

| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| MDGDNQSENSQFLLLGISESPEQQRILFWMFLSMYLVTVLGNVLIILAISSDSHLHTPMYFFLANLSFTDLFFVTNTIPKMLVNFQSQNKAISYAGCLTQLYFLVSLVTLDNLILAVMAYDRYVATCCPLHYVTAMSPGLCVLLLSLCWGLSVLYGLLLTFLLTRVTFCGPREIHYLFCDMYILLWLACSNTHIIHTALIATGCFIFLTPLGFMTTSYVRIVRTILQMPSASKKYKTFSTCASHLGVVSLFYGTLAMVYLQPLHTYSMKDSVATVMYAVLTPMMNPFIYRLRNKDMHGAPGRVLWRPFQRPK | |
| CCCCCCCCCHHHHSSCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCHHHHHHHHHHHHHHHHCCCHHHHHHHHHCCCCSSCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCSSCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCSSCCCCCCHHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCCCSSSCCHHHHHHHHHHHCCHHSSSSCCCCCCCCCCCSSSSSCCCCCCCCCCHHHCCCCHHHHHHHHHHHHHHCCCCC | |
| 989888616334022699984277899999999999999999999999971787556299998889987654341578999899726896573888999999999999999999999987514412665437852377189999999999999999999999962278998944882678689986715353699999999999999997999999999999884014576876651046256888799985724336707898788877479984004362434564146628899999999853216899 |
| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| MDGDNQSENSQFLLLGISESPEQQRILFWMFLSMYLVTVLGNVLIILAISSDSHLHTPMYFFLANLSFTDLFFVTNTIPKMLVNFQSQNKAISYAGCLTQLYFLVSLVTLDNLILAVMAYDRYVATCCPLHYVTAMSPGLCVLLLSLCWGLSVLYGLLLTFLLTRVTFCGPREIHYLFCDMYILLWLACSNTHIIHTALIATGCFIFLTPLGFMTTSYVRIVRTILQMPSASKKYKTFSTCASHLGVVSLFYGTLAMVYLQPLHTYSMKDSVATVMYAVLTPMMNPFIYRLRNKDMHGAPGRVLWRPFQRPK | |
| 865723120020000101442621200001003012202332210000010023000000000001011011120031030020121433403020000011000200220100000003100000000020000000200000012002100200110000020001037240000001232003000110210100012213330332330032010100000010214623210000000000000011100100000002233432200000001211330010000203102300230033326368 |
| Rank | PDB Hit | Iden1 | Iden2 | Cov. | Norm. Z-score | Download Align. | 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | | | | | | | | | | | | | | | | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sec.Str Seq | CCCCCCCCCHHHHSSCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCHHHHHHHHHHHHHHHHCCCHHHHHHHHHCCCCSSCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCSSCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCSSCCCCCCHHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCCCSSSCCHHHHHHHHHHHCCHHSSSSCCCCCCCCCCCSSSSSCCCCCCCCCCHHHCCCCHHHHHHHHHHHHHHCCCCC MDGDNQSENSQFLLLGISESPEQQRILFWMFLSMYLVTVLGNVLIILAISSDSHLHTPMYFFLANLSFTDLFFVTNTIPKMLVNFQSQNKAISYAGCLTQLYFLVSLVTLDNLILAVMAYDRYVATCCPLHYVTAMSPGLCVLLLSLCWGLSVLYGLLLTFLLTRVTFCGPREIHYLFCDMYILLWLACSNTHIIHTALIATGCFIFLTPLGFMTTSYVRIVRTILQMPSASKKYKTFSTCASHLGVVSLFYGTLAMVYLQPLHTYSMKDSVATVMYAVLTPMMNPFIYRLRNKDMHGAPGRVLWRPFQRPK | |||||||||||||||||||||||||
| 1 | 3emlA | 0.21 | 0.23 | 0.89 | 3.59 | Download | --------------------IMGSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAITIST--GFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGGCGEGQVACLFEDVVP-----------MNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAIIVGLFALCWLPLHIINCFT-FFCPDCSHAPWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVLRQ- | |||||||||||||||||||
| 2 | 5tgzA | 0.22 | 0.23 | 0.90 | 2.19 | Download | -RGENFMDIECF----MVLNPSQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRCPSYHFIGSLAVADLLGSVIFVYSFIDF-HVFHRKDSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIAVLPLL-----------------GWNCEKLQSVCSDIFPHDKTYLMFWIGVVSVLLLFIVYAYMYILWKAHSHDQARMDIELAKTLVLILVVLIICWGPLLAIMVYDVFGKMNKLIKTVFMLCLLNSTVNPIIYALRSKDLRHAFRSMF-------- | |||||||||||||||||||
| 3 | 5tgzA | 0.21 | 0.23 | 0.87 | 2.07 | Download | GRGENFMD---------IENPSQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRCPSYHFIGSLAVADLLGSVIFVYSFIDFH-VFHRKDSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIAVLPLLG----WNCEKLIFPHID------------------KTYLMFWIGVVSVLLLFIVYAYMYILWKAHSHAQARMDIELAKTLVLILVVLIICWGPLLAIMVYDVFGKMNKLIKTVFAFCSMLCTVNPIIYALRSKDLRHAFRSMF-------- | |||||||||||||||||||
| 4 | 4ib4 | 0.19 | 0.23 | 0.88 | 1.54 | Download | -----------------EEQGNKLHWAALLILMVIIPTIGGNTLVILAVSLEKKLQYATNYFLMSLAVADLLVGLFVMPIALLTIMFAMWPLPLVLCPAWLFLDVLFSTASIWHLCAISVDRYIAIKKPIQANQYNSRATAFIKITVVWLISIGIAIPVPIK--GIE----TNPNNITC----------VLTKEFGDFMLFGSLAAFFTPLAIMIVTYFLTIHALQKKAADSNEQRASKVLGIVFFLFLLMWCPFFITNILVLCDSNTQLLEIFVWIGYVSSGVNPLVYTLFNKTFRDAFGRYITCNYR--- | |||||||||||||||||||
| 5 | 4yay | 0.17 | 0.24 | 0.90 | 1.24 | Download | KTTRN-AYIQKYLILNSSDCPYIFVMIPTLYSIIFVVGIFGNSLVVIVIYFYMKLKTVASVFLLNLALADLCFLLTLPLWAVYTAMEYRWPFGNYLCKIASASVSFNLYASVFLLTCLSIDRYLAIVHPMKSRLRRTMLVAKVTCIIIWLLAGLASLPAIIHR-NVFF------IITVCAFHYE--------TLPIGLGLTKNILGFLFPFLIILTSYTLIWKALK------KNDDIFKIIMAIVLFFFFSWIPHQIFTFLDVGIRDVTAMPITICIAYFNNCLNPLFYGFLGKKFKRYFLQLL-------- | |||||||||||||||||||
| 6 | 3emlA | 0.21 | 0.23 | 0.88 | 3.83 | Download | ---------------------MGSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAIT--ISTGFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGGCGEGQVACLFEDVVP-----------MNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAIIVGLFALCWLPLHIINCF-TFFCPDCSHAPWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVLRQ- | |||||||||||||||||||
| 7 | 4iaq | 0.20 | 0.24 | 0.85 | 1.73 | Download | -------------------SLPWKVLLVMLLALITLATTLSNAFVIATVYRTRKLHTPANYLIASLAVTDLLVSILVMPISTMYTVTGRWTLGQVVCDFWLSSDITCCTASIWHLCVIALDRYWAITDAVEYSAKRTPKRAAVMIALVWVFSISISLPPFFW-RQASECVVN----------T-------DH---ILYTVYSTVGAFYFPTLLLIALYGRIYVEARSRIADARERKATKTLGIILGAFIVCWLPFFIISLVMPIH-L-AIFDFFTWLGYLNSLINPIIYTMSNEDFKQAFHKLIRFK----- | |||||||||||||||||||
| 8 | 2z73A | 0.19 | 0.22 | 0.94 | 2.94 | Download | -ETYNPSIVVHPHWREFDQPDAVYYSLGIFIGICGIIGCGGNGIVIYLFTKTKSLQTPANMFIINLAFSDFTFSLVNGFPLMTISCFLKKIFGFAACKVYGFIGGIFGFMSIMTMAMISIDRYNVIGRPMAASKKMSHRRAFIMIIFVWLWSVLWAIGPIFG------WGAYTLEGVLCD---------STTRSNILCMFILG---FFGPILIIFFCYFNIVMAQAGANAEMRLAKISIVIVSQFLLSWSPYAVVALLAFGPLEWVTPYAAQLPVMFAKASAIHNPMIYSVSHPKFREAISQTFPWVLTCCQ | |||||||||||||||||||
| 9 | 3emlA | 0.21 | 0.23 | 0.88 | 4.94 | Download | --------------------IMGSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAITIST--GFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLNCGQSQGCGEGQVACLFEDVVP-----------MNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAIIVGLFALCWLPLHI-INCFTFFCPDCSHAPL-MYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVLRQ- | |||||||||||||||||||
| 10 | 2ydoA | 0.17 | 0.24 | 0.92 | 5.72 | Download | -----------------S------SVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSAAAADILVGVLAIPFAIA--ISTGFCAACHGCLFIACFVLVLTASSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGQPKEGKAHSQGCGEGQVACLFEDVVPMNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLKQMESTLQKEVHAAKSLAIIVGLFAINCFTFFCPDCSHAPWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVLRQQ | |||||||||||||||||||
| ||||||||||||||||||||||||||
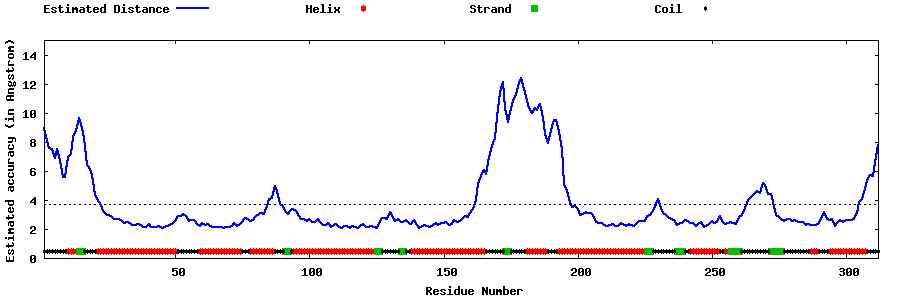
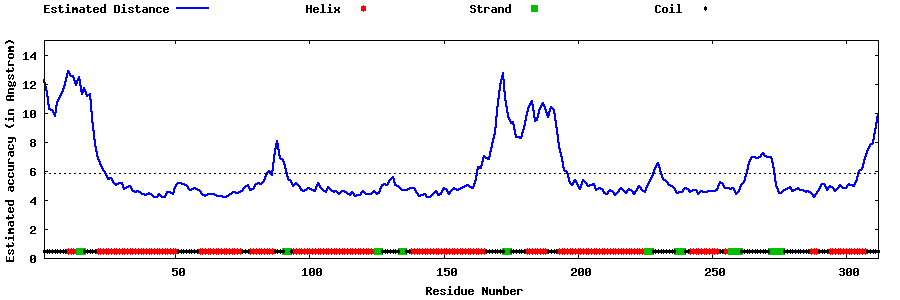
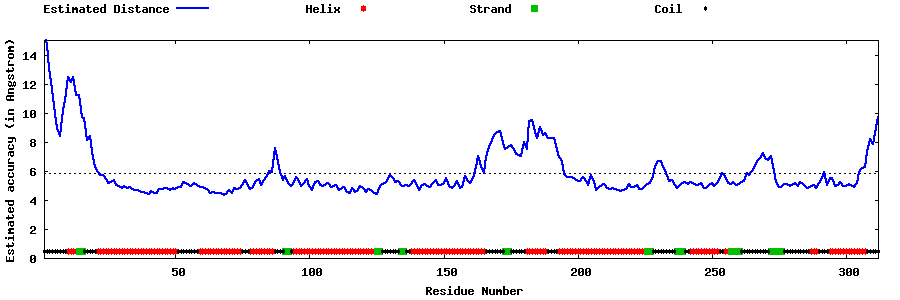
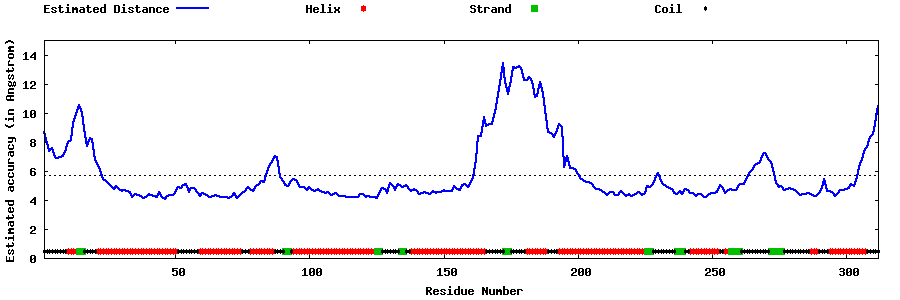
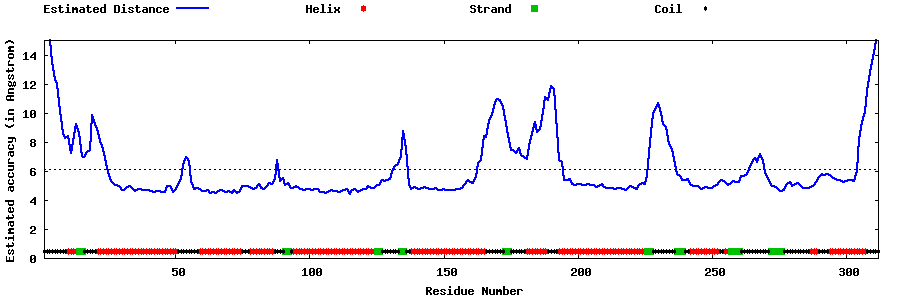
| Generated 3D models | Estimated local accuracy of models | ||
 | |||
|
|
|||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||