

| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 | |
| | | | | | | | | | | | | | | | | | |
| MQPSPPPTELVPSERAVVLLSCALSALGSGLLVATHALWPDLRSRARRLLLFLSLADLLSAASYFYGVLQNFAGPSWDCVLQGALSTFANTSSFFWTVAIALYLYLSIVRAARGPRTDRLLWAFHVVSWGVPLVITVAAVALKKIGYDASDVSVGWCWIDLEAKDHVLWMLLTGKLWEMLAYVLLPLLYLLVRKHINRAHTALSEYRPILSQEHRLLRHSSMADKKLVLIPLIFIGLRVWSTVRFVLTLCGSPAVQTPVLVVLHGIGNTFQGGANCIMFVLCTRAVRTRLFSLCCCCCSSQPPTKSPAGTPKAPAPSKPGESQESQGTPGELPST | |
| CCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCSSHHHHHHHHHHHHHHHHHHHHHHHHHHHSSSSSCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCSSSCCCCCCCCSSSSHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHCCHHHHHHHHHHHHHHCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCC | |
| 98999977888999999999999999999999999999867247557999999999999999999989862479998007999999999999999999999999828888536665205777788887447899999999870125788888877476758998818874069899999999999999999999999999988621762113378889999999999999999999999999999999845763058999999999999999999787530499999999999885569999998899999998999988788889999999998 |
| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 | |
| | | | | | | | | | | | | | | | | | |
| MQPSPPPTELVPSERAVVLLSCALSALGSGLLVATHALWPDLRSRARRLLLFLSLADLLSAASYFYGVLQNFAGPSWDCVLQGALSTFANTSSFFWTVAIALYLYLSIVRAARGPRTDRLLWAFHVVSWGVPLVITVAAVALKKIGYDASDVSVGWCWIDLEAKDHVLWMLLTGKLWEMLAYVLLPLLYLLVRKHINRAHTALSEYRPILSQEHRLLRHSSMADKKLVLIPLIFIGLRVWSTVRFVLTLCGSPAVQTPVLVVLHGIGNTFQGGANCIMFVLCTRAVRTRLFSLCCCCCSSQPPTKSPAGTPKAPAPSKPGESQESQGTPGELPST | |
| 47455347413300300011001001310220110000023013321100010000002321000001023256430000000230010120001000000320020001113373132000000010112112100000002231432343110000023633210000112332122223333221222012102212333343344344444133312303220101133233223323331000001363121100000000000311220000000124400420241023224444445345444544344444445445434464458 |
| Rank | PDB Hit | Iden1 | Iden2 | Cov. | Norm. Z-score | Download Align. | 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 | | | | | | | | | | | | | | | | | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sec.Str Seq | CCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCSSHHHHHHHHHHHHHHHHHHHHHHHHHHHSSSSSCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCSSSCCCCCCCCSSSSHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHCCHHHHHHHHHHHHHHCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCC MQPSPPPTELVPSERAVVLLSCALSALGSGLLVATHALWPDLRSRARRLLLFLSLADLLSAASYFYGVLQNFAGPSWDCVLQGALSTFANTSSFFWTVAIALYLYLSIVRAARGPRTDRLLWAFHVVSWGVPLVITVAAVALKKIGYDASDVSVGWCWIDLEAKDHVLWMLLTGKLWEMLAYVLLPLLYLLVRKHINRAHTALSEYRPILSQEHRLLRHSSMADKKLVLIPLIFIGLRVWSTVRFVLTLCGSPAVQTPVLVVLHGIGNTFQGGANCIMFVLCTRAVRTRLFSLCCCCCSSQPPTKSPAGTPKAPAPSKPGESQESQGTPGELPST | |||||||||||||||||||||||||
| 1 | 4l6rA | 0.13 | 0.17 | 0.84 | 1.94 | Download | -EVQKEVAKMYSSFQVMYTVGYSLSLGALLLALAILGGLSKLHCTRNAIHANLFASFVLKASSVLVIDGLLRTLSDAGCRVAAVFMQYGIVANYCWLLVEGLYLHNLLGLATLPE--RSFFSLYLGIGWGAPMLFVVPWAVVKCLFENVQCW--------TSNDNMGFWWILRFPVFLAILINFFIFVRIVQLLVAKLRAR----------QMHHTDYKFRLAKSTLTLIPLLGVHEVVFAFVTD----EHAQGTLRSAKLFFDLFLSSFQGLLVAVLYCFLNKEVQSELRRRWHRWRLGKVLWEERN--------------------------- | |||||||||||||||||||
| 2 | 5ee7A | 0.17 | 0.16 | 0.73 | 3.45 | Download | -----------SSFQVMYTVGYSLSLAALLLALAILGGLSKLHCTANAIHANLFLSFVLKASAVLFIDGLLRTVSVAACRVAAVFMQYGIVANYCWLLVEGLYLHNLL-------GLRSFFSLYLGIGWGAPALFVVPWAVVKCLFENVQ------CWTNMG-----FWWILRFPV--FLAILINFFIFVRIVQLLVAKLR--------ARQMHHTDYAFRLAKSTLTLIPLLGV-HFVVFAFVT--------DEHRSAKLFFDLALSSFQGLLVAVLYCFLNKEVQSELRRRW----------------------------------------- | |||||||||||||||||||
| 3 | 5uz7R | 0.16 | 0.18 | 0.79 | 2.63 | Download | -AFTPEKLKNAYVLYYLAIVGHSLSIFTLVISLGIFVFFRSLGCQRVTLHKNMFLTYILNSMIIIIHLVEV-RRDPVSCKILHFFHQYMMACNYFWMLCEGIYLHTLIVVAVFTEKQ--RLRWYYLLGWGFPLVPTTIHAITRAVYFNDN------CWLSVETHLL---YIIHGPVMAALVVNFFFLLNIVRVLVTK--------MRETHMYL-------KAVKATMILVPLLGIQFVVF------------PKMLGKIYDYVMHSLIHFQGFFVATIYCFCNNEVQTTVKRQWAQFKIQWNQRWG----------------------------- | |||||||||||||||||||
| 4 | 2ziy | 0.12 | 0.19 | 0.98 | 1.50 | Download | NPSDQVPDAVYYSLGIFIGICGIIGCGGNGIVIYLFTKTKSLQTPANMFIINLAFSDFTFSLVNFPLMTIKWIFGFAACKVYGFIGGIFGFMSIMTMAMISIDRYNVIGRPMAASKSHRRAFIMIIFVWLWSVLWAIGPIFGWAYTLEG---VLCNCSFDYISRDSTSNILCMFILGFFGPILIIFFCYFNIVMSVSNHEKEMAAMAKRLNKAQAGANAEMRLAKISIVIVSQFLL--SWSPYAVVAAQFGPLEWVTPYAAQLPVMFAKASAIHNPMIYSVSHPKFREAISQTFPWVLTFDDKETEDDKDAETEIPAGESSDAAPSADAAQMKE- | |||||||||||||||||||
| 5 | 5vaiA | 0.19 | 0.18 | 0.80 | 2.00 | Download | ----SSPEERLLSLYIIYTVGYALSFSALVIASAILLGFRHLHCTRNYIHLNLFASFILRALSVFIKDAASTYQDSLGCRLVFLLMQYCVAANYYWLLVEGAYLYTLLAFAVFSE--QRIFKLYLSIGWGVPLLFVIPWGIVKYL-YE-----DEGCWTRNSNM--NYWLIIRLPILFAIGVNFLI--FIRVICIVVSKLKANLMCK----T-DI---KCRLAKSTLTLIPLLGT-HEVI----FAFVMDEHAGTLRFVKLFTELSFTSFQGLMVAILYCFVNNEVQMEFRKSWERWR------------------------------------- | |||||||||||||||||||
| 6 | 4l6rA | 0.13 | 0.17 | 0.84 | 2.39 | Download | -EVQKEVAKMYSSFQVMYTVGYSLSLGALLLALAILGGLSKLHCTRNAIHANLFASFVLKASSVLVIDGLLRDGAVAGCRVAAVFMQYGIVANYCWLLVEGLYLHNLLGLATLPE--RSFFSLYLGIGWGAPMLFVVPWAVVKCLFENVQ--------CWTSNDNMGFWWILRFPVFLAILINFFIFVRIVQLLVAKLRARQMHHTD----------YKFRLAKSTLTLIPLLGVHEVVFAFV----TDEHAQGTLRSAKLFFDLFLSSFQGLLVAVLYCFLNKEVQSELRRRWHRWRLGKVLWEERN--------------------------- | |||||||||||||||||||
| 7 | 2ziy | 0.13 | 0.19 | 0.87 | 1.74 | Download | -----VPDAVYYSLGIFIGICGIIGCGGNGIVIYLFTKTKSLQTPANMFIINLAFSDFTFSLVNFPLMTIKWIFGFAACKVYGFIGGIFGFMSIMTMAMISIDRYNVIGRPMAASKKMSHRRAFIMIIFVWLWSVLWAIGPIFGWGAYTLEGVLCNCSFDYISRDSRSNILCMFILGFFGPILIIFFCYFNIVMSVSNHEKEMAAMAKRLRKAQAGANAEMRLAKISIVIVSQFLLSWSPYAVVALLAQFGPLEWVTPYAAQLPVMFAKASAIHNPMIYSVSHPKFREAISQTFPW--------------------------------------- | |||||||||||||||||||
| 8 | 1u19A | 0.13 | 0.19 | 0.93 | 3.54 | Download | PQYYLAEPWQFSMLAAYMFLLIMLGFPINFLTLYVTVQHKKLRTPLNYILLNLAVADLFMVFGGFTTTLYTSLHGYFGCNLEGFFATLGGEIALWSLVVLAIERYVVVCKPMSNFRFGENHAIMGVAFTWVMALACAAPPLVGWSRYIPEGMQC-SCGIDYYTPHEETFVIYMFVVHFIIPLIVIFFCYGQLVFTVK-------EAAAQQQESATTQKAEKEVTRMVIIMVIAFLICWLPYAGVAFYIFTHQGSDFGPIFMTIPAFFAKTSAVYNPVIYIMMNKQFRNCMVTTLCCGKNPLGDDEA---------STTVSKTETSQVAPA----- | |||||||||||||||||||
| 9 | 4l6rA | 0.13 | 0.17 | 0.84 | 2.13 | Download | -EVQKEVAKMYSSFQVMYTVGYSLSLGALLLALAILGGLSKLHCTRNAIHANLFASFVLKASSVLVIDGLLRTLSDAGCRVAAVFMQYGIVANYCWLLVEGLYLHNLLGLATLPE--RSFFSLYLGIGWGAPMLFVVPWAVVKCLFENVQC-----WTSNDNMG---FWWILRFPVFLAILINFFIFVRIVQLLVAKLRARQMHHTD----------YKFRLAKSTLTLIPLLGVHEVVFAF----VTDEHAQGTLRSAKLFFDLFLSSFQGLLVAVLYCFLNKEVQSELRRRWHRWRLGKVLWEERN--------------------------- | |||||||||||||||||||
| 10 | 5uz7R | 0.16 | 0.18 | 0.79 | 2.69 | Download | -AFTPEKLKNAYVLYYLAIVGHSLSIFTLVISLGIFVFFRSLGCQRVTLHKNMFLTYILNSMIIIIHLVE-VRRDPVSCKILHFFHQYMMACNYFWMLCEGIYLHTLIVVAVFTEKQR--LRWYYLLGWGFPLVPTTIHAITRAVYFNDN------CWLSVETH---LLYIIHGPVMAALVVNFFFLLNIVRVLVT--------KMRETH-------MYLKAVKATMILVPLLGIQFVVF------------PKMLGKIYDYVMHSLIHFQGFFVATIYCFCNNEVQTTVKRQWAQFKIQWNQRWG----------------------------- | |||||||||||||||||||
| ||||||||||||||||||||||||||
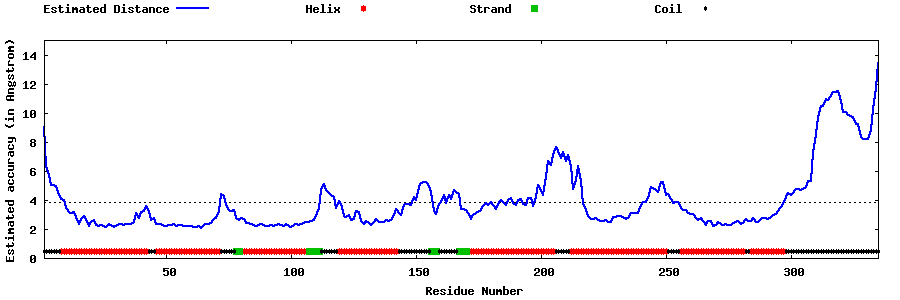
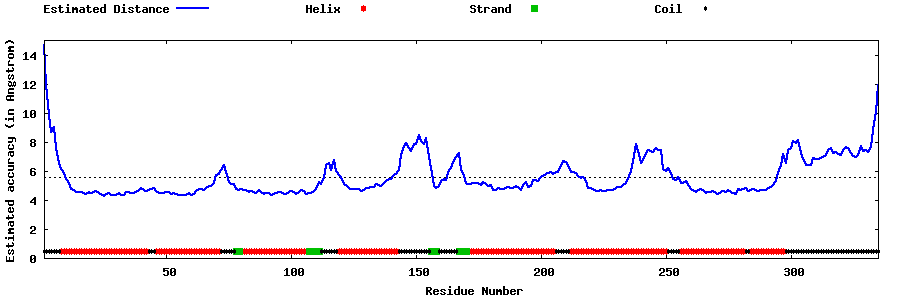
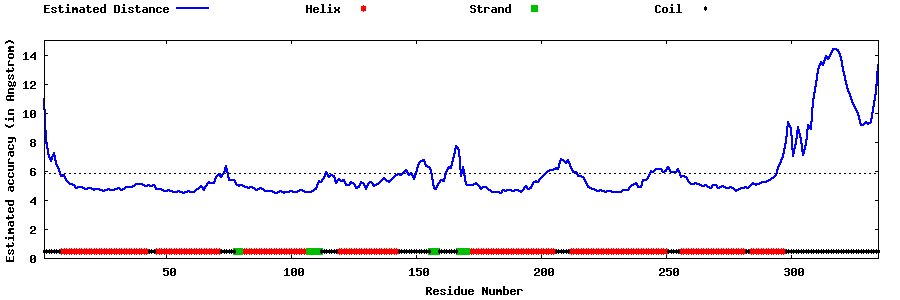
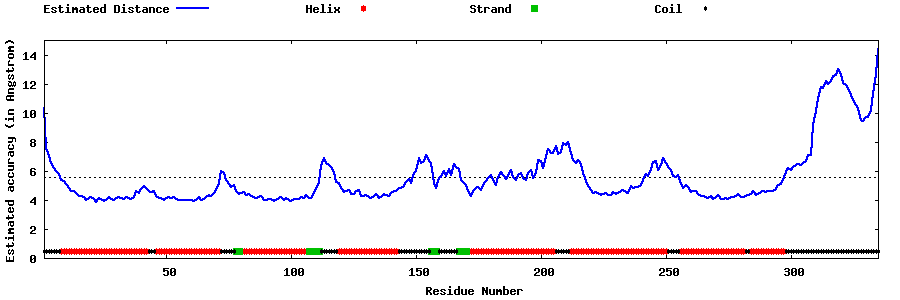
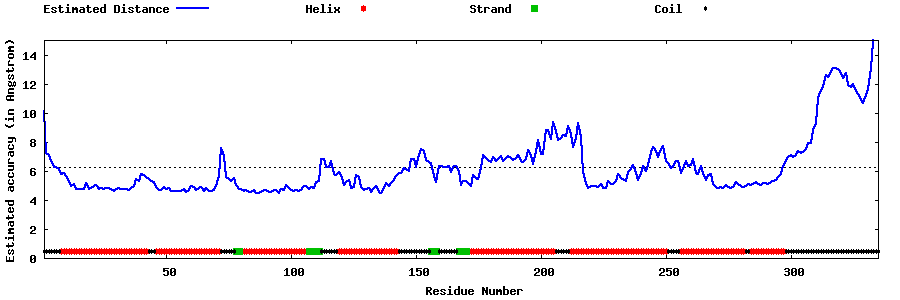
| Generated 3D models | Estimated local accuracy of models | ||
 | |||
|
|
|||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||