGALLRTCVKAMGIWVVSVLLAVPEAVFSEVARISSLDNSSFTACIPYPQTDELHPKIHSVLIFLVYFLIPLAIISIYYYHIAKTLIKSAHNLPGEYNEHTKKQMETRKRLAKIVLVFVGCFIFCWFPNHILYMYRSFNYNEIDPSLGHMIVTLVARVLSFGNSCVNPFALYLLSESFRRHFNSQLCCGRKSYQERGTSYLLSSSAVRMTSLKSNAKNMVTNSVLLNGHSMKQEMAL | |
| CCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSSSSCCCCCCCSSSSSCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCHHHHHHHHHHCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCC | |
| 999878876778998878888888765556656665405789999999999999999888987404302079889698999999999999999999999999997398889708887899999999999999999999971735004063210322676886527999999999989999834477034699727999616982669999999999999999999999999999999995405889862007789988877689999999999999998999999999997444477047999999999999999999999999981999999999981987878887888877788866268888999864466753689875513579 |
| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 340 360 380 | |
| | | | | | | | | | | | | | | | | | | | | |
| MPSKSLSNLSVTTGANESGSVPEGWERDFLPASDGTTTELVIRCVIPSLYLLIITVGLLGNIMLVKIFITNSAMRSVPNIFISNLAAGDLLLLLTCVPVDASRYFFDEWMFGKVGCKLIPVIQLTSVGVSVFTLTALSADRYRAIVNPMDMQTSGALLRTCVKAMGIWVVSVLLAVPEAVFSEVARISSLDNSSFTACIPYPQTDELHPKIHSVLIFLVYFLIPLAIISIYYYHIAKTLIKSAHNLPGEYNEHTKKQMETRKRLAKIVLVFVGCFIFCWFPNHILYMYRSFNYNEIDPSLGHMIVTLVARVLSFGNSCVNPFALYLLSESFRRHFNSQLCCGRKSYQERGTSYLLSSSAVRMTSLKSNAKNMVTNSVLLNGHSMKQEMAL | |
| 664732412223333231232343333322333432313100000002011200100130110000000003402100010010002012010001111100200133100020010000000100000000000000000010001002223332101000000000220010010000002023143274320000000033652031000001022101300000020000000101333453436445434433442110000000000000000212000000100223413321000001020110002000100101000142004002300202333346544444344433343434444543333333324433456358 |
| Rank | PDB Hit | Iden1 | Iden2 | Cov. | Norm. Z-score | Download Align. | 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 340 360 380 | | | | | | | | | | | | | | | | | | | | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sec.Str Seq | CCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSSSSCCCCCCCSSSSSCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCHHHHHHHHHHCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCC MPSKSLSNLSVTTGANESGSVPEGWERDFLPASDGTTTELVIRCVIPSLYLLIITVGLLGNIMLVKIFITNSAMRSVPNIFISNLAAGDLLLLLTCVPVDASRYFFDEWMFGKVGCKLIPVIQLTSVGVSVFTLTALSADRYRAIVNPMDMQTSGALLRTCVKAMGIWVVSVLLAVPEAVFSEVARISSLDNSSFTACIPYPQTDELHPKIHSVLIFLVYFLIPLAIISIYYYHIAKTLIKSAHNLPGEYNEHTKKQMETRKRLAKIVLVFVGCFIFCWFPNHILYMYRSFNYNEIDPSLGHMIVTLVARVLSFGNSCVNPFALYLLSESFRRHFNSQLCCGRKSYQERGTSYLLSSSAVRMTSLKSNAKNMVTNSVLLNGHSMKQEMAL | |||||||||||||||||||||||||
| 1 | 4n6hA | 0.25 | 0.24 | 0.77 | 3.12 | Download | ------------------------------SPGARSASSLALAIAITALYSAVCAVGLLGNVLVMFGIVRYTKMKTATNIYIFNLALADALATSTLPFQSAKYLM--TWPFGELLCKAVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKALDFRTPAKAKLINICIWVLASGVGVPIMVMAVTRPRD----GAVVCMLQFPSPSWYWDTVTKICVFLFAFVVPILIITVCYGLMLLRLRSVRLLSGS------KEKDRSLRRITRMVLVVVGAFVVCWAPIHIFVIVWTLVDID-RRDPLVVAALHLCIALGYANSSLNPVLYAFLDENFKRCFRQLCRKPCG---------------------------------------------- | |||||||||||||||||||
| 2 | 5gliA | 0.32 | 0.28 | 0.80 | 3.85 | Download | ---------------------------SPPPCQGPIEIKETFKYINTVVSCLVFVLGIIGNSTLLYIIYKNKCMRNGPNILIASLALGDLLHIVIAIPINVYKLLAEDWPFGAEMCKLVPFIQKASVGITVLSLCALSIDRYRAVASWSRIKGIGVPKWTAVEIVLIWVVSVVLAVPEAIGFDIITMDY-KGSYLRICLLHPVQKQFYATAKDWWLFSFYFCLPLAITAFFYTLMTCEMLRIDEGGGSGGDEAEKDHLKQRREVAKTVFCLVLVFALCWLPLHLARILKLTLYNQNDPLSFLLVLDYIGINMASLNSCANPIALYLVSKRFKNAFKSALCC------------------------------------------------- | |||||||||||||||||||
| 3 | 4n6hA | 0.25 | 0.24 | 0.77 | 3.64 | Download | ------------------------------SPGARSASSLALAIAITALYSAVCAVGLLGNVLVMFGIVRYTKMKTATNIYIFNLALADALATST-LPFQSAKYLMETWPFGELLCKAVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKALDFRTPAKAKLINICIWVLASGVGVPIMVMAVTRPRDG----AVVCMLQFPSPSWYWDTVTKICVFLFAFVVPILIITVCYGLMLLRLRSVRLLSGS------KEKDRSLRRITRMVLVVVGAFVVCWAPIHIFVIVWTLVDID-RRDPLVVAALHLCIALGYANSSLNPVLYAFLDENFKRCFRQLCRKPCG---------------------------------------------- | |||||||||||||||||||
| 4 | 4djh | 0.25 | 0.25 | 0.75 | 1.54 | Download | --------------------------------------SPAIPVIITAVYSVVFVVGLVGNSLVMFVIIRYTKMKTATNIYIFNLALADALVTT-TMPFQSTVYLMNSWPFGDVLCKIVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKALDFRTPLKAKIINICIWLLSSSVGISAIVLGGTKVR---EDVDVIECSLQFPDDDYWDLFMKICVFIFAFVIPVLIIIVCYTLMILRLKSVRLLSGNIFKTPYREKDRNLRRITRLVLVVVAVFVVCWTPIHIFILVEALGSA-------ALSSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCFP------------------------------------------------- | |||||||||||||||||||
| 5 | 4djh | 0.25 | 0.25 | 0.75 | 1.17 | Download | --------------------------------------SPAIPVIITAVYSVVFVVGLVGNSLVMFVIIRYTKMKTATNIYIFNLALADALVTT-TMPFQSTVYLMNSWPFGDVLCKIVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKALDFRTPLKAKIINICIWLLSSSVGISAIVLGGTKVRED---VDVIECSLQFPDDSWWDLFMKICVFIFAFVIPVLIIIVCYTLMILRLKSVRLLSGNIKTPNYREKDRNLRRITRLVLVVVAVFVVCWTPIHIFILVEALGSA---A----LSSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCFP------------------------------------------------- | |||||||||||||||||||
| 6 | 4n6hA | 0.25 | 0.24 | 0.77 | 3.29 | Download | --------------------------------GARSASSLALAIAITALYSAVCAVGLLGNVLVMFGIVRYTKMKTATNIYIFNLALADALA-TSTLPFQSAKYLMETWPFGELLCKAVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKALDFRTPAKAKLINICIWVLASGVGVPIMVMAVTRPRD----GAVVCMLQFPSPSWYWDTVTKICVFLFAFVVPILIITVCYGLMLLRLRSVRLLSG------SKEKDRSLRRITRMVLVVVGAFVVCWAPIHIFVIVWTLVDID-RRDPLVVAALHLCIALGYANSSLNPVLYAFLDENFKRCFRQLCRKPCG---------------------------------------------- | |||||||||||||||||||
| 7 | 4djh | 0.26 | 0.25 | 0.74 | 1.68 | Download | ----------------------------------------AIPVIITAVYSVVFVVGLVGNSLVMFVIIRYTKMKTATNIYIFNLALADALVTT-TMPFQSTVYLMNSWPFGDVLCKIVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKALDFRTPLKAKIINICIWLLSSSVGISAIVLGGTKVREDV--DVIECSLQFPDDDSWWDLFMKICVFIFAFVIPVLIIIVCYTLMILRLKSVRLLSGNITWDAYREKDRNLRRITRLVLVVVAVFVVCWTPIHIFILVEALGSA-------ALSSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCF-------------------------------------------------- | |||||||||||||||||||
| 8 | 4n6hA | 0.25 | 0.24 | 0.78 | 4.75 | Download | -----------------------------GSPGARSASSLALAIAITALYSAVCAVGLLGNVLVMFGIVRYTKMKTATNIYIFNLALADALATST-LPFQSAKYLMETWPFGELLCKAVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKALDFRTPAKAKLINICIWVLASGVGVPIMVMAVTRPRD----GAVVCMLQFPSPSWYWDTVTKICVFLFAFVVPILIITVCYGLMLLRLRSVRLLSGS------KEKDRSLRRITRMVLVVVGAFVVCWAPIHIFVIVWTLVDIDRRD-PLVVAALHLCIALGYANSSLNPVLYAFLDENFKRCFRQLCRKPCG---------------------------------------------- | |||||||||||||||||||
| 9 | 4n6hA | 0.25 | 0.24 | 0.77 | 3.19 | Download | ------------------------------SPGARSASSLALAIAITALYSAVCAVGLLGNVLVMFGIVRYTKMKTATNIYIFNLALADALATSTLPFQSAKYLM-ETWPFGELLCKAVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKALDFRTPAKAKLINICIWVLASGVGVPIMVMAVTRPR----DGAVVCMLQFPSPSWYWDTVTKICVFLFAFVVPILIITVCYGLMLLRLRSVRLLSGS------KEKDRSLRRITRMVLVVVGAFVVCWAPIHIFVIVWTLV-DIDRRDPLVVAALHLCIALGYANSSLNPVLYAFLDENFKRCFRQLCRKPCG---------------------------------------------- | |||||||||||||||||||
| 10 | 5c1mA | 0.24 | 0.21 | 0.76 | 3.73 | Download | ---------------------------GSHSLPQTGSPSMVTAITIMALYSIVCVVGLFGNFLVMYVIVRYTKMKTATNIYIFNLALADALAT-STLPFQSVNYLMGTWPFGNILCKIVISIDYYNMFTSIFTLCTMSVDRYIAVCHPVKALDFRTPRNAKIVNVCNWILSSAIGLPVMFMATTKYR----QGSIDCTLTFSHPTWYWENLLKICVFIFAFIMPVLIITVCYGLMILRLKSVRMLSGS------KEKDRNLRRITRMVLVVVAVFIVCWTPIHIYVIIKALI--TIPETTFQTVSWHFCIALGYTNSCLNPVLYAFLDENFKRCF------------------------------------------------------- | |||||||||||||||||||
| ||||||||||||||||||||||||||
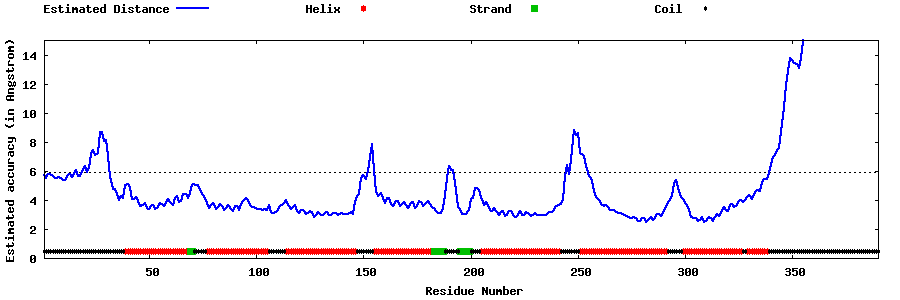
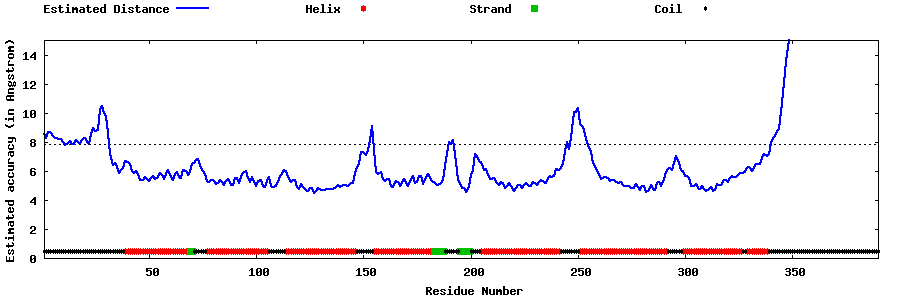
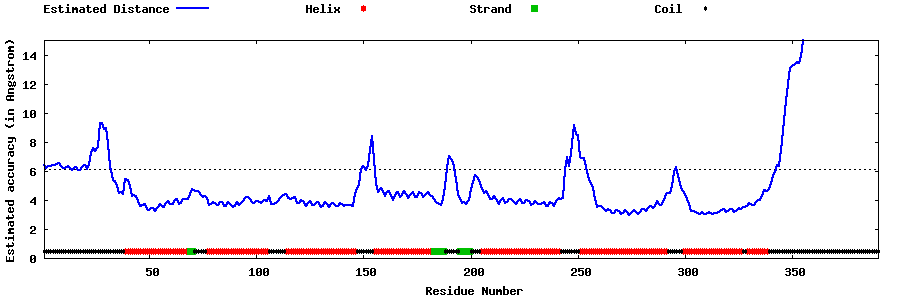
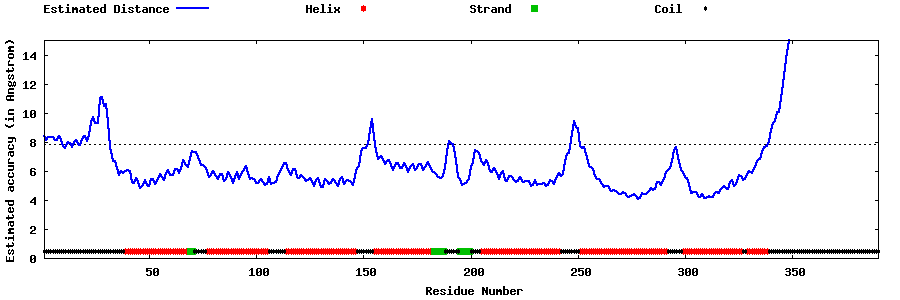
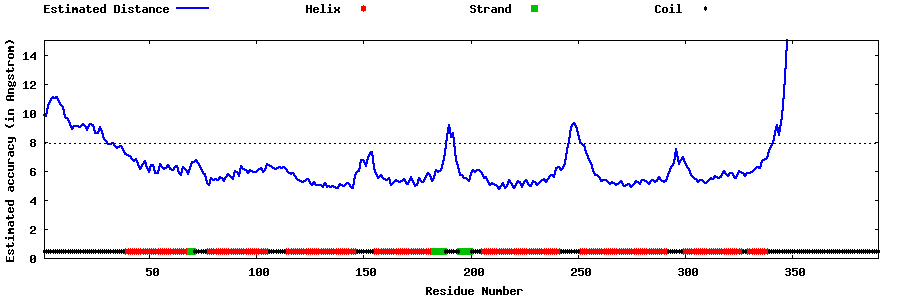
| Generated 3D models | Estimated local accuracy of models | ||
 | |||
|
|
|||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||