

| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 340 360 380 400 420 440 | |
| | | | | | | | | | | | | | | | | | | | | | | | |
| MVPEPGPTANSTPAWGAGPPSAPGGSGWVAAALCVVIALTAAANSLLIALICTQPALRNTSNFFLVSLFTSDLMVGLVVMPPAMLNALYGRWVLARGLCLLWTAFDVMCCSASILNLCLISLDRYLLILSPLRYKLRMTPLRALALVLGAWSLAALASFLPLLLGWHELGHARPPVPGQCRLLASLPFVLVASGLTFFLPSGAICFTYCRILLAARKQAVQVASLTTGMASQASETLQVPRTPRPGVESADSRRLATKHSRKALKASLTLGILLGMFFVTWLPFFVANIVQAVCDCISPGLFDVLTWLGYCNSTMNPIIYPLFMRDFKRALGRFLPCPRCPRERQASLASPSLRTSHSGPRPGLSLQQVLPLPLPPDSDSDSDAGSGGSSGLRLTAQLLLPGEATQDPPLPTRAAAAVNFFNIDPAEPELRPHPLGIPTN | |
| CCCCCCCCCCCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHSSSSSSSCCCCCCCCSSSSHHHHHHHHHHHHHHHHHHHHHHHCSSSCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCCSSSSSSCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCHHHHHHHHHHHHHHHHHHHHHHHCCCHHHHHHHHHHHCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCC | |
| 97889999887888888888882589999999999999999847870787882300058777487339999999999997999999982826477789999999999999999999999998524157761037865158999999999999999999999985660257766888998789307548642466899999999999999999999675202433344433332223344555555445443321111023456777888999999999999899999999977188867999999999999980624846457999999999984551468878776677542101468888865346768888998888888888887787765333687754678999987876546300367776557776899999 |
| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 340 360 380 400 420 440 | |
| | | | | | | | | | | | | | | | | | | | | | | | |
| MVPEPGPTANSTPAWGAGPPSAPGGSGWVAAALCVVIALTAAANSLLIALICTQPALRNTSNFFLVSLFTSDLMVGLVVMPPAMLNALYGRWVLARGLCLLWTAFDVMCCSASILNLCLISLDRYLLILSPLRYKLRMTPLRALALVLGAWSLAALASFLPLLLGWHELGHARPPVPGQCRLLASLPFVLVASGLTFFLPSGAICFTYCRILLAARKQAVQVASLTTGMASQASETLQVPRTPRPGVESADSRRLATKHSRKALKASLTLGILLGMFFVTWLPFFVANIVQAVCDCISPGLFDVLTWLGYCNSTMNPIIYPLFMRDFKRALGRFLPCPRCPRERQASLASPSLRTSHSGPRPGLSLQQVLPLPLPPDSDSDSDAGSGGSSGLRLTAQLLLPGEATQDPPLPTRAAAAVNFFNIDPAEPELRPHPLGIPTN | |
| 73023343222232444333222001000001112102202311200000001033013100000000010001000000201001001220110200010000000000000020000000000000020030333122300000000011100200110000101334354342331020102310011002200300000002000000100121023244244323322323243344433432233333223333342210010000000001000100000000200011011100000002002000100000000042003001200012214453344234332323334434333344334341454344444444444444424353334444465434445444323133064144424434253548 |
| Rank | PDB Hit | Iden1 | Iden2 | Cov. | Norm. Z-score | Download Align. | 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 340 360 380 400 420 440 | | | | | | | | | | | | | | | | | | | | | | | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sec.Str Seq | CCCCCCCCCCCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHSSSSSSSCCCCCCCCSSSSHHHHHHHHHHHHHHHHHHHHHHHCSSSCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCCSSSSSSCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCHHHHHHHHHHHHHHHHHHHHHHHCCCHHHHHHHHHHHCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCC MVPEPGPTANSTPAWGAGPPSAPGGSGWVAAALCVVIALTAAANSLLIALICTQPALRNTSNFFLVSLFTSDLMVGLVVMPPAMLNALYGRWVLARGLCLLWTAFDVMCCSASILNLCLISLDRYLLILSPLRYKLRMTPLRALALVLGAWSLAALASFLPLLLGWHELGHARPPVPGQCRLLASLPFVLVASGLTFFLPSGAICFTYCRILLAARKQAVQVASLTTGMASQASETLQVPRTPRPGVESADSRRLATKHSRKALKASLTLGILLGMFFVTWLPFFVANIVQAVCDCISPGLFDVLTWLGYCNSTMNPIIYPLFMRDFKRALGRFLPCPRCPRERQASLASPSLRTSHSGPRPGLSLQQVLPLPLPPDSDSDSDAGSGGSSGLRLTAQLLLPGEATQDPPLPTRAAAAVNFFNIDPAEPELRPHPLGIPTN | |||||||||||||||||||||||||
| 1 | 3sn6R | 0.31 | 0.22 | 0.64 | 2.32 | Download | ----------------------EVWVVGMGIVMSLIVLAIVFGNVLVITAIAKFERLQTVTNYFITSLACADLVMGLAVVPFGAAHILTKTWTFGNFWCEFWTSIDVLCVTASIETLCVIAVDRYFAITSPFKYQSLLTKNKARVIILMVWIVSGLTSFLPIQMHWYRQEAINYAEETCCDFFTNQAYAIASSIVSFYVPLVIMVFVYSRVFQEAKRQLQKIDKSEGR-------------------------------CLKEHKALKTLGIIMGTFTLCWLPFFIVNIVHVIQNLIRKEVYILLNWIGYVNSGFNPLIYC-RSPDFRIAFQELL-C------------------------------------------------------------------------------------------------------- | |||||||||||||||||||
| 2 | 2rh1A | 0.30 | 0.26 | 0.71 | 3.73 | Download | ---------------------DEVWVVGMGIVMSLIVLAIVFGNVLVITAIAKFERLQTVTNYFITSLACADLVMGLAVVPFGAAHILMKMWTFGNFWCEFWTSIDVLCVTASIETLCVIAVDRYFAITSPFKYQSLLTKNKARVIILMVWIVSGLTSFLPIQMHWYRATHQEAIEETCCDFFTNQAYAIASSIVSFYVPLVIMVFVYSRVFQEAKRQLNIFEMLRIDEGLRLKIYKDTEGYYTIGIGHRTGTWDAYKFCLKEHKALKTLGIIMGTFTLCWLPFFIVNIVHVIQDLIRKEVYILLNWIGYVNSGFNPLIYC-RSPDFRIAFQELL--------------------------------------------------------------------------------------------------------- | |||||||||||||||||||
| 3 | 3sn6R | 0.30 | 0.23 | 0.69 | 3.17 | Download | YNQTPNRAKRVITTFRTGTWDAEVWVVGMGIVMSLIVLAIVFGNVLVITAIAKFERLQTVTNYFITSLACADLVMGLAVVPFGAAHILTKTWTFGNFWCEFWTSIDVLCVTASIETLCVIAVDRYFAITSPFKYQSLLTKNKARVIILMVWIVSGLTSFLPIQMHWYRQAINCYAEETCCDFFTNQAYAIASSIVSFYVPLVIMVFVYSRVFQEAKRQLQKIDKSEGRC-------------------------------LKEHKALKTLGIIMGTFTLCWLPFFIVNIVHVIQNLIRKEVYILLNWIGYVNSGFNPLIYC-RSPDFRIAFQELLC-------------------------------------------------------------------------------------------------------- | |||||||||||||||||||
| 4 | 2ziy | 0.18 | 0.19 | 0.81 | 1.54 | Download | LRDNETWWYNPSIIVREFDQVPDAVYYSLGIFIGICGIIGCGGNGIVIYLFTKTKSLQTPANMFIINLAFSDFTFSLVNFPLMTISCFLKKWIFGFAACKVYGFIGGIFGFMSIMTMAMISIDRYNVIGRPMAASKKMSHRRAFIMIIFVWLWSVLWAIGP-IFGWGAYTLE--GVLCNCSFDYTRSNILCMFILGFFGPILIIFFCYFNIVMSVSNHEKEMAAMAKRL-------------------NAKELRKAQAGANAEMRLAKISIVIVSQFLLSWSPYAVVALLAQFGPLVTPYAAQLPVMFAKASAIHNPMIYSVSHPKFREAISQTFPWVLTCCQFDDKETE----DDKDA-------ETEIP----AGESSDA------------------------APSADAAQMKE----------------------- | |||||||||||||||||||
| 5 | 3d4s | 0.29 | 0.26 | 0.71 | 1.23 | Download | ------------------------WVVGMGIVMSLIVLAIVFGNVLVITAIAKFERLQTVTNYFITSLACADLVMGLAVVPFGAAHILMKMWTFGNFWCEFWTSIDVLCVTASIWTLCVIAVDRYFAITSPFKYQSLLTKNKARVIILMVWIVSGLTSFLPIQMHWYRATHQEYAEETCCDFFTNQAYAIASSIVSFYVPLVIMVFVYSRVFQEAKRQLNIFEMLRIDEGLRLDKAIGRNTNGVIEQQKSTGTWDAYKFCLKEHKALKTLGIIMGTFTLCWLPFFIVNIVHVIQDNIRKEVYILLNWIGYVNSGFNPLIYCR-SPDFRIAFQELLCL------------------------------------------------------------------------------------------------------- | |||||||||||||||||||
| 6 | 3sn6R | 0.31 | 0.22 | 0.64 | 2.79 | Download | -----------------------VWVVGMGIVMSLIVLAIVFGNVLVITAIAKFERLQTVTNYFITSLACADLVMGLAVVPFGAAHILTKTWTFGNFWCEFWTSIDVLCVTASIETLCVIAVDRYFAITSPFKYQSLLTKNKARVIILMVWIVSGLTSFLPIQMHWYRQAINCYAEETCCDFFTNQAYAIASSIVSFYVPLVIMVFVYSRVFQEAKRQLQKIDKSEG-------------------------------RCLKEHKALKTLGIIMGTFTLCWLPFFIVNIVHVIQDLIRKEVYILLNWIGYVNSGFNPLIYC-RSPDFRIAFQELLC-------------------------------------------------------------------------------------------------------- | |||||||||||||||||||
| 7 | 4djh | 0.20 | 0.19 | 0.71 | 1.74 | Download | ----------------------PAIPVIITAVYSVVFVVGLVGNSLVMFVIIRYTKMKTATNIYIFNLALADALVTT-TMPFQSTVYLMNSWPFGDVLCKIVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKALDFRTPLKAKIINICIWLLSSSVGISAIVLGGTKVREDVDVIECSLQFPDDLFMKICVFIFAFVIPVLIIIVCYTLMILRLKSVRLLSGNIFEMLRIDEGKAIGRNTQKRWNQTPTGTWDAYREKDRNLRRITRLVLVVVAVFVVCWTPIHIFILVEALGSA-ALSSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCF-------------------------------------------------------------------------------------------------------- | |||||||||||||||||||
| 8 | 2rh1A | 0.29 | 0.26 | 0.70 | 4.39 | Download | ---------------------DEVWVVGMGIVMSLIVLAIVFGNVLVITAIAKFERLQTVTNYFITSLACADLVMGLAVVPFGAAHILMKMWTFGNFWCEFWTSIDVLCVTASIETLCVIAVDRYFAITSPFKYQSLLTKNKARVIILMVWIVSGLTSFLPIQMHWYRATHEETCCDFF----TNQAYAIASSIVSFYVPLVIMVFVYSRVFQEAKRQL-NIFEMLRIDEGLRLKIYKDTEGKRVITTFRTGTWDAYKFCLKEHKALKTLGIIMGTFTLCWLPFFIVNIVHVIQDLIRKEVYILLNWIGYVNSGFNPLIYC-RSPDFRIAFQELLCL------------------------------------------------------------------------------------------------------- | |||||||||||||||||||
| 9 | 3sn6R | 0.31 | 0.22 | 0.64 | 2.61 | Download | ----------------------EVWVVGMGIVMSLIVLAIVFGNVLVITAIAKFERLQTVTNYFITSLACADLVMGLAVVPFGAAHILTKTWTFGNFWCEFWTSIDVLCVTASIETLCVIAVDRYFAITSPFKYQSLLTKNKARVIILMVWIVSGLTSFLPIQMHWYRQEAINCYEETCCDFFTNQAYAIASSIVSFYVPLVIMVFVYSRVFQEAKRQLQKIDKSEGR-------------------------------CLKEHKALKTLGIIMGTFTLCWLPFFIVNIVHVIQDLIRKEVYILLNWIGYVNSGFNPLIYC-RSPDFRIAFQELLC-------------------------------------------------------------------------------------------------------- | |||||||||||||||||||
| 10 | 3zpqA | 0.37 | 0.26 | 0.64 | 3.32 | Download | ----------------GAELLSQQWEAGMSLLMALVVLLIVAGNVLVIAAIGSTQRLQTLTNLFITSLACADLVVGLLVVPFGATLVVRGTWLWGSFLCELWTSLDVLCVTASIETLCVIAIDRYLAITSPFRYQSLMTRARAKVIICTVWAISALVSFLPIMMHWWRDEDPQYQDPGCCDFVTNRAYAIASSIISFYIPLLIMIFVALRVYREAKEQSR-------------------------------------VMLMREHKALKTLGIIMGVFTLCWLPFFLVNIVNVFNDLVPDWLFVAFNWLGYANSAMNPIIYC-RSPDFRKAFKRLLA-------------------------------------------------------------------------------------------------------- | |||||||||||||||||||
| ||||||||||||||||||||||||||
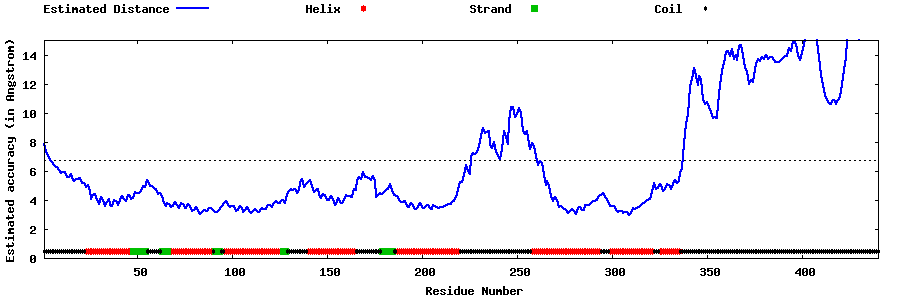
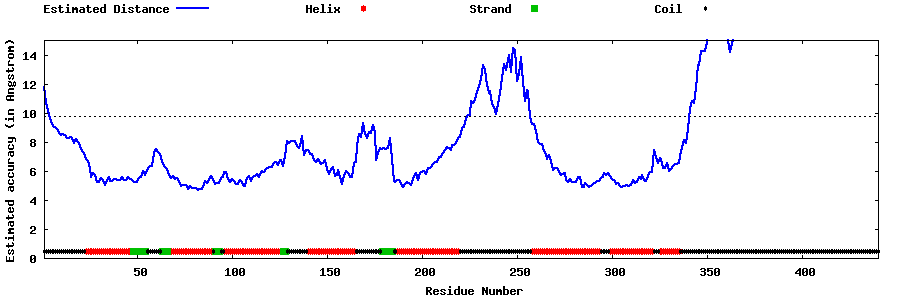
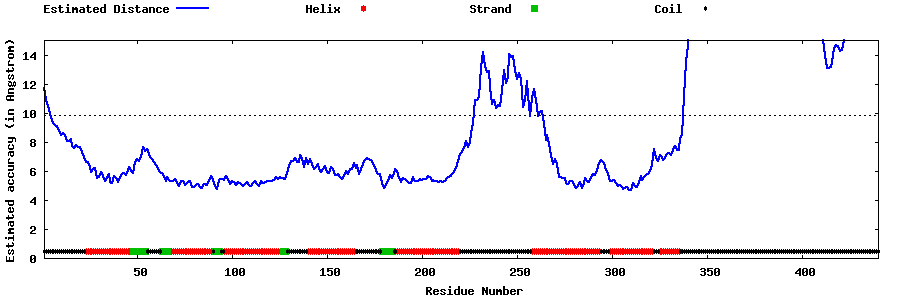
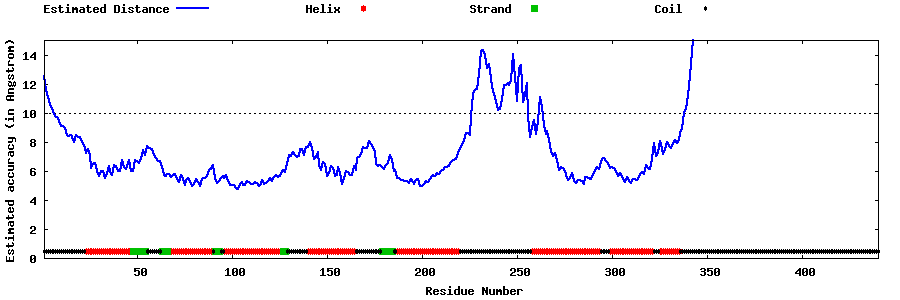
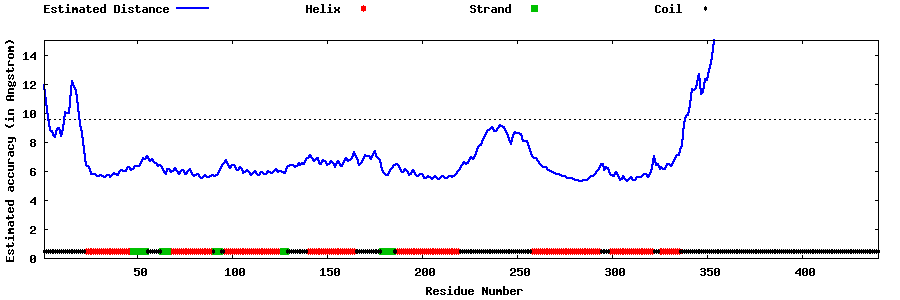
| Generated 3D models | Estimated local accuracy of models | ||
 | |||
|
|
|||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||