

| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 | |
| | | | | | | | | | | | | | | | | | |
| MLTLTRIRTVSYEVRSTFLFISVLEFAVGFLTNAFVFLVNFWDVVKRQALSNSDCVLLCLSISRLFLHGLLFLSAIQLTHFQKLSEPLNHSYQAIIMLWMIANQANLWLAACLSLLYCSKLIRFSHTFLICLASWVSRKISQMLLGIILCSCICTVLCVWCFFSRPHFTVTTVLFMNNNTRLNWQIKDLNLFYSFLFCYLWSVPPFLLFLVSSGMLTVSLGRHMRTMKVYTRNSRDPSLEAHIKALKSLVSFFCFFVISSCAAFISVPLLILWRDKIGVMVCVGIMAACPSGHAAILISGNAKLRRAVMTILLWAQSSLKVRADHKADSRTLC | |
| CCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCHHHHHHHHHHHHHHHHSSHHHHHHHHHHHCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSSCCCCCHHHHHHHHHHHCHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCSSSSSCCCCSSSSSSSCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCHHHHHHHHHHHHHHHHHHHHHHHCCCHHHHHHHHHHHHHHCCCCCCCCCCCCCCCCCC | |
| 998886101069999999999999999999999999999999999369898788999999999888365089979999967352555443579999999999399999999999998456368896699999987448276999999999999999999984146551035553169659998741168999999999999999999999999999999999999861878999999809999999999999999999999999999998713566999999999999789867998608739999999999860411468999999877789 |
| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 | |
| | | | | | | | | | | | | | | | | | |
| MLTLTRIRTVSYEVRSTFLFISVLEFAVGFLTNAFVFLVNFWDVVKRQALSNSDCVLLCLSISRLFLHGLLFLSAIQLTHFQKLSEPLNHSYQAIIMLWMIANQANLWLAACLSLLYCSKLIRFSHTFLICLASWVSRKISQMLLGIILCSCICTVLCVWCFFSRPHFTVTTVLFMNNNTRLNWQIKDLNLFYSFLFCYLWSVPPFLLFLVSSGMLTVSLGRHMRTMKVYTRNSRDPSLEAHIKALKSLVSFFCFFVISSCAAFISVPLLILWRDKIGVMVCVGIMAACPSGHAAILISGNAKLRRAVMTILLWAQSSLKVRADHKADSRTLC | |
| 735434344032102220132133123313311220021001000453303101100000011112002113332210000021323331000000010133123001000000000000002231100000133043200110132223233111110121233332323343334321223142332220211123333333233213313310110022023203432544411314002200100020222133333212310012233443001000112002100100101013143013001200220202032447463445443 |
| Rank | PDB Hit | Iden1 | Iden2 | Cov. | Norm. Z-score | Download Align. | 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 | | | | | | | | | | | | | | | | | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sec.Str Seq | CCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCHHHHHHHHHHHHHHHHSSHHHHHHHHHHHCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSSCCCCCHHHHHHHHHHHCHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCSSSSSCCCCSSSSSSSCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCHHHHHHHHHHHHHHHHHHHHHHHCCCHHHHHHHHHHHHHHCCCCCCCCCCCCCCCCCC MLTLTRIRTVSYEVRSTFLFISVLEFAVGFLTNAFVFLVNFWDVVKRQALSNSDCVLLCLSISRLFLHGLLFLSAIQLTHFQKLSEPLNHSYQAIIMLWMIANQANLWLAACLSLLYCSKLIRFSHTFLICLASWVSRKISQMLLGIILCSCICTVLCVWCFFSRPHFTVTTVLFMNNNTRLNWQIKDLNLFYSFLFCYLWSVPPFLLFLVSSGMLTVSLGRHMRTMKVYTRNSRDPSLEAHIKALKSLVSFFCFFVISSCAAFISVPLLILWRDKIGVMVCVGIMAACPSGHAAILISGNAKLRRAVMTILLWAQSSLKVRADHKADSRTLC | |||||||||||||||||||||||||
| 1 | 4grvA | 0.11 | 0.16 | 0.86 | 1.31 | Download | --NSDLDVNTDIYSKVLVTAIYLALFVVGTVGNSVTLFTLA----RKSLQSTVHYHLGSLALSDLLILLLAMPVELYNFIWVHHPWAGDAGCRGYYFLRDACTYATALNVASLSVARYLAIC---HPFKAK--LMSRSRTKKFISAIWLASALLAIPMLFTMGLQNRSADGT--HPGGLVCTPIVDTATVKVVIQVNTFMSFLFPMLVISILNTVIANKLTVM--------SGSVQALRHGVLVARAVVIAFVVCWLPYHVRRLMFCYISDEQWTHYFYMLTNALAYASSAINPILYNLVSANFRQV-------------------------- | |||||||||||||||||||
| 2 | 5tjvA | 0.12 | 0.19 | 0.83 | 2.82 | Download | ---------LNPSQQLAIAVLSLTLGTFTVLENLLVLCVILHS--RSLRCRPSYHFIGSLAVADLLGSVIFVYSFIDFHVF--HRKDSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLA----YKRIVTRPKAVVAFCLMWTIAIVIAVLPLLGWNCEKLQSVCSDIFPHID------------ETYLMFWIGVTSVLLLFIVYAYMYILWKAGKRAMSFSD--------QARMDIRLAKTLVLILVVLIICWGPLLAIMVYDVFGKMKTVFAFCSMLCLLNSTVNPIIYALRSKDLRHAFRSMF--------------------- | |||||||||||||||||||
| 3 | 4n6hA | 0.11 | 0.22 | 0.88 | 2.26 | Download | --SPGARSASSLALAIAITALYSAVCAVGLLGNVLVMFGIVRY--TKMK-TATNIYIFNLALADALATST--LPFQSAKYLMETWPFGELLCKAVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKA----LDFRTPAKAKLINICIWVLASGVGVPIMVMAVTRPRDG-AVVCMLQ------FPSPSWYWDTVTKICVFLFVVPILIITVCYGLMLLRLRSV------RLLSGSKEKDRSLRRITRMVLVVVGAFVVCWAPIHIFVIVWTLVDPLVVAALHLCIALGYSSLNPVLYAFLDENFKRCFRQLCRKPCG---------------- | |||||||||||||||||||
| 4 | 4djh | 0.10 | 0.22 | 0.87 | 1.56 | Download | ----------SPAIPVIITAVYSVVFVVGLVGNSLVMFVIIR---YTKMKTATNIYIFNLALADALVTTTMPFQST-VYLMNSWPF-GDVLCKIVLSIDYYNMFTSIFTLTMMSVDRYIAV---CHPVKAL-DFRTPLKAKIINICIWLLSSSVGISAIVLGGTKVR-E--DVDVECSLQFPDDDYSDLFMKI--CVFIFAFVIPVLIIIVCYTLMILRLKSVRLLSGNIFTPAYREKDRNLRRITRLVLVVVAVFVVCWTPIHIFILVEALGAALSSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCFP------------------- | |||||||||||||||||||
| 5 | 5glh | 0.09 | 0.16 | 0.86 | 1.17 | Download | -PPCQGPIEIKETFKYINTVVSCLVFVLGIIGNSTLLYIIYK--------NGPNILIASLALGDLLHIVIAIPINVYKLL-AEDWPFGAEMCKLVPFIQKASVGITVLSLCALSIDRYRAVAS---W-----------WTAVEIVLIWVVSVVLAVPEAIGFDIITMD-Y--KGSYLRICLLHPVQKAFMQFYDWWLFSFYFCLPLAITAFFYTLMTCEMLRKNEGLRLTWDAYLNDHLKQRREVAKTVFCLVLVFALCWLPLHLARILKLTLYNLVLDYIGINMASLNSCANPIALYLVSKRFKNAFKSAL--------------------- | |||||||||||||||||||
| 6 | 4djhA | 0.09 | 0.20 | 0.84 | 1.46 | Download | -----------PAIPVIITAVYSVVFVVGLVGNSLVMFVIIRY---TKMKTATNIYIFNLALADALVTTTMPFQSTVYLM--NSWPFGDVLCKIVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKALDFRTPLKAKIINICIWLLSSSVGISAIVLGGTKVREDVDVIECSLQFPDD------DYSWWDLFMKICVFIFAFVIPVLIIIVCYTLMILRLKSVRLLSD-----------RNLRRITRLVLVVVAVFVVCWTPIHIFILVEALGSALSSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCFP------------------- | |||||||||||||||||||
| 7 | 4ib4 | 0.13 | 0.17 | 0.85 | 1.71 | Download | ---------------HWAALLILMVIIPTIGGNTLVILAVSL---EKKLQYATNYFLMSLAVADLLVGLFVMPIALLTIMFEAMWPLPLVLCPAWLFLDVLFSTASIWHLCAISVDRYIAIKKPIQANQYNSRATAFIKITVVWLISIGIAIPVPI---KGIETNP---NNITCVLTK------ER---FGDFMLFGSLAAFFTPLAIMIVTYFLTIHALQKKAADLEDNYIQKYLQTISNEQRASKVLGIVFFLFLLMWCPFFITNITLVLCDLQMLLEIFVWIGYVSSGVNPLVYTLFNKTFRDAFGRYITCN------------------ | |||||||||||||||||||
| 8 | 4n6hA | 0.11 | 0.22 | 0.88 | 2.81 | Download | -GSPGARSASSLALAIAITALYSAVCAVGLLGNVLVMFGIVRY---TKMKTATNIYIFNLALADALA--TSTLPFQSAKYLMETWPFGELLCKAVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVK----ALDFRTPAKAKLINICIWVLASGVGVPIMVM--------AVTRPRDGAVVCMLQPSWYWDTVTKICVFLFAFVVPILIITVCYGLMLLRLRSV------RLLSGSKEKDRSLRRITRMVLVVVGAFVVCWAPIHIFVIVWTLVDIVAALHLCIALGYANSSLNPVLYAFLDENFKRCFRQLCRK---PCG------------- | |||||||||||||||||||
| 9 | 4djhA | 0.11 | 0.20 | 0.83 | 1.46 | Download | ----------SPAIPVIITAVYSVVFVVGLVGNSLVMFVIIR----TKMKTATNIYIFNLALADALV--TTTMPFQSTVYLMNSWPFGDVLCKIVLSIDYYNMFTSIFTLTMMSVDRYIAVC---HPVKA-LDFRTPLKAKIINICIWLLSSSVGISAIVLKVREDVDVIECSLQFPDD-----DYSWWDLFMKICVFIFAFVIPVLIIIVCYTLMILRLKSVRLL------------DRNLRRITRLVLVVVAVFVVCWTPIHIFILVEALGSALSSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCFP------------------- | |||||||||||||||||||
| 10 | 5c1mA | 0.10 | 0.15 | 0.86 | 1.37 | Download | SHSLPQTGSPSMVTAITIMALYSIVCVVGLFGNFLVMYVIVR---YTKMKTATNIYIFNLALADALA--TSTLPFQSVNYLMGTWPFGNILCKIVISIDYYNMFTSIFTLCTMSVDRYIAVC---HPVK-ALDFRTPRNAKIVNVCNWILSSAIGLPVMFMATTKYRQGSIDCTLTFSHPTW-----YWENLLKICVFIFAFIMPVLIITVCYGLMILRLKS------VRMLSGSKEKDRNLRRITRMVLVVVAVFIVCWTPIHIYVIIKALITQTVSWHFCIALGYTNSCLNPVLYAFLDENFKRCF------------------------- | |||||||||||||||||||
| ||||||||||||||||||||||||||
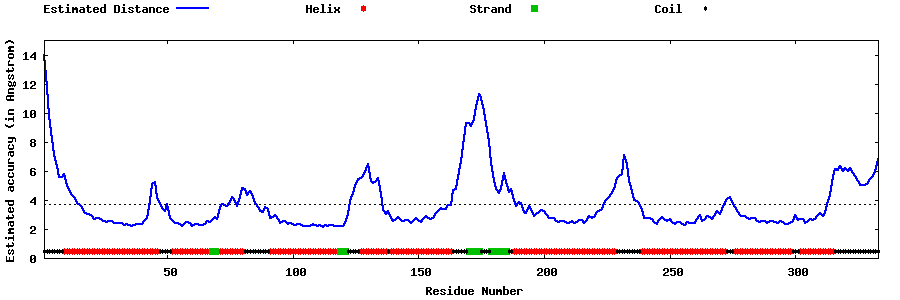
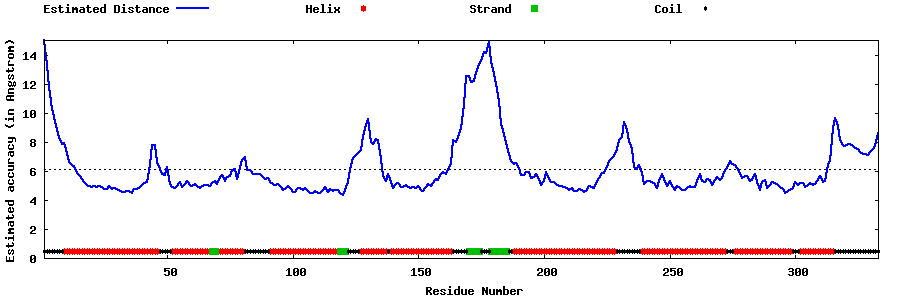
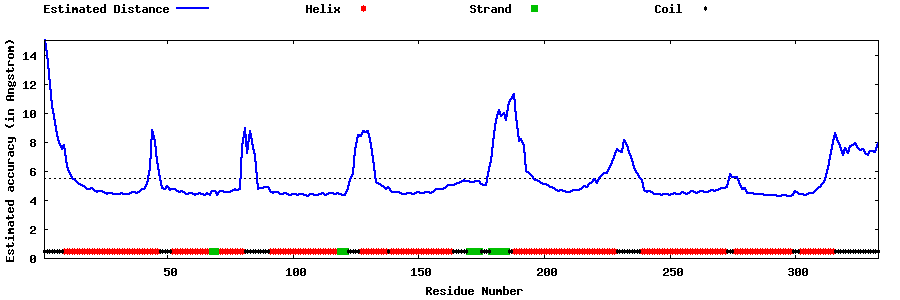
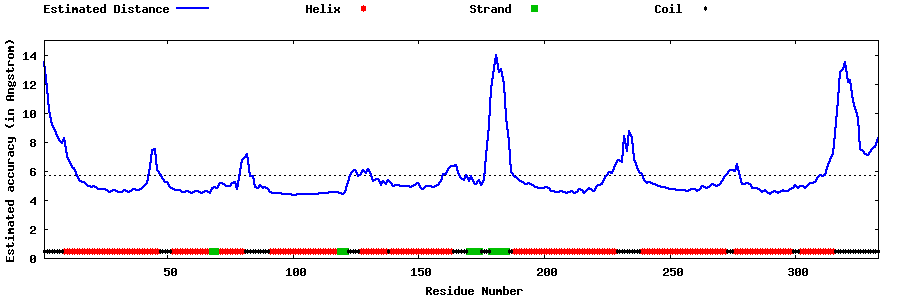
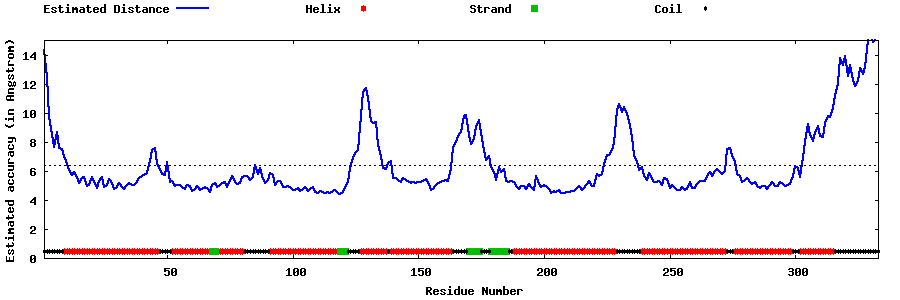
| Generated 3D models | Estimated local accuracy of models | ||
 | |||
|
|
|||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||