

| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 340 | |
| | | | | | | | | | | | | | | | | | | |
| MTSNFSQPVVQLCYEDVNGSCIETPYSPGSRVILYTAFSFGSLLAVFGNLLVMTSVLHFKQLHSPTNFLIASLACADFLVGVTVMLFSMVRTVESCWYFGAKFCTLHSCCDVAFCYSSVLHLCFICIDRYIVVTDPLVYATKFTVSVSGICISVSWILPLTYSGAVFYTGVNDDGLEELVSALNCVGGCQIIVSQGWVLIDFLLFFIPTLVMIILYSKIFLIAKQQAIKIETTSSKVESSSESYKIRVAKRERKAAKTLGVTVLAFVISWLPYTVDILIDAFMGFLTPAYIYEICCWSAYYNSAMNPLIYALFYPWFRKAIKLILSGDVLKASSSTISLFLE | |
| CCCCCCCCCCCCCCCCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHSSSSSCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCSSCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCCCCCCCCCSSSCCSSSSSSSHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHCCHHHHHHHHHHHCCCCCCCCCCCCCCCCC | |
| 998778652002456678778899998389999999999999999994898177577568888378999999999999999999899999999491557637886899999999999999999999972224665145864454888888476999999999999998615446767776346789877515224077641378999999999999999999998334533344433222113345566665542540135888899983469999999986388781899999999999998086999995789999999999476458999985136679 |
| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 340 | |
| | | | | | | | | | | | | | | | | | | |
| MTSNFSQPVVQLCYEDVNGSCIETPYSPGSRVILYTAFSFGSLLAVFGNLLVMTSVLHFKQLHSPTNFLIASLACADFLVGVTVMLFSMVRTVESCWYFGAKFCTLHSCCDVAFCYSSVLHLCFICIDRYIVVTDPLVYATKFTVSVSGICISVSWILPLTYSGAVFYTGVNDDGLEELVSALNCVGGCQIIVSQGWVLIDFLLFFIPTLVMIILYSKIFLIAKQQAIKIETTSSKVESSSESYKIRVAKRERKAAKTLGVTVLAFVISWLPYTVDILIDAFMGFLTPAYIYEICCWSAYYNSAMNPLIYALFYPWFRKAIKLILSGDVLKASSSTISLFLE | |
| 744534473031014322121344422301100000111210231333120000000113300100000000001001100000021100200222011032002000100010000003000000000000002003143312330000000000220231021000000234325423342412320201323300310030132002000200210020013203333433443432323221323434341030000000001000210000000200051000310000001201310130010000104301300120001200244034240258 |
| Rank | PDB Hit | Iden1 | Iden2 | Cov. | Norm. Z-score | Download Align. | 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 340 | | | | | | | | | | | | | | | | | | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sec.Str Seq | CCCCCCCCCCCCCCCCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHSSSSSCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCSSCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCCCCCCCCCSSSCCSSSSSSSHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHCCHHHHHHHHHHHCCCCCCCCCCCCCCCCC MTSNFSQPVVQLCYEDVNGSCIETPYSPGSRVILYTAFSFGSLLAVFGNLLVMTSVLHFKQLHSPTNFLIASLACADFLVGVTVMLFSMVRTVESCWYFGAKFCTLHSCCDVAFCYSSVLHLCFICIDRYIVVTDPLVYATKFTVSVSGICISVSWILPLTYSGAVFYTGVNDDGLEELVSALNCVGGCQIIVSQGWVLIDFLLFFIPTLVMIILYSKIFLIAKQQAIKIETTSSKVESSSESYKIRVAKRERKAAKTLGVTVLAFVISWLPYTVDILIDAFMGFLTPAYIYEICCWSAYYNSAMNPLIYALFYPWFRKAIKLILSGDVLKASSSTISLFLE | |||||||||||||||||||||||||
| 1 | 3sn6R | 0.30 | 0.26 | 0.83 | 2.79 | Download | ---------------------------EVWVVGMGIVMSLIVLAIVFGNVLVITAIAKFERLQTVTNYFITSLACADLVMGLAVVPFGAAHILTKTWTFGNFWCEFWTSIDVLCVTASIETLCVIAVDRYFAITSPFKYQSLLTKNKARVIILMVWIVSGLTSFLPIQMHWYRQEAINC---YAEETCCDFFTNQYAIASSIVSFYVPLVIMVFVYSRVFQEAKRQLQKIDKSEGR------------CLKEHKALKTLGIIMGTFTLCWLPFFIVNIVHVIQDNLIRKEVYILLNWIGYVNSGFNPLIYC-RSPDFRIAFQELLC---------------- | |||||||||||||||||||
| 2 | 4amjA | 0.31 | 0.28 | 0.85 | 4.48 | Download | --------------------------SQQWEAGMSLLMALVVLLIVAGNVLVIAAIGSTQRLQTLTNLFITSLACADLVVGLLVVPFGATLVVRGTWLWGSFLCELWTSLDVLCVTASIETLCVIAIDRYLAITSPFRYQSLMTRARAKVIICTVWAISALVSFLPIMMHWWRDEDPQALKCYQDPGCCDFVTNRAYAIASIISFYIPLLIMIFVALRVYREAKEQIRKIDRASKRKT------SRVMLMREHKALKTLGIIMGVFTLCWLPFFLVNIVNVFNRDLVPDWLFVAFNWLGYANSAMNPIIYCR-SPDFRKAFKRLL----------------- | |||||||||||||||||||
| 3 | 3sn6R | 0.30 | 0.26 | 0.83 | 3.26 | Download | ---------------------------EVWVVGMGIVMSLIVLAIVFGNVLVITAIAKFERLQTVTNYFITSLACADLVMGLAVVPFGAAHILTKTWTFGNFWCEFWTSIDVLCVTASIETLCVIAVDRYFAITSPFKYQSLLTKNKARVIILMVWIVSGLTSFLPIQMHWYRQEAINCYAE---ETCCDFFTNQAYAIASSIVSYVPLVIMVFVYSRVFQEAKRQLQKIDKSEGRC------------LKEHKALKTLGIIMGTFTLCWLPFFIVNIVHVIQDNLIRKEVYILLNWIGYVNSGFNPLIYC-RSPDFRIAFQELLC---------------- | |||||||||||||||||||
| 4 | 4ib4 | 0.25 | 0.25 | 0.87 | 1.53 | Download | ---------------------EE---QGNKLHWAALLILMVIIPTIGGNTLVILAVSLEKKLQYATNYFLMSLAVADLLVGLFVMPIALLTIMEAMWPLPLVLCPAWLFLDVLFSTASIWHLCAISVDRYIAIKKPIQANQYNSRATAFIKITVVWLISIGIAIPVPIKGIETN-PN----N----ITCVLTKEGDFMFGSLAAFFTPLAIMIVTYFLTIHALQKKAADLEDNWETADKAYIQKYLQTISNEQRASKVLGIVFFLFLLMWCPFFITNITLVLCDSCNLQMLLEIFVWIGYVSSGVNPLVYTLFNKTFRDAFGRYITCNYR------------ | |||||||||||||||||||
| 5 | 3d4s | 0.28 | 0.28 | 0.87 | 1.18 | Download | -----------------------------WVVGMGIVMSLIVLAIVFGNVLVITAIAKFERLQTVTNYFITSLACADLVMGLAVVPFGAAHILMKMWTFGNFWCEFWTSIDVLCVTASIWTLCVIAVDRYFAITSPFKYQSLLTKNKARVIILMVWIVSGLTSFLPIQMHWYRATHQAINCYAE-ETCCDFFTNQAYAIASIVSFYVPLVIMVFVYSRVFQEAKRQLNIFEMAIGRNTNGVTWDAYKFCLKEHKALKTLGIIMGTFTLCWLPFFIVNIVHVIQDNLIRKEVYILLNWIGYVNSGFNPLIYCR-SPDFRIAFQELLCL--------------- | |||||||||||||||||||
| 6 | 3sn6R | 0.30 | 0.26 | 0.82 | 3.05 | Download | ----------------------------VWVVGMGIVMSLIVLAIVFGNVLVITAIAKFERLQTVTNYFITSLACADLVMGLAVVPFGAAHILTKTWTFGNFWCEFWTSIDVLCVTASIETLCVIAVDRYFAITSPFKYQSLLTKNKARVIILMVWIVSGLTSFLPIQMHWY---RQEAINCYAEETCCDFFTNQAYAISSIVSFYVPLVIMVFVYSRVFQEAKRQLQKIDKSEGR------------CLKEHKALKTLGIIMGTFTLCWLPFFIVNIVHVIQDNLIRKEVYILLNWIGYVNSGFNPLIYC-RSPDFRIAFQELLC---------------- | |||||||||||||||||||
| 7 | 3uon | 0.29 | 0.29 | 0.86 | 1.73 | Download | ------------------------------VVFIVLVAGSLSLVTIIGNILVMVSIKVNRHLQTVNNYFLFSLACADLIIGVFSMNLYTLYTVIGYWPLGPVVCDLWLALDYVVSNASVMNLLIISFDRYFCVTKPLTYPVKRTTKMAGMMIAAAWVLSFILWAPAILFWQFIVGVRT-VEDGECYIQFFSN-AAVTFGTAIAAFYLPVIIMTVLYWHISRASKSRINIFEMLGPAATNTGTWDAYPPPSREKKVTRTILAILLAFIITWAPYNVMVLINTFCAPCIPNTVWTIGYWLCYINSTINPACYALCNATFKKTFKHLLM---------------- | |||||||||||||||||||
| 8 | 2z73A | 0.21 | 0.25 | 0.91 | 3.74 | Download | --------------ETWNPSIVVDQVPDAVYYSLGIFIGICGIIGCGGNGIVIYLFTKTKSLQTPANMFIINLAFSDFTFSLVNGPLMTISCFLKKWIFGFAACKVYGFIGGIFGFMSIMTMAMISIDRYNVIGRPMAASKKMSHRRAFIMIIFVWLWSVLWAIGPIFGWG------------AEGVLCNCSFDYISLCMFILGFFGPILIIFFCYFNIVMSVSNHEKEMAAMAKRLNAKELRKAQAGANAEMRLAKISIVIVSQFLLSWSPYAVVALLAQFGPLWVTPYAAQLPVMFAKASAIHNPMIYSVSHPKFREAISQTFPWVLTCCQFD------D | |||||||||||||||||||
| 9 | 3sn6R | 0.30 | 0.26 | 0.83 | 2.96 | Download | ---------------------------EVWVVGMGIVMSLIVLAIVFGNVLVITAIAKFERLQTVTNYFITSLACADLVMGLAVVPFGAAHILTKTWTFGNFWCEFWTSIDVLCVTASIETLCVIAVDRYFAITSPFKYQSLLTKNKARVIILMVWIVSGLTSFLPIQMHWYRQEAINCYAEETC---CDFFTNQAYAIASIVSFYVPLVIMVFVYSRVFQEAKRQLQKIDKSEGR------------CLKEHKALKTLGIIMGTFTLCWLPFFIVNIVHVIQDNLIRKEVYILLNWIGYVNSGFNPLIYC-RSPDFRIAFQELLC---------------- | |||||||||||||||||||
| 10 | 3zpqA | 0.31 | 0.28 | 0.84 | 3.86 | Download | ---------------------GAELLSQQWEAGMSLLMALVVLLIVAGNVLVIAAIGSTQRLQTLTNLFITSLACADLVVGLLVVPFGATLVVRGTWLWGSFLCELWTSLDVLCVTASIETLCVIAIDRYLAITSPFRYQSLMTRARAKVIICTVWAISALVSFLPIMMHWWRDEDPQALKCYQDPGCCDFVTNRAYAIASSISFYIPLLIMIFVALRVYREAKEQS------------------RVMLMREHKALKTLGIIMGVFTLCWLPFFLVNIVNVFNRDLVPDWLFVAFNWLGYANSAMNPIIYCR-SPDFRKAFKRLLA---------------- | |||||||||||||||||||
| ||||||||||||||||||||||||||
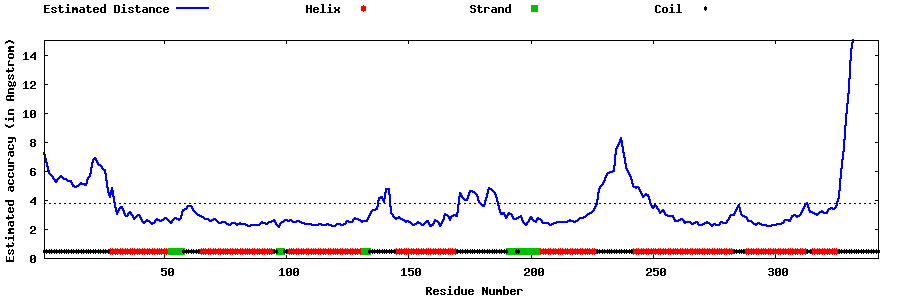
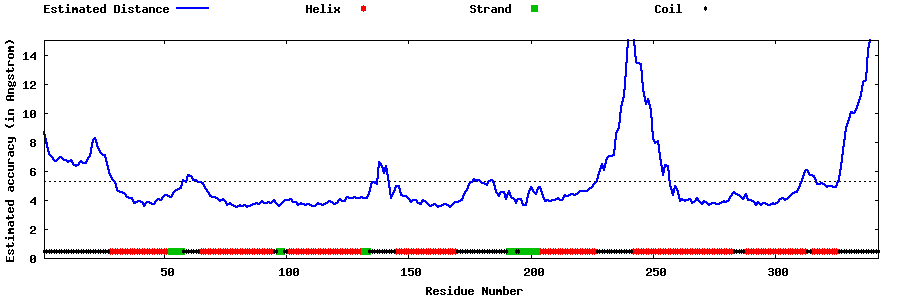
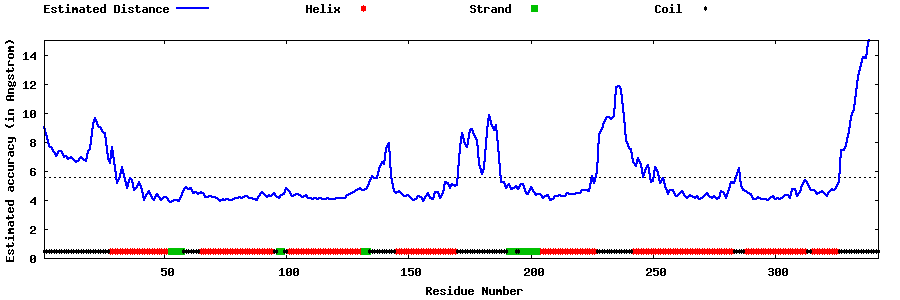
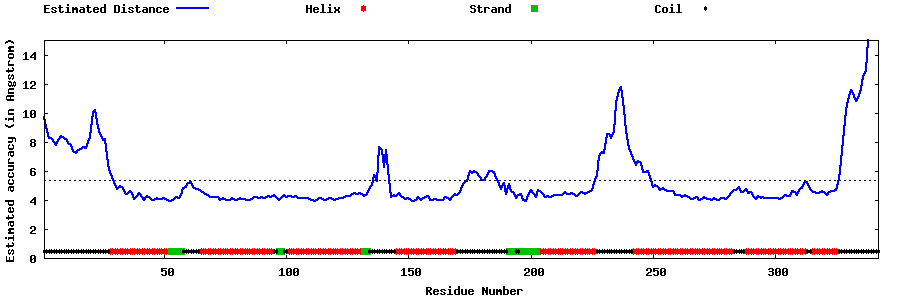
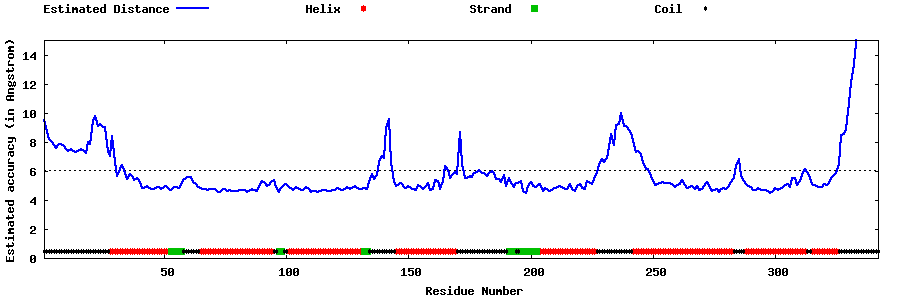
| Generated 3D models | Estimated local accuracy of models | ||
 | |||
|
|
|||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||