

| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 340 360 | |
| | | | | | | | | | | | | | | | | | | | |
| MANYSHAADNILQNLSPLTAFLKLTSLGFIIGVSVVGNLLISILLVKDKTLHRAPYYFLLDLCCSDILRSAICFPFVFNSVKNGSTWTYGTLTCKVIAFLGVLSCFHTAFMLFCISVTRYLAIAHHRFYTKRLTFWTCLAVICMVWTLSVAMAFPPVLDVGTYSFIREEDQCTFQHRSFRANDSLGFMLLLALILLATQLVYLKLIFFVHDRRKMKPVQFVAAVSQNWTFHGPGASGQAAANWLAGFGRGPTPPTLLGIRQNANTTGRRRLLVLDEFKMEKRISRMFYIMTFLFLTLWGPYLVACYWRVFARGPVVPGGFLTAAVWMSFAQAGINPFVCIFSNRELRRCFSTTLLYCRKSRLPREPYCVI | |
| CCCCCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCSSCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCSSSCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCHHHHHHHHHHHCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHCCHHHHHHHHHHHCCCCCCCCCCCCCCCC | |
| 9888899887688888399999999999999999998898522035566666544899999999999999999779999998289700771078789999999999999999999999999902702045345689999999999999999999999815864467888726604776320352011010006988999999999999999998550232101002322356676654332234455567776543334444332111002455557778878865568999999986419999999986377668578999999999999848489998458999999999956546688789988329 |
| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 340 360 | |
| | | | | | | | | | | | | | | | | | | | |
| MANYSHAADNILQNLSPLTAFLKLTSLGFIIGVSVVGNLLISILLVKDKTLHRAPYYFLLDLCCSDILRSAICFPFVFNSVKNGSTWTYGTLTCKVIAFLGVLSCFHTAFMLFCISVTRYLAIAHHRFYTKRLTFWTCLAVICMVWTLSVAMAFPPVLDVGTYSFIREEDQCTFQHRSFRANDSLGFMLLLALILLATQLVYLKLIFFVHDRRKMKPVQFVAAVSQNWTFHGPGASGQAAANWLAGFGRGPTPPTLLGIRQNANTTGRRRLLVLDEFKMEKRISRMFYIMTFLFLTLWGPYLVACYWRVFARGPVVPGGFLTAAVWMSFAQAGINPFVCIFSNRELRRCFSTTLLYCRKSRLPREPYCVI | |
| 4003023334334424101000102011200220332320000000123301100000000201010000000010000000343302102000000000001000000100000023330000200313332131000000000000000000000000232433454320203342020000011332022212212200100000022233334343243343333344443444434433443443333443332444343334333333441334220000000001111100200000000200044230030000000110020001000000000440040024002202464342401006 |
| Rank | PDB Hit | Iden1 | Iden2 | Cov. | Norm. Z-score | Download Align. | 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 340 360 | | | | | | | | | | | | | | | | | | | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sec.Str Seq | CCCCCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCSSCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCSSSCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCHHHHHHHHHHHCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHCCHHHHHHHHHHHCCCCCCCCCCCCCCCC MANYSHAADNILQNLSPLTAFLKLTSLGFIIGVSVVGNLLISILLVKDKTLHRAPYYFLLDLCCSDILRSAICFPFVFNSVKNGSTWTYGTLTCKVIAFLGVLSCFHTAFMLFCISVTRYLAIAHHRFYTKRLTFWTCLAVICMVWTLSVAMAFPPVLDVGTYSFIREEDQCTFQHRSFRANDSLGFMLLLALILLATQLVYLKLIFFVHDRRKMKPVQFVAAVSQNWTFHGPGASGQAAANWLAGFGRGPTPPTLLGIRQNANTTGRRRLLVLDEFKMEKRISRMFYIMTFLFLTLWGPYLVACYWRVFARGPVVPGGFLTAAVWMSFAQAGINPFVCIFSNRELRRCFSTTLLYCRKSRLPREPYCVI | |||||||||||||||||||||||||
| 1 | 5tjvA | 0.17 | 0.17 | 0.78 | 2.40 | Download | -ENFMDIECFMV--LNPSQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRCPSYHFIGSLAVADLLGSVIFVYSFIDFHVFH--RKDSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIAVLPLLGWNCEKLQSV---CSDIFPHIDETYLMFWIGVTSVLLLFIVYAYMYILWKA---------------------------------------------------------GKRAMSFSDQARMDIRLAKTLVLILVVLIICWGPLLAIMVYDVFGKMNKLIKTVFAFCSMLCLLNSTVNPIIYALRSKDLRHAFRSMF---------------- | |||||||||||||||||||
| 2 | 5uenA | 0.16 | 0.20 | 0.93 | 4.33 | Download | ----------------SAFQAAYIGIEVLIALVSVPGNVLVIWAVKVNQALRDATFCFIVSLAVADVAVGALVIPLAILINIGPQ-TYFH--TCLMVACPVLILTQSSILALLAIAVDRYLRVKIPLRYKMVVTPRRAAVAIAGCWILSFVVGLTPMFGWNNLSAVGSMGEPVIKCESMEYMVYFNFFVWVLPPLLLMVLIYLEVFYLIRKQLADLEDNWETLNDNVKDALTKMRAAALDAPEMKDFRHEAQAAAEQLKTTRNAYIQKYLERARSTLQKELKIAKSLALILFLFALSWLPLHILNCITLFCPSCHKPSILTYIAIFLTHGNSAMNPIVYAFRIQKFRVTFLKIWNDHFRCQPLE------ | |||||||||||||||||||
| 3 | 5uenA | 0.16 | 0.20 | 0.95 | 3.24 | Download | --------------SISAFQAAYIGIEVLIALVSVPGNVLVIWAVKVNQALRDATFCFIVSLAVADVAVGALVIPLAILINIGP-QTYF--HTCLMVACPVLILTQSSILALLAIAVDRYLRVKIPLRYKMVVTPRRAAVAIAGCWILSFVVGLTPMFGWNNLSGSMGEPVIKCEFESMEYMVYFNFFVWVLPPLLLMVLIYLEVFYLIRKQLADLEWETLNDNVKDALTKMAPEMKDFRHGFDILVGQIDDALKLANEGKVNAYIQKYLERARSTLQKELKIAKSLALILFLFALSWLPLHILNCITLFCPSCHKPSILTYIAIFLTHGNSAMNPIVYAFRIQKFRVTFLKIWNDHFRCQPLEVLF--- | |||||||||||||||||||
| 4 | 4ib4 | 0.20 | 0.23 | 0.93 | 1.57 | Download | -------EE------QGNKLHWAALLILMVIIPTIGGNTLVILAVSLEKKLQYATNYFLMSLAVADLLVGLFVMPIALLTIMFEAMWPLPLVLCPAWLFLDVLFSTASIWHLCAISVDRYIAIKKPIQANQYNSRATAFIKITVVWLISIGIAIPVIKGIETN---PNNITCVLTKERFGDFMLFGSLAAFFTPLAIMIVTYFLTIHALQKKAADLEDNWETLNDNLEKADNAAQTKMRAAALDAQKNEGKVKEAQAAAEQLKTTRNAYIQKYLQTISNEQRASKVLGIVFFLFLLMWCPFFITNITLVLCDSCNTLQMLLEIFVWIGYVSSGVNPLVYTLFNKTFRDAFGRYITCNYR----------- | |||||||||||||||||||
| 5 | 3uon | 0.18 | 0.21 | 0.91 | 1.23 | Download | ----------------TFEVVFIVLVAGSLSLVTIIGNILVMVSIKVNRHLQTVNNYFLFSLACADLIIGVFSMNLYTLYTVIGY-WPLGPVVCDLWLALDYVVSNASVMNLLIISFDRYFCVTKPLTYPVKRTTKMAGMMIAAAWVLSFILWAPAILFWQFIVGTVEDGECYIQFFSNAAVTFGTAIAAFYLPVIIMTVLYWHISRASKSRINIFEMLRIDEGLRLKDGHAIGRNTNGVITKDRGILRNAKLAKRWDEVNLAKSFRTGTWDAYPPPSREKKVTRTILAILLAFIITWAPYNVMVLINTFCAPC-IPNTVWTIGYWLCYINSTINPACYALCNATFKKTFKHLLM--------------- | |||||||||||||||||||
| 6 | 5uenA | 0.16 | 0.20 | 0.94 | 3.08 | Download | ----------------SAFQAAYIGIEVLIALVSVPGNVLVIWAVKVNQALRDATFCFIVSLAVADVAVGALVIPLAILINIGPQ-TYFH--TCLMVACPVLILTQSSILALLAIAVDRYLRVKIPLRYKMVVTPRRAAVAIAGCWILSFVVGLTPMFGWNNLSGSMGEPVIKCEFEKMEYMVYFNFFVWVLPPLLLMVLIYLEVFYLIRKQLADLEDNWETLNDNVKDALTKGQIDDALKLANEGKVKEAQAAAEQLKTTRNAYIQKYLERARSTLQKELKIAKSLALILFLFALSWLPLHILNCITLFCPSCHKPSILTYIAIFLTHGNSAMNPIVYAFRIQKFRVTFLKIWNDHFRCQPLEVLF--- | |||||||||||||||||||
| 7 | 4ib4 | 0.21 | 0.23 | 0.91 | 1.77 | Download | -------------------LHWAALLILMVIIPTIGGNTLVILAVSLEKKLQYATNYFLMSLAVADLLVGLFVMPIALLTIMFEAMWPLPLVLCPAWLFLDVLFSTASIWHLCAISVDRYIAIKKPIQANQYNSRATAFIKITVVWLISIGIAIPPIKGIET---NPNNITCVLTKERFGDFMLFGSLAAFFTPLAIMIVTYFLTIHALQKKAADLEDNWETLNDNLKEKADNAAQVRAALDAQKKDNEGKVKEAQAAAEQLKTTRNAYIQKYLQTISNEQRASKVLGIVFFLFLLMWCPFFITNITLVLCDSCTTLQMLLEIFVWIGYVSSGVNPLVYTLFNKTFRDAFGRYITCNYR----------- | |||||||||||||||||||
| 8 | 4ib4A | 0.20 | 0.23 | 0.93 | 4.06 | Download | ------------EEQGNKLHWAALLILMV-IIPTIGGNTLVILAVSLEKKLQYATNYFLMSLAVADLLVGLFVMPIALLTIMFEAMWPLPLVLCPAWLFLDVLFSTASIWHLCAISVDRYIAIKKPIQANQYNSRATAFIKITVVWLISIGIAIPVPIKG--IETNPNNITCVLTKERFGDFMLFGSLAAFFTPLAIMIVTYFLTIHALQKKAADLEDNWETLNDNLKVIEKADNLVGQIDDALKLANEGKVKEAQAAAEQLKTTRNAYIQKYLQTISNEQRASKVLGIVFFLFLLMWCPFFITNITLVLCDSCNTLQMLLEIFVWIGYVSSGVNPLVYTLFNKTFRDAFGRYITCNYR----------- | |||||||||||||||||||
| 9 | 4ib4A | 0.19 | 0.23 | 0.93 | 2.71 | Download | -------------EEQGNKLHWAALLILMVIIPTIGGNTLVILAVSLEKKLQYATNYFLMSLAVADLLVGLFVMPIALLTIMFEAMWPLPLVLCPAWLFLDVLFSTASIWHLCAISVDRYIAIKKPIQANQYNSRATAFIKITVVWLISIGIAIPVPIKGIETNPNNITCVL--TKERFGDFMLFGSLAAFFTPLAIMIVTYFLTIHALQKKAADLEDNWETLNDNLKVIEKADNAAQVKDALTKMRAAGKVKEAQAAAEQLKTTRNAYIQKYLQTISNEQRASKVLGIVFFLFLLMWCPFFITNITLVLCDSCNTLQMLLEIFVWIGYVSSGVNPLVYTLFNKTFRDAFGRYITCNYR----------- | |||||||||||||||||||
| 10 | 4nc3A | 0.19 | 0.23 | 0.94 | 2.91 | Download | -------------EEQGNKLHWAALLILMVIIPTIGGNTLVILAVSLEKKLQYATNYFLMSLAVADLLVGLFVMPIALLTIMFEAMWPLPLVLCPAWLFLDVLFSTASIWHLCAISVDRYIAIKKPIQANQYNSRATAFIKITVVWLISIGIAIPVPIKGIETVDNPNNITCVLTKERFGDFMLFGSLAAFFTPLAIMIVTYFLTIHALQKKAADLEDNWETLNDNLKVIEKADNAAQVKDALTKMRNEGKVKEAQAAAEQLKTTRNAYIQKYLQTISNEQRASKVLGIVFFLFLLMWCPFFITNITLVLCDSCNTLQMLLEIFVWIGYVSSGVNPLVYTLFNKTFRDAFGRYITCNYR----------- | |||||||||||||||||||
| ||||||||||||||||||||||||||
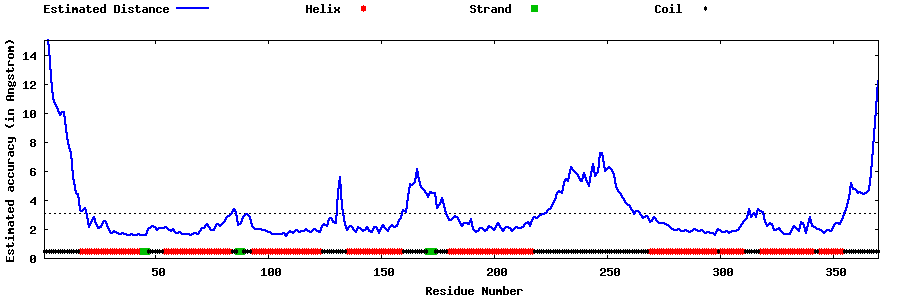
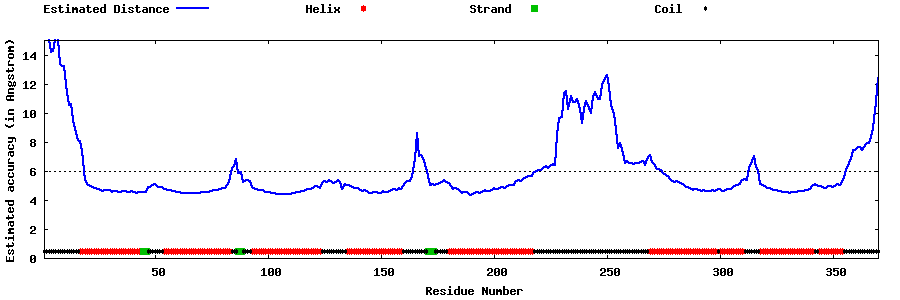
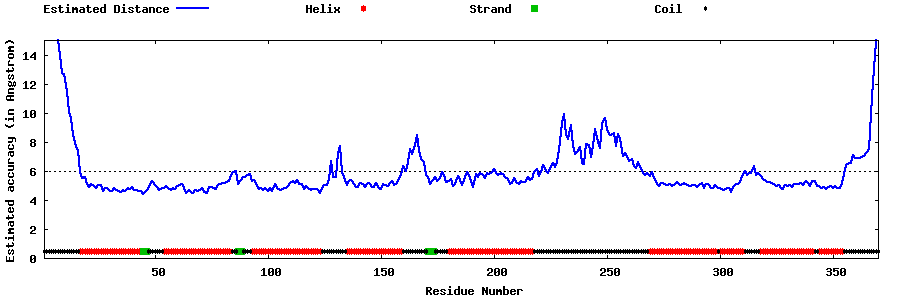
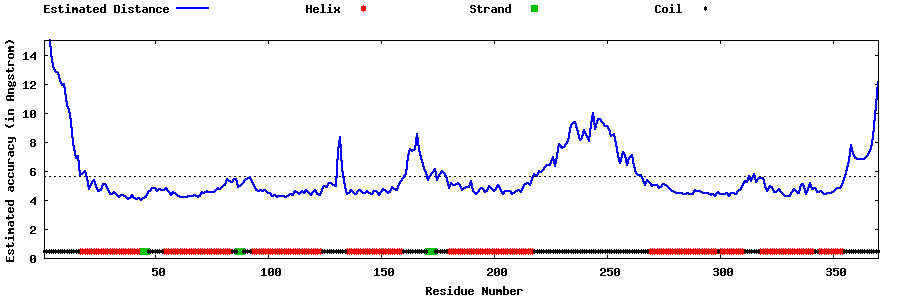
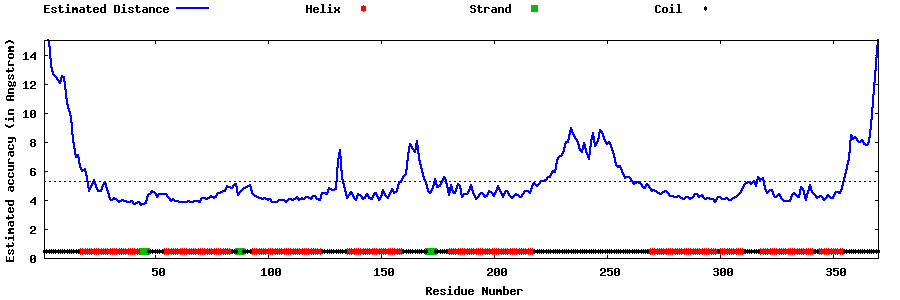
| Generated 3D models | Estimated local accuracy of models | ||
 | |||
|
|
|||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||