

| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 | |
| | | | | | | | | | | | | | | | | | |
| MNSWDAGLAGLLVGTMGVSLLSNALVLLCLLHSADIRRQAPALFTLNLTCGNLLCTVVNMPLTLAGVVAQRQPAGDRLCRLAAFLDTFLAANSMLSMAALSIDRWVAVVFPLSYRAKMRLRDAALMVAYTWLHALTFPAAALALSWLGFHQLYASCTLCSRRPDERLRFAVFTGAFHALSFLLSFVVLCCTYLKVLKVARFHCKRIDVITMQTLVLLVDLHPSVRERCLEEQKRRRQRATKKISTFIGTFLVCFAPYVITRLVELFSTVPIGSHWGVLSKCLAYSKAASDPFVYSLLRHQYRKSCKEILNRLLHRRSIHSSGLTGDSHSQNILPVSE | |
| CCHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSSCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCSSSSSCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHCCHHHHHHHHHHHCCCCCCCCCCCCCCCCCCCCCCCCCCCC | |
| 9631999999999999999998998725412107788985289999999999999999999999999948404864898599999999999999999999997262235557776657599898967899999999999999982551257876347880789615565899999999999999999999999999999998877521111110123455665331026788999999999999999999999899999999999848887389999999999999989899998289999999999847303799987788888887788799999 |
| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 | |
| | | | | | | | | | | | | | | | | | |
| MNSWDAGLAGLLVGTMGVSLLSNALVLLCLLHSADIRRQAPALFTLNLTCGNLLCTVVNMPLTLAGVVAQRQPAGDRLCRLAAFLDTFLAANSMLSMAALSIDRWVAVVFPLSYRAKMRLRDAALMVAYTWLHALTFPAAALALSWLGFHQLYASCTLCSRRPDERLRFAVFTGAFHALSFLLSFVVLCCTYLKVLKVARFHCKRIDVITMQTLVLLVDLHPSVRERCLEEQKRRRQRATKKISTFIGTFLVCFAPYVITRLVELFSTVPIGSHWGVLSKCLAYSKAASDPFVYSLLRHQYRKSCKEILNRLLHRRSIHSSGLTGDSHSQNILPVSE | |
| 7420200011201210330331120000000103403330000000000200010000012010011027200003003300000002001000300000001000000100313430233000000000122132201101100213236323312022230333232200001103312331120002002200210131034144344434444453443345443544333420000000000011123113100000010004030132010100030123002200010001741140014002221234424444333434445443468 |
| Rank | PDB Hit | Iden1 | Iden2 | Cov. | Norm. Z-score | Download Align. | 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 | | | | | | | | | | | | | | | | | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sec.Str Seq | CCHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSSCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCSSSSSCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHCCHHHHHHHHHHHCCCCCCCCCCCCCCCCCCCCCCCCCCCC MNSWDAGLAGLLVGTMGVSLLSNALVLLCLLHSADIRRQAPALFTLNLTCGNLLCTVVNMPLTLAGVVAQRQPAGDRLCRLAAFLDTFLAANSMLSMAALSIDRWVAVVFPLSYRAKMRLRDAALMVAYTWLHALTFPAAALALSWLGFHQLYASCTLCSRRPDERLRFAVFTGAFHALSFLLSFVVLCCTYLKVLKVARFHCKRIDVITMQTLVLLVDLHPSVRERCLEEQKRRRQRATKKISTFIGTFLVCFAPYVITRLVELFSTVPIGSHWGVLSKCLAYSKAASDPFVYSLLRHQYRKSCKEILNRLLHRRSIHSSGLTGDSHSQNILPVSE | |||||||||||||||||||||||||
| 1 | 3sn6R | 0.18 | 0.17 | 0.84 | 3.18 | Download | -EVWVVGMGIVMSLIVLAIVFGNVLVITAIAKFERLQT-VTNYFITSLACADLVMGLAVVPFGAAHILTKTWTFGNFWCEFWTSIDVLCVTASIETLCVIAVDRYFAITSPFKYQSLLTKNKARVIILMVWIVSGLTSFLPIQMHWYR---QEAINCYAEETCCDFFTNQAYAIASSIVSFYVPLVIMVFVYSRVFQEAKRQLQKIDKSEGR--------------------CLKEHKALKTLGIIMGTFTLCWLPFFIVNIVHVIQDNLIRKEVYILLNWIGYVNSGFNPLIYC-RSPDFRIAFQELLC--------------------------- | |||||||||||||||||||
| 2 | 4amjA | 0.20 | 0.19 | 0.87 | 4.36 | Download | --QWEAGMSLLMALVVLLIVAGNVLVIAAIGSTQRLQ-TLTNLFITSLACADLVVGLLVVPFGATLVVRGTWLWGSFLCELWTSLDVLCVTASIETLCVIAIDRYLAITSPFRYQSLMTRARAKVIICTVWAISALVSFLPIMMHWWRDEDPQALKCYQDPGCCDFVTNRAYAIASSIISFYIPLLIMIFVALRVYREAKEQ--------------IRKIDRASKRKTSRVMLMREHKALKTLGIIMGVFTLCWLPFFLVNIVNVFNRDLVPDWLFVAFNWLGYANSAMNPIIYCR-SPDFRKAFKRLLA--------------------------- | |||||||||||||||||||
| 3 | 3sn6R | 0.18 | 0.17 | 0.84 | 3.24 | Download | -EVWVVGMGIVMSLIVLAIVFGNVLVITAIAKFERLQTV-TNYFITSLACADLVMGLAVVPFGAAHILTKTWTFGNFWCEFWTSIDVLCVTASIETLCVIAVDRYFAITSPFKYQSLLTKNKARVIILMVWIVSGLTSFLPIQMHWYRAINCYAE-----ETCCDFFTNQAYAIASSIVSFYVPLVIMVFVYSRVFQEAKRQLQKIDKSEGRC--------------------LKEHKALKTLGIIMGTFTLCWLPFFIVNIVHVIQDNLIRKEVYILLNWIGYVNSGFNPLIYC-RSPDFRIAFQELLC--------------------------- | |||||||||||||||||||
| 4 | 2ziy | 0.16 | 0.19 | 0.96 | 1.55 | Download | PDAVYYSLGIFIGICGIIGCGGNGIVIYLFTKTKSLQ-TPANMFIINLAFSDFTFSLVGFPLMTISCFLKKWIFGFAACKVYGFIGGIFGFMSIMTMAMISIDRYNVIGRPMAASKKMSHRRAFIMIIFVWLWSVLWAIGPI-FGWGAYTLVLCNCSFDYISRDS--TTRSNILCMFILGFFGPILIIFFCYFNIVMSVSNHEKEMAAMAKRL--------NAKELRKAQAGANAEMRLAKISIVIVSQFLLSWSPYAVVALLAQFGPLEVTPYAAQLPVMFAKASAIHNPMIYSVSHPKFREAISQTFPWVLTCCQFDDKDAETEIPAGESSQMKE | |||||||||||||||||||
| 5 | 3d4s | 0.16 | 0.20 | 0.90 | 1.21 | Download | ---WVVGMGIVMSLIVLAIVFGNVLVITAIAKFERLQ-TVTNYFITSLACADLVMGLAVVPFGAAHILMKMWTFGNFWCEFWTSIDVLCVTASIWTLCVIAVDRYFAITSPFKYQSLLTKNKARVIILMVWIVSGLTSFLPIQMHWYRATHQ--EAINCYATCCDFFTNQAYAIASSIVSFYVPLVIMVFVYSRVFQEAKRQLNIFEMLRIDKGRNTNGVIDETGTWDAYKFCLKEHKALKTLGIIMGTFTLCWLPFFIVNIVHVIQDNLIRKEVYILLNWIGYVNSGFNPLIYCR-SPDFRIAFQELLCL-------------------------- | |||||||||||||||||||
| 6 | 3sn6R | 0.18 | 0.17 | 0.84 | 3.14 | Download | -EVWVVGMGIVMSLIVLAIVFGNVLVITAIAKFERLQ-TVTNYFITSLACADLVMGLAVVPFGAAHILTKTWTFGNFWCEFWTSIDVLCVTASIETLCVIAVDRYFAITSPFKYQSLLTKNKARVIILMVWIVSGLTSFLPIQMHWYR---QEAINCYAEETCCDFFTNQAYAIASSIVSFYVPLVIMVFVYSRVFQEAKRQLQKIDKSEGR--------------------CLKEHKALKTLGIIMGTFTLCWLPFFIVNIVHVIQDNLIRKEVYILLNWIGYVNSGFNPLIYC-RSPDFRIAFQELLC--------------------------- | |||||||||||||||||||
| 7 | 3d4s | 0.16 | 0.20 | 0.91 | 1.78 | Download | ---WVVGMGIVMSLIVLAIVFGNVLVITAIAKFERLQT-VTNYFITSLACADLVMGLAVVPFGAAHILMKMWTFGNFWCEFWTSIDVLCVTASIWTLCVIAVDRYFAITSPFKYQSLLTKNKARVIILMVWIVSGLTSFLPIQMHWYRATHEAINCYAEETCCDF-FTNQAYAIASSIVSFYVPLVIMVFVYSRVFQEAKRQLNIFEMLRIDETNGVITKDESTGTWDAYKFCLKEHKALKTLGIIMGTFTLCWLPFFIVNIVHVIQDNLIRKEVYILLNWIGYVNSGFNPLIYCR-SPDFRIAFQELLCL-------------------------- | |||||||||||||||||||
| 8 | 4bvnA | 0.20 | 0.19 | 0.85 | 4.04 | Download | LQQWEAGMSLLMALVVLLIVAGNVLVIAAIGSTQRLQT-LTNLFITSLACADLVVGLLVVPFGATLVVRGTWLWGSFLCELWTSLDVLCVTASVETLCVIAIDRYLAITSPFRYQSLMTRARAKVIICTVWAISALVSFLPIMMHWWRDEDPQALKCYQDPGCCDFVTNRAYAIASSIISFYIPLLIMIFVALRVYREAKEQIRKRKTS--------------------RVMLMREHKALKTLGIIMGVFTLCWLPFFLVNIVNVFNRDLVPKWLFVAFNWLGYANSAMNPIILC-RSPDFRKAFKRLLA--------------------------- | |||||||||||||||||||
| 9 | 3sn6R | 0.17 | 0.17 | 0.84 | 3.22 | Download | -EVWVVGMGIVMSLIVLAIVFGNVLVITAIAKFERLQT-VTNYFITSLACADLVMGLAVVPFGAAHILTKTWTFGNFWCEFWTSIDVLCVTASIETLCVIAVDRYFAITSPFKYQSLLTKNKARVIILMVWIVSGLTSFLPIQMHWYRQEAIN---CYAEETCCDFFTNQAYAIASSIVSFYVPLVIMVFVYSRVFQEAKRQLQKIDKSEGR--------------------CLKEHKALKTLGIIMGTFTLCWLPFFIVNIVHVIQDNLIRKEVYILLNWIGYVNSGFNPLIYC-RSPDFRIAFQELLC--------------------------- | |||||||||||||||||||
| 10 | 4bvnA | 0.20 | 0.19 | 0.85 | 4.38 | Download | LQQWEAGMSLLMALVVLLIVAGNVLVIAAIGSTQRLQT-LTNLFITSLACADLVVGLLVVPFGATLVVRGTWLWGSFLCELWTSLDVLCVTASVETLCVIAIDRYLAITSPFRYQSLMTRARAKVIICTVWAISALVSFLPIMMHWWRDEDPQALKCYQDPGCCDFVTNRAYAIASSIISFYIPLLIMIFVALRVYREAKEQIR--------------------KRKTSRVMLMREHKALKTLGIIMGVFTLCWLPFFLVNIVNVFNRDLVPKWLFVAFNWLGYANSAMNPIILC-RSPDFRKAFKRLLA--------------------------- | |||||||||||||||||||
| ||||||||||||||||||||||||||
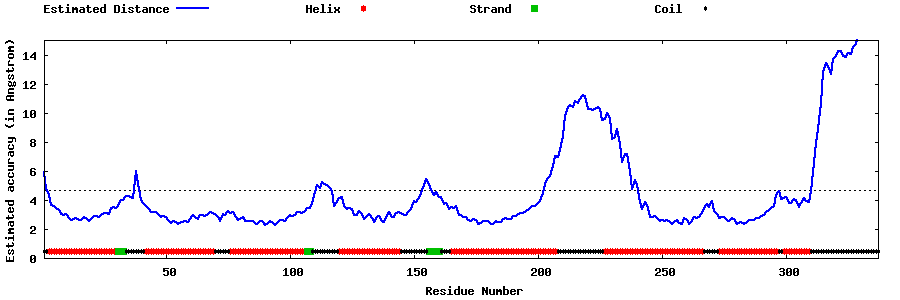
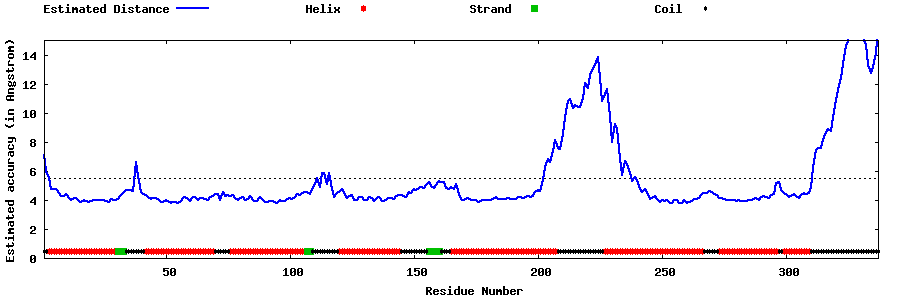
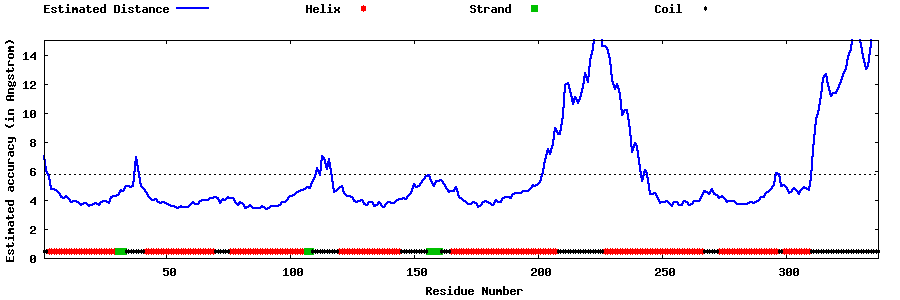
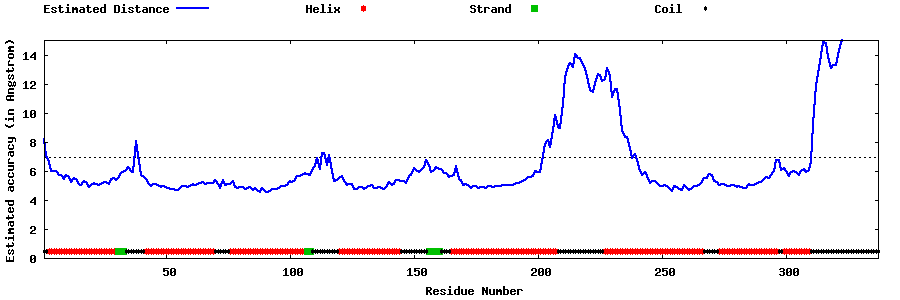
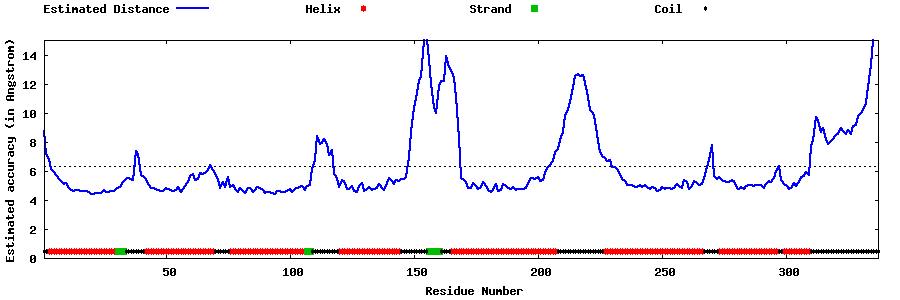
| Generated 3D models | Estimated local accuracy of models | ||
 | |||
|
|
|||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||