

| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| MGTDNQTWVSEFILLGLSSDWDTRVSLFVLFLVMYVVTVLGNCLIVLLIRLDSRLHTPMYFFLTNLSLVDVSYATSVVPQLLAHFLAEHKAIPFQSCAAQLFFSLALGGIEFVLLAVMAYDRYVAVCDALRYSAIMHGGLCARLAITSWVSGFISSPVQTAITFQLPMCRNKFIDHISCELLAVVRLACVDTSSNEVTIMVSSIVLLMTPFCLVLLSYIQIISTILKIQSREGRKKAFHTCASHLTVVALCYGVAIFTYIQPHSSPSVLQEKLFSVFYAILTPMLNPMIYSLRNKEVKGAWQKLLWKFSGLTSKLAT | |
| CCCCCCCSSSSSSSSCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCHHHHHHHHHHHHHHHHCCCHHHHHHHHCCCCCSSCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCCCCCCCCCHHHHHHHHCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCHHHCHHHHHHHHHHHHHHHHHSSCCCCCCCCCCCCCSSSSSHHCHHHCCCCHHHCCCCHHHHHHHHHHHHHHHHCHHHCCC | |
| 99988731456678779998618999999999999999998999999998677755748999878999987776247899999847899678389999999999999999999999997650776165445773148879999999999999999999999990348999894788604808888886247509999999999999999899999999999999823866254751431658789979999864773462789999877780899754154433124633146499999999999975202220379 |
| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| MGTDNQTWVSEFILLGLSSDWDTRVSLFVLFLVMYVVTVLGNCLIVLLIRLDSRLHTPMYFFLTNLSLVDVSYATSVVPQLLAHFLAEHKAIPFQSCAAQLFFSLALGGIEFVLLAVMAYDRYVAVCDALRYSAIMHGGLCARLAITSWVSGFISSPVQTAITFQLPMCRNKFIDHISCELLAVVRLACVDTSSNEVTIMVSSIVLLMTPFCLVLLSYIQIISTILKIQSREGRKKAFHTCASHLTVVALCYGVAIFTYIQPHSSPSVLQEKLFSVFYAILTPMLNPMIYSLRNKEVKGAWQKLLWKFSGLTSKLAT | |
| 87462302021000000033240001001303331331333332002001000300000010020000000001001002000101166220101002201120131023002000200200000002102000100330000000100210031031001100101103513010000032000100021032002102331233133122102301320020001031361231011000000000000220000000103263354340000011023203300300002042013002300232431355148 |
| Rank | PDB Hit | Iden1 | Iden2 | Cov. | Norm. Z-score | Download Align. | 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | | | | | | | | | | | | | | | | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sec.Str Seq | CCCCCCCSSSSSSSSCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCHHHHHHHHHHHHHHHHCCCHHHHHHHHCCCCCSSCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCCCCCCCCCHHHHHHHHCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCHHHCHHHHHHHHHHHHHHHHHSSCCCCCCCCCCCCCSSSSSHHCHHHCCCCHHHCCCCHHHHHHHHHHHHHHHHCHHHCCC MGTDNQTWVSEFILLGLSSDWDTRVSLFVLFLVMYVVTVLGNCLIVLLIRLDSRLHTPMYFFLTNLSLVDVSYATSVVPQLLAHFLAEHKAIPFQSCAAQLFFSLALGGIEFVLLAVMAYDRYVAVCDALRYSAIMHGGLCARLAITSWVSGFISSPVQTAITFQLPMCRNKFIDHISCELLAVVRLACVDTSSNEVTIMVSSIVLLMTPFCLVLLSYIQIISTILKIQSREGRKKAFHTCASHLTVVALCYGVAIFTYIQPHSSPSVLQEKLFSVFYAILTPMLNPMIYSLRNKEVKGAWQKLLWKFSGLTSKLAT | |||||||||||||||||||||||||
| 1 | 3emlA | 0.18 | 0.21 | 0.88 | 3.52 | Download | --------------------IMGSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAITISTG--FCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGGCGEGQVACLFEDVVP-----------MNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAI-IVGLFALCWLPLHIINCFTFFCPDCSHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVLRQ----- | |||||||||||||||||||
| 2 | 5tgzA | 0.19 | 0.22 | 0.89 | 2.16 | Download | -RGENFMDIECF----MVLNPSQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRCPSYHFIGSLAVADLLGSVIFVYSFIDFHVFHRK-DSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIAVL--------PLLGWNCEKL---------QSVCSDIFPHDKTYLMFWIGVVSVLLLFIVYAYMYILWKAHSHAPDQARMELAKTLVLILVVLIICWGPLLAIMVYDVFGKMNKLIKTVFAMLCLLNSTVNPIIYALRSKDLRHAFRSM------------- | |||||||||||||||||||
| 3 | 4iaqA | 0.21 | 0.22 | 0.85 | 2.19 | Download | ------------YIYQDSISLPWKVLLVMLLALITLATTLSNAFVIATVYRTRKLHTPANYLIASLAVTDLLVSILVMPISTMYTVTGR----WVVCDFWLSSDITCCTASIWHLCVIALDRYWAITDAVEYSAKRTPKRAAVMIALVWVFSISISL---------PPFFWRE---------------CVVNTDHILYTVYSTVGAFYFPTLLLIALYGRIYVEARSRIILLMAARERKATKTLGIILGAFIVCWLPFFIISLVMPIHLAIFDFFTWLGYLNSLINPIIYTMSNEDFKQAFHKLIRFK--------- | |||||||||||||||||||
| 4 | 4djh | 0.16 | 0.18 | 0.89 | 1.54 | Download | -------------------SPAIPVIITAVYSVVFVVGLVGNSLVMFVIIRYTKMKTATNIYIFNLALADALVTT-TMPFQSTVYLMNSWPFGDVLCKIVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKALDFRTPLKAKIINICIWLLSSSVGISAIVLGGTKVR---EDVDVIECSLQFPDD---DYSWWDLFMKICVFIFAFVIPVLIIIVCYTLMILRLKSVRLLRNLRRITRLVLVVVAVFVVCWTPIHIFILVEALGSAALSSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCFP---------- | |||||||||||||||||||
| 5 | 4yay | 0.18 | 0.22 | 0.89 | 1.24 | Download | KTTRN-AYIQKYLILNSSDCNYIFVMIPTLYSIIFVVGIFGNSLVVIVIYFYMKLKTVASVFLLNLALADLCFLLTLPLWAVYTAMEYRWPFGNYLCKIASASVSFNLYASVFLLTCLSIDRYLAIVHPMKSRLRRTMLVAKVTCIIIWLLAGLASLPAIIHRNVFF-------IITVCAFHYE--------TLPIGLGLTKNILGFLFPFLIILTSYTLIWKALK------KNDDIFKIIMAIVLFFFFSWIPHQIFTFLQLGIDIVDTAMPITICIAYFNNCLNPLFYGFLGKKFKRYFLQLL------------ | |||||||||||||||||||
| 6 | 3emlA | 0.18 | 0.21 | 0.87 | 3.73 | Download | ---------------------MGSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAIT--ISTGFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGGCGEGQVACLFEDVVP-----------MNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAIIVGLFALCWLPLHIINCF-TFFCPDCSHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVLRQ----- | |||||||||||||||||||
| 7 | 4iaq | 0.21 | 0.24 | 0.83 | 1.72 | Download | ---------------------PWKVLLVMLLALITLATTLSNAFVIATVYRTRKLHTPANYLIASLAVTDLLVSILVMPISTMYTVTGRWTLGQVVCDFWLSSDITCCTASIWHLCVIALDRYWAITDAVEYSAKRTPKRAAVMIALVWVFSISISLPPF-FWRQASE----------CVVNT-------D---HILYTVYSTVGAFYFPTLLLIALYGRIYVEARSRIADARERKATKTLGIILGAFIVCWLPFFIISLVMPIH--L-AIFDFFTWLGYLNSLINPIIYTMSNEDFKQAFHKLIRFK--------- | |||||||||||||||||||
| 8 | 4buoA | 0.17 | 0.18 | 0.89 | 2.92 | Download | ----NSD-------LDVNTDIYSKVLVTAIYLALFVVGTVGNSVTLFTLARKKSLQSTVDYYLGSLALSDLLILLLAMPVELYNFIWVHHPWAFAGCRGYYFLRDACTYATALNVVSLSVELYLAICHPFKAKTLMSRSRTKKFISAIWLASALLAIPMLFTMGLQNLSGDGTHPG------------GVDTATLKVVIQVNTFMSFLFPMLVASILNTVIANKLTVMVHQPGRVQALRRGVLVLRAVVIAFVVCWLPYHVRRLMFCYISDEQWTNALVYVSAAINPILYNLVSANFRQVFLSTL------------ | |||||||||||||||||||
| 9 | 3emlA | 0.18 | 0.21 | 0.88 | 4.73 | Download | --------------------IMGSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAITIS--TGFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLNCGQSQGCGEGQVACLFEDVVP-----------MNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAII-VGLFALCWLPLHIINCFTFFCPDCSHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVLRQ----- | |||||||||||||||||||
| 10 | 2ydoA | 0.16 | 0.21 | 0.92 | 5.70 | Download | -----------------------SSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSAAAADILVGVLAIPFAIA--ISTGFCAACHGCLFIACFVLVLTASSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGQPKEGKAHSQGCGEGQVACLFEDVVPMNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLKQMESTLQKEVHAAKSLAIIVGLFALCWLPFFCPDCSHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVLRQQEPFK | |||||||||||||||||||
| ||||||||||||||||||||||||||
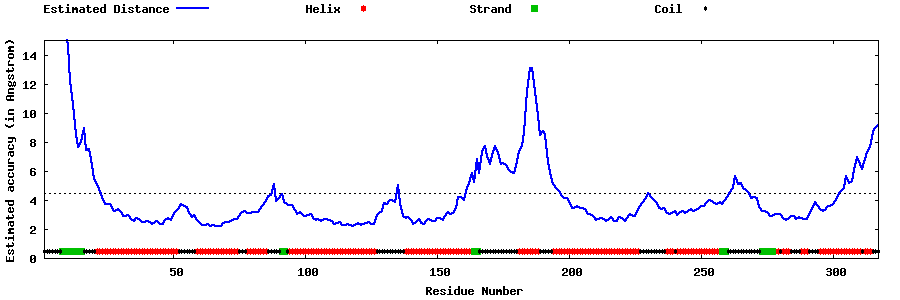
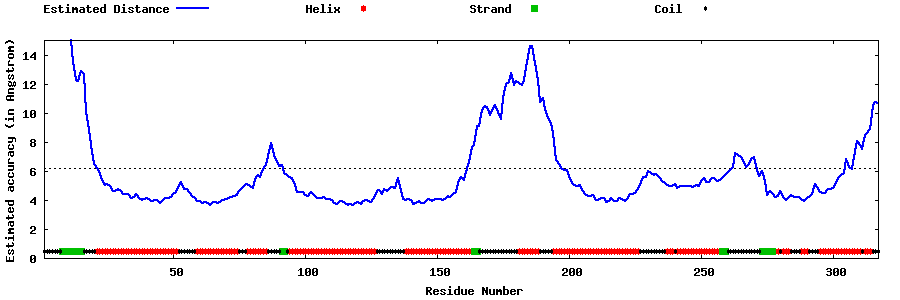
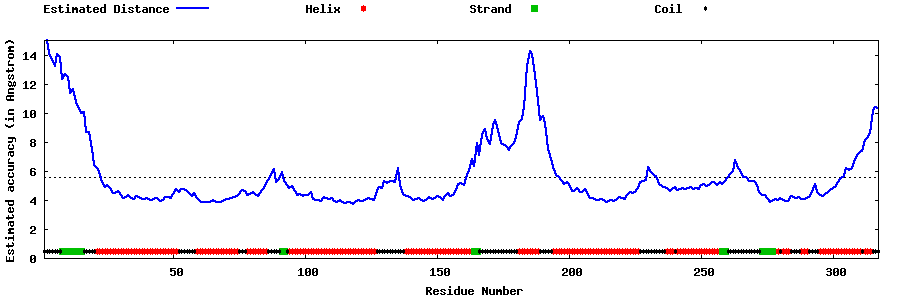
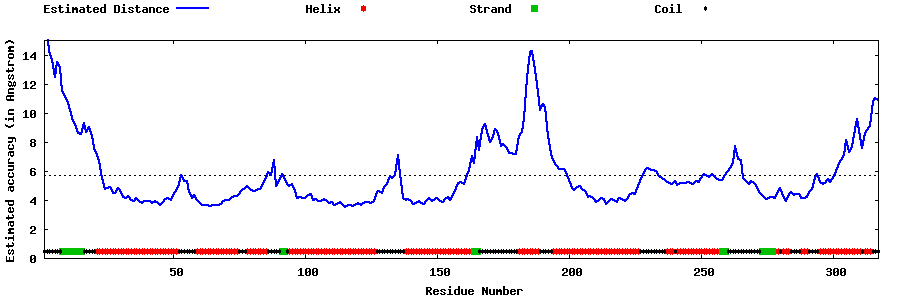
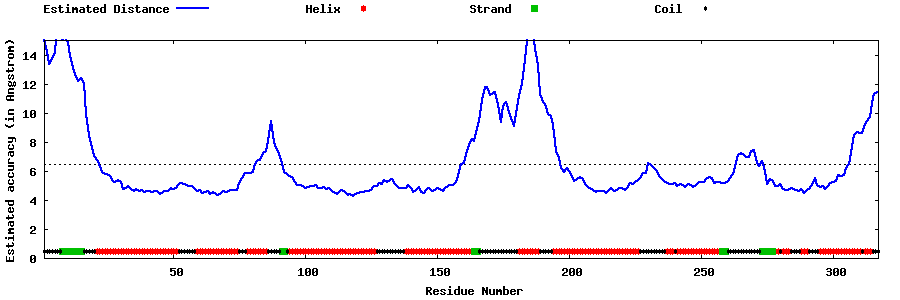
| Generated 3D models | Estimated local accuracy of models | ||
 | |||
|
|
|||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||