

| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 | |
| | | | | | | | | | | | | | | | | | |
| MLGSKPRVHLYILPCASQQVSTMGDRGTSNHSEMTDFILAGFRVRPELHILLFLLFLFVYAMILLGNVGMMTIIMTDPRLNTPMYFFLGNLSFIDLFYSSVIEPKAMINFWSENKSISFAGCVAQLFLFALLIVTEGFLLAAMAYDRFIAICNPLLYSVQMSTRLCTQLVAGSYFCGCISSVIQTSMTFTLSFCASRAVDHFYCDSRPLQRLSCSDLFIHRMISFSLSCIIILPTIIVIIVSYMYIVSTVLKIHSTEGHKKAFSTCSSHLGVVSVLYGAVFFMYLTPDRFPELSKVASLCYSLVTPMLNPLIYSLRNKDVQEALKKFLEKKNIIL | |
| CCCCCCCCSSSSSCCCHHHHHHHHHHHCCCCCCCSSSSSSCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCHHHHHHHHHHHHCCCCCCCHHHHHHHHHCCCCSSCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHSSSCCCCCCCCCCCCCHHHHHHHCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCCCHHCCHHHHHHHHHHHHHHHSSSSSCCCCCCCCCCSSSSSSCCHHHCCCCHHHCCCCHHHHHHHHHHHHHCCCCC | |
| 99988760378822525440478875167883231568964789936089999999999999998887999998508887774988876799884371014629999987238978858999999999999999999999999754234006210188126788999999999999999999999998354078999058541581888777156862667778888899999989999999999999981366757675430164888998999974441468168985667787988822014124356440564989999999997436049 |
| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 | |
| | | | | | | | | | | | | | | | | | |
| MLGSKPRVHLYILPCASQQVSTMGDRGTSNHSEMTDFILAGFRVRPELHILLFLLFLFVYAMILLGNVGMMTIIMTDPRLNTPMYFFLGNLSFIDLFYSSVIEPKAMINFWSENKSISFAGCVAQLFLFALLIVTEGFLLAAMAYDRFIAICNPLLYSVQMSTRLCTQLVAGSYFCGCISSVIQTSMTFTLSFCASRAVDHFYCDSRPLQRLSCSDLFIHRMISFSLSCIIILPTIIVIIVSYMYIVSTVLKIHSTEGHKKAFSTCSSHLGVVSVLYGAVFFMYLTPDRFPELSKVASLCYSLVTPMLNPLIYSLRNKDVQEALKKFLEKKNIIL | |
| 76545330100000023632531451375020200100000004324010100130333132133233200300100030000000001000000010000000200000015423010300000010000000000100000000000000200102030033000000000001000001000000020201442201000003210020001213010100010013113303300330231001000103126323100000000200000232011000000423432400000210320023003000021330140023005333016 |
| Rank | PDB Hit | Iden1 | Iden2 | Cov. | Norm. Z-score | Download Align. | 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 | | | | | | | | | | | | | | | | | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sec.Str Seq | CCCCCCCCSSSSSCCCHHHHHHHHHHHCCCCCCCSSSSSSCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCHHHHHHHHHHHHCCCCCCCHHHHHHHHHCCCCSSCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHSSSCCCCCCCCCCCCCHHHHHHHCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCCCHHCCHHHHHHHHHHHHHHHSSSSSCCCCCCCCCCSSSSSSCCHHHCCCCHHHCCCCHHHHHHHHHHHHHCCCCC MLGSKPRVHLYILPCASQQVSTMGDRGTSNHSEMTDFILAGFRVRPELHILLFLLFLFVYAMILLGNVGMMTIIMTDPRLNTPMYFFLGNLSFIDLFYSSVIEPKAMINFWSENKSISFAGCVAQLFLFALLIVTEGFLLAAMAYDRFIAICNPLLYSVQMSTRLCTQLVAGSYFCGCISSVIQTSMTFTLSFCASRAVDHFYCDSRPLQRLSCSDLFIHRMISFSLSCIIILPTIIVIIVSYMYIVSTVLKIHSTEGHKKAFSTCSSHLGVVSVLYGAVFFMYLTPDRFPELSKVASLCYSLVTPMLNPLIYSLRNKDVQEALKKFLEKKNIIL | |||||||||||||||||||||||||
| 1 | 3emlA | 0.17 | 0.17 | 0.82 | 3.25 | Download | ---------------------------------------------IMGSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAITIS---GFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGGCGEGQVACLFEDVVP-----------MNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAI-IVGLFALCWLPLHIINCFTFFCPDCSHAWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVLRQ | |||||||||||||||||||
| 2 | 5tgzA | 0.21 | 0.25 | 0.81 | 1.87 | Download | ---------------------------GENFMDIECF----MVLNPSQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRCPSYHFIGSLAVADLLGSVIFVYSFIDFHVFHRK-DSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIAVL------------------------PLLGWVCSDIFPHDKTYLMFWIGVVSVLLLFIVYAYMYILWKAHSHAPDQARMELAKTLVLILVVLIICWGPLLAIMVYDVFGKMNKLIKCSMLCLLNSTVNPIIYALRSKDLRHAFRSM-------- | |||||||||||||||||||
| 3 | 4iaqA | 0.15 | 0.19 | 0.81 | 2.11 | Download | -------------------------------------YIYQDSISLPWKVLLVMLLALITLATTLSNAFVIATVYRTRKLHTPANYLIASLAVTDLLVSILVMPISTMYTVTGRWTLGQVVCDFWLSSDITCCTASIWHLCVIALDRYWAITDAVEYSAKRTPKRAAVMIALVWVFSISISL------------------------PPFFWRECVVNTDHILYTVYSTVGAFYFPTLLLIALYGRIYVEARSRIIQLMAARERKATKTLGIILGAFIVCWLPFFIISLVMPLAIFDFFTWLGYLNSLINPIIYTMSNEDFKQAFHKLIRFK---- | |||||||||||||||||||
| 4 | 4djh | 0.15 | 0.21 | 0.83 | 1.52 | Download | --------------------------------------------SPAIPVIITAVYSVVFVVGLVGNSLVMFVIIRYTKMKTATNIYIFNLALADALVTT-TMPFQSTVYLMNSWPFGDVLCKIVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKALDFRTPLKAKIINICIWLLSSSVGISAIVLGGTKVR---EDVDVIECSLQFPDD---DYSWWDLFMKICVFIFAFVIPVLIIIVCYTLMILRLKSVRLDRNLRRITRLVLVVVAVFVVCWTPIHIFILVEALGALSSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCFP----- | |||||||||||||||||||
| 5 | 4yay | 0.14 | 0.20 | 0.92 | 1.26 | Download | LEDKSPEMKDFRHGFDEGKVAAAEQLKTTRNAYIQKYLILNSSDCNYIFVMIPTLYSIIFVVGIFGNSLVVIVIYFYMKLKTVASVFLLNLALADLCFLLTLPLWAVYTAMEYRWPFGNYLCKIASASVSFNLYASVFLLTCLSIDRYLAIVHPMKSRLRRTMLVAKVTCIIIWLLAGLASLPAIIH-RNVFF------IITVCAFHYE--------TLPIGLGLTKNILGFLFPFLIILTSYTLIWKALK------KNDDIFKIIMAIVLFFFFSWIPHQIFTFLDVGDVDTAMPITICIAYFNNCLNPLFYGFLGKKFKRYFLQLL------- | |||||||||||||||||||
| 6 | 3emlA | 0.17 | 0.17 | 0.82 | 3.46 | Download | ----------------------------------------------MGSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAIT--ISTGFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGGCGEGQVACLFEDVVP-----------MNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAIIVGLFALCWLPLHIINCF-TFFCPDCSHAPLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVLRQ | |||||||||||||||||||
| 7 | 4iaq | 0.18 | 0.20 | 0.79 | 1.73 | Download | ----------------------------------------------PWKVLLVMLLALITLATTLSNAFVIATVYRTRKLHTPANYLIASLAVTDLLVSILVMPISTMYTVTGRWTLGQVVCDFWLSSDITCCTASIWHLCVIALDRYWAITDAVEYSAKRTPKRAAVMIALVWVFSISISLPPFFW-RQASE----------CVVN----------TDHILYTVYSTVGAFYFPTLLLIALYGRIYVEARSRIADARERKATKTLGIILGAFIVCWLPFFIISLVMPIHLAIF-DFFTWLGYLNSLINPIIYTMSNEDFKQAFHKLIRFK---- | |||||||||||||||||||
| 8 | 4ea3A | 0.19 | 0.18 | 0.77 | 2.70 | Download | --------------------------------------------PLGLKVTIVGLYLAVCVGGLLGNCLVMYVILRHTKMKTATNIYIFNLALADTLVLLT-LPFQGTDILLGFWPFGNALCKTVIAIDYYNMFTSTFTLTAMSVDRYVAICHPTSSKAQAVNVAIWALASVVGVPVAIMGSAQVEDEIECLVEIPTPQDYWGPVFAICIFLFSF-----------------IVPVLVISVCYSLMIRRLRGVRLLSGSVAVFVGCWTPVQVFVLAQGLG----VQPSSTAVAILRFCTALGYVNSCLNPILYAFLDENFKACFR---------- | |||||||||||||||||||
| 9 | 3emlA | 0.17 | 0.17 | 0.81 | 4.87 | Download | -----------------------------------------------ISSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAITIS---GFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLNCGQSQGCGEGQVACLFEDVVP-----------MNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAI-IVGLFALCWLPLHIINCFTFFCPDCSHAPLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVLRQ | |||||||||||||||||||
| 10 | 2ydoA | 0.16 | 0.17 | 0.85 | 5.51 | Download | ------------------------------------------------SSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSAAAADILVGVLAIPFAIA--ISTGFCAACHGCLFIACFVLVLTASSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGQPKEGKAHSQGCGEGQVACLFEDVVPMNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLKQMESTLQKEVHAAKSLAIIVGLFALCWFTFFCPDCSHAPLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVLRQ | |||||||||||||||||||
| ||||||||||||||||||||||||||
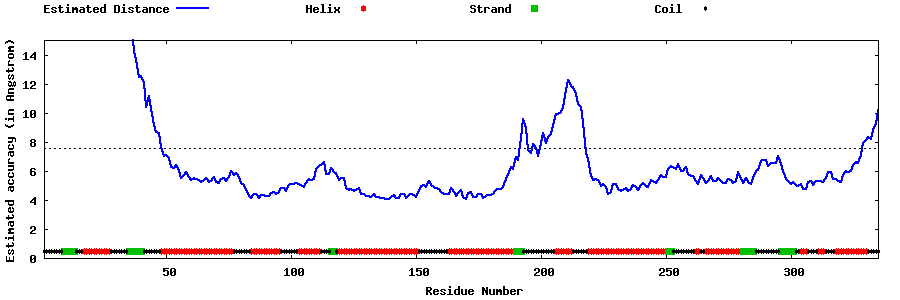
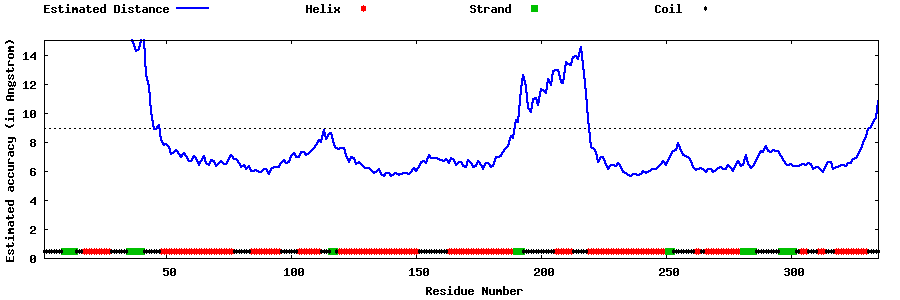
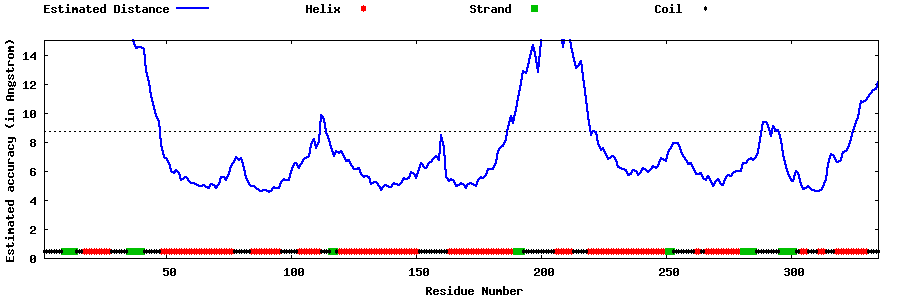
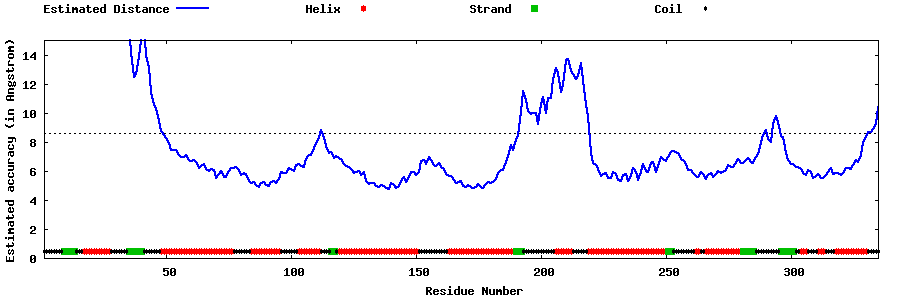
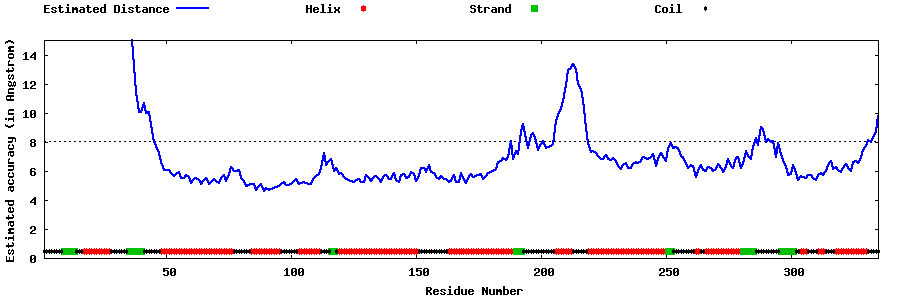
| Generated 3D models | Estimated local accuracy of models | ||
 | |||
|
|
|||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||