

| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 340 360 | |
| | | | | | | | | | | | | | | | | | | | |
| MANTTGEPEEVSGALSPPSASAYVKLVLLGLIMCVSLAGNAILSLLVLKERALHKAPYYFLLDLCLADGIRSAVCFPFVLASVRHGSSWTFSALSCKIVAFMAVLFCFHAAFMLFCISVTRYMAIAHHRFYAKRMTLWTCAAVICMAWTLSVAMAFPPVFDVGTYKFIREEDQCIFEHRYFKANDTLGFMLMLAVLMAATHAVYGKLLLFEYRHRKMKPVQMVPAISQNWTFHGPGATGQAAANWIAGFGRGPMPPTLLGIRQNGHAASRRLLGMDEVKGEKQLGRMFYAITLLFLLLWSPYIVACYWRVFVKACAVPHRYLATAVWMSFAQAAVNPIVCFLLNKDLKKCLRTHAPCWGTGGAPAPREPYCVM | |
| CCCCCCCCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHSSSSSCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCSSCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCSSSSCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHCCHHHHHHHHHHHCCCCCCCCCCCCCCCCCC | |
| 9888999877787778970899999999999999999988873110355577663768999999999999999998899999980895347734887899999999999999999999986985246111665464898999799999999999999998467766688998357258762015556566667889999999999999999999873431033332333223577643233344445667777655332123333233321245788877888999999999999999839999999999737777877999999999999998989999836899999999980754889999999985279 |
| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 340 360 | |
| | | | | | | | | | | | | | | | | | | | |
| MANTTGEPEEVSGALSPPSASAYVKLVLLGLIMCVSLAGNAILSLLVLKERALHKAPYYFLLDLCLADGIRSAVCFPFVLASVRHGSSWTFSALSCKIVAFMAVLFCFHAAFMLFCISVTRYMAIAHHRFYAKRMTLWTCAAVICMAWTLSVAMAFPPVFDVGTYKFIREEDQCIFEHRYFKANDTLGFMLMLAVLMAATHAVYGKLLLFEYRHRKMKPVQMVPAISQNWTFHGPGATGQAAANWIAGFGRGPMPPTLLGIRQNGHAASRRLLGMDEVKGEKQLGRMFYAITLLFLLLWSPYIVACYWRVFVKACAVPHRYLATAVWMSFAQAAVNPIVCFLLNKDLKKCLRTHAPCWGTGGAPAPREPYCVM | |
| 6404134344344434233011001011112002203321200000001233011000000000010000000000100000002333021030010000000000000001000000233300002003133311220000000000000000000000002223334542101013321100000013211120120013000200210222233343342433333233333433444344334434434332232323243333433334344333421000000103101210131110000000004323013001000000012000100000000044004001400202224645433301006 |
| Rank | PDB Hit | Iden1 | Iden2 | Cov. | Norm. Z-score | Download Align. | 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 340 360 | | | | | | | | | | | | | | | | | | | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sec.Str Seq | CCCCCCCCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHSSSSSCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCSSCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCSSSSCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHCCHHHHHHHHHHHCCCCCCCCCCCCCCCCCC MANTTGEPEEVSGALSPPSASAYVKLVLLGLIMCVSLAGNAILSLLVLKERALHKAPYYFLLDLCLADGIRSAVCFPFVLASVRHGSSWTFSALSCKIVAFMAVLFCFHAAFMLFCISVTRYMAIAHHRFYAKRMTLWTCAAVICMAWTLSVAMAFPPVFDVGTYKFIREEDQCIFEHRYFKANDTLGFMLMLAVLMAATHAVYGKLLLFEYRHRKMKPVQMVPAISQNWTFHGPGATGQAAANWIAGFGRGPMPPTLLGIRQNGHAASRRLLGMDEVKGEKQLGRMFYAITLLFLLLWSPYIVACYWRVFVKACAVPHRYLATAVWMSFAQAAVNPIVCFLLNKDLKKCLRTHAPCWGTGGAPAPREPYCVM | |||||||||||||||||||||||||
| 1 | 4iaqA | 0.16 | 0.21 | 0.90 | 2.38 | Download | ----------YIYQDSISLPWKVLLVMLLALITLATTLSNAFVIATVYRTRKLHTPANYLIASLAVTDLLVSILVMPISTMYTVTG-RWTLGQVVCDFWLSSDITCCTASIWHLCVIALDRYWAITDAVEYSAKRTPKRAAVMIALVWVFSISISLPPFFWRQASECVVNTD----HILYTVYSTVGAFYFPTLLLIALYGRIYVEARSRIADLEDNWETLNDNLKVIQKATPDSPEMKDFRHGFDILVGGKVKEAQAAAEQLKTTRNAYIQKYLLMAARERKATKTLGIILGAFIVCWLPFFIISLVMP------IHLAIFDFFTWLGYLNSLINPIIYTMSNEDFKQAFHKLIRFK--------------- | |||||||||||||||||||
| 2 | 5uenA | 0.17 | 0.23 | 0.92 | 4.36 | Download | ------------------SAFQAAYIGIEVLIALVSVPGNVLVIWAVKVNQALRDATFCFIVSLAVADVAVGALVIPLAILINIGPQ-TYFH--TCLMVACPVLILTQSSILALLAIAVDRYLRVKIPLRYKMVVTPRRAAVAIAGCWILSFVVGLTPMFGWNNLSAVERAWKVISMEYMVYFNFFVWVLPPLLLMVLIYLEVFYLIRKQLADLEDNWETLNDNVKDALTKMRAAALDAPEMKDFRGKVKEAQAAAEQLKTTRNAYIQKYLERARSTLQKELKIAKSLALILFLFALSWLPLHILNCITLFCPSCHKPSILTYIAIFLTHGNSAMNPIVYAFRIQKFRVTFLKIWNDHFRCQPLE-------- | |||||||||||||||||||
| 3 | 5uenA | 0.17 | 0.23 | 0.94 | 3.23 | Download | ----------------SISAFQAAYIGIEVLIALVSVPGNVLVIWAVKVNQALRDATFCFIVSLAVADVAVGALVIPLAILINIGP-QTYF--HTCLMVACPVLILTQSSILALLAIAVDRYLRVKIPLRYKMVVTPRRAAVAIAGCWILSFVVGLTPMFGWNNLSASMGEPVIISMEYMVYFNFFVWVLPPLLLMVLIYLEVFYLIRKQLADLEDNNVKDALTKMRAAALDAPEMKDFRHGFDILVGQGKVKEAQAAAEQLKNAYIQKYLERARSTLQKELKIAKSLALILFLFALSWLPLHILNCITLFCPSCHKPSILTYIAIFLTHGNSAMNPIVYAFRIQKFRVTFLKIWNDHFRCQPLEVLF----- | |||||||||||||||||||
| 4 | 4ib4 | 0.20 | 0.24 | 0.92 | 1.56 | Download | ---------------EEQGNKLHWAALLILMVIIPTIGGNTLVILAVSLEKKLQYATNYFLMSLAVADLLVGLFVMPIALLTIMFEAMWPLPLVLCPAWLFLDVLFSTASIWHLCAISVDRYIAIKKPIQANQYNSRATAFIKITVVWLISIGIAIPVIKGIETN---PNNITCVLTKERFGDFMLFGSLAAFFTPLAIMIVTYFLTIHALQKKAADLEDNWETLNDNLKVIEKADNAAQRAAALDAQKNEGKVKEAQAAAEQLKTTNAYIQKYLQTISNEQRASKVLGIVFFLFLLMWCPFFITNITLVLCDSCNTLQMLLEIFVWIGYVSSGVNPLVYTLFNKTFRDAFGRYITCNYR------------- | |||||||||||||||||||
| 5 | 3uon | 0.18 | 0.22 | 0.90 | 1.21 | Download | ------------------TFEVVFIVLVAGSLSLVTIIGNILVMVSIKVNRHLQTVNNYFLFSLACADLIIGVFSMNLYTLYTVIG-YWPLGPVVCDLWLALDYVVSNASVMNLLIISFDRYFCVTKPLTYPVKRTTKMAGMMIAAAWVLSFILWAPAILFWQFIVGTVEDGECYIQFFSNAAVTIAAFYLPVIIMTVLYWHISRASKSRINIFEMLRIDEGLRLKIYDELTKSPSAAGRNTNGVIGILRNAKLQQKRWDLAKSRNQTPNTWDAYPPPSREKKVTRTILAILLAFIITWAPYNVMVLINTFCAPC-IPNTVWTIGYWLCYINSTINPACYALCNATFKKTFKHLLM----------------- | |||||||||||||||||||
| 6 | 4ib4A | 0.18 | 0.24 | 0.92 | 3.00 | Download | ----------------EQGNKLHWAALLILMVIIPTIGGNTLVILAVSLEKKLQYATNYFLMSLAVADLLVGLFVMPIALLTIMFEAMWPLPLVLCPAWLFLDVLFSTASIWHLCAISVDRYIAIKKPIQANQYNSRATAFIKITVVWLISIGIAIPVPIKGIETNPNNITCVLTKERDFMLFGSLAAFFTPLAIMIVTYFLTIHALQKKAADLEDNWETLNDNLKVIEKADNADILVGQIDDALKLANEGKVKEAQAAAEQLKTTRNAYIQKYLQTISNEQRASKVLGIVFFLFLLMWCPFFITNITLVLCDSCTTLQMLLEIFVWIGYVSSGVNPLVYTLFNKTFRDAFGRYITCNYR------------- | |||||||||||||||||||
| 7 | 4ib4 | 0.21 | 0.24 | 0.90 | 1.72 | Download | ---------------------LHWAALLILMVIIPTIGGNTLVILAVSLEKKLQYATNYFLMSLAVADLLVGLFVMPIALLTIMFEAMWPLPLVLCPAWLFLDVLFSTASIWHLCAISVDRYIAIKKPIQANQYNSRATAFIKITVVWLISIGIAIPPIKGIETN---PNNITCVLTKERMLFGSLAAFFTPLAIMIVTYFLTIHALQKKAADLEDNWETLNDNKADNAAQVKDALTKMRAALKDRHGFNEGKVKEAQAAAEQLKTRNAYIQKYLQTISNEQRASKVLGIVFFLFLLMWCPFFITNITLVLCDSCNTLQMLLEIFVWIGYVSSGVNPLVYTLFNKTFRDAFGRYITCNY-------------- | |||||||||||||||||||
| 8 | 2rh1A | 0.18 | 0.24 | 0.84 | 4.28 | Download | -----------------DEVWVVGMGIVMSLIVLAIVFGNVLVITAIAKFERLQTVTNYFITSLACADLVMGLAVVPFGAAHILMK-MWTFGNFWCEFWTSIDVLCVTASIETLCVIAVDRYFAITSPFKYQSLLTKNKARVIILMVWIVSGLTSFLPIQMHWYRATHAEETCC--DFFTNQAYAIASSIVSFYVPLVIMVFVYSRVFQEAKRQLQKRWDEAAVNLAKSRWYNQTPNRAKRVITTFRTGTWDAYKFCL---------------------KEHKALKTLGIIMGTFTLCWLPFFIVNIVHVIQDNL-IRKEVYILLNWIGYVNSGFNPLIYCR-SPDFRIAFQELL-CL--------------- | |||||||||||||||||||
| 9 | 4ib4A | 0.18 | 0.24 | 0.92 | 2.65 | Download | ---------------EEQGNKLHWAALLILMVIIPTIGGNTLVILAVSLEKKLQYATNYFLMSLAVADLLVGLFVMPIALLTIMFEAMWPLPLVLCPAWLFLDVLFSTASIWHLCAISVDRYIAIKKPIQANQYNSRATAFIKITVVWLISIGIAIPVPIKGIETNPNNITCVLTKERDFMLFGSLAAFFTPLAIMIVTYFLTIHALQKKAADLEDNWETLNDNLKVIEKADNAAQVKDALTKMRAAALDGKVKEAQAAAEQLKTTRNAYIQKYLQTISNEQRASKVLGIVFFLFLLMWCPFFITNITLVLCDSCNTLQMLLEIFVWIGYVSSGVNPLVYTLFNKTFRDAFGRYITCNYR------------- | |||||||||||||||||||
| 10 | 4nc3A | 0.18 | 0.23 | 0.92 | 2.76 | Download | ---------------EEQGNKLHWAALLILMVIIPTIGGNTLVILAVSLEKKLQYATNYFLMSLAVADLLVGLFVMPIALLTIMFEAMWPLPLVLCPAWLFLDVLFSTASIWHLCAISVDRYIAIKKPIQANQYNSRATAFIKITVVWLISIGIAIPVPIKGIETVDNPNNITCVLTKDFMLFGSLAAFFTPLAIMIVTYFLTIHALQKKAADLEDNWETLNDNLKVIEKADNAAQVKDALTKMRAKANEGKVKEAQAAAEQLKTTRNAYIQKYLQTISNEQRASKVLGIVFFLFLLMWCPFFITNITLVLCDSCNTLQMLLEIFVWIGYVSSGVNPLVYTLFNKTFRDAFGRYITCNYR------------- | |||||||||||||||||||
| ||||||||||||||||||||||||||
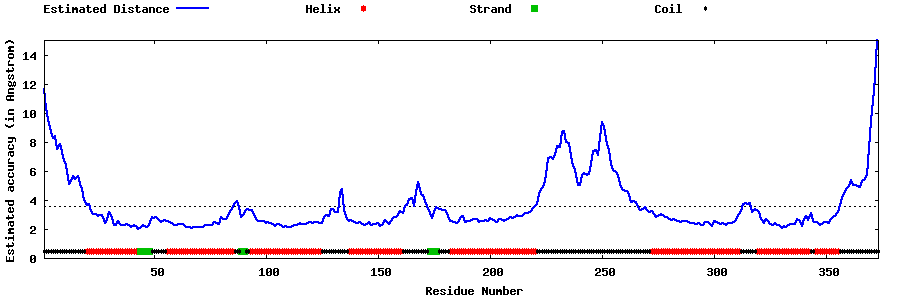
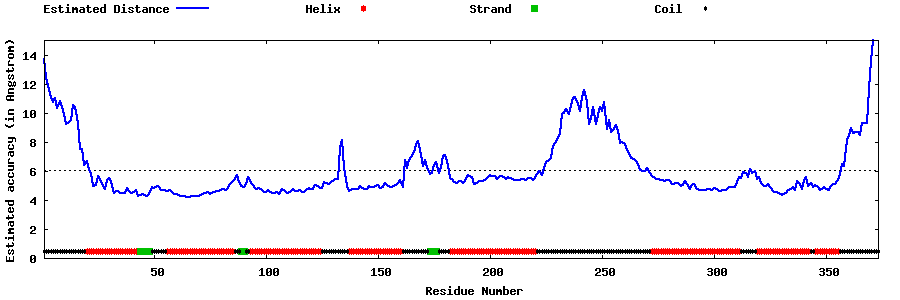
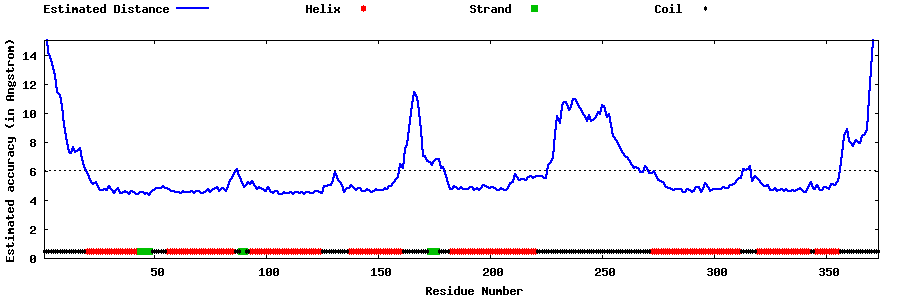
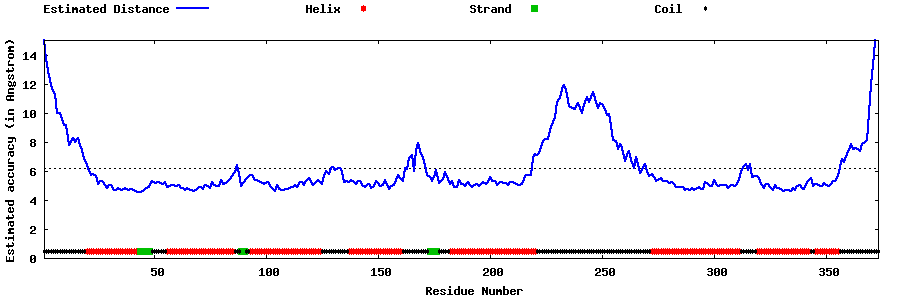
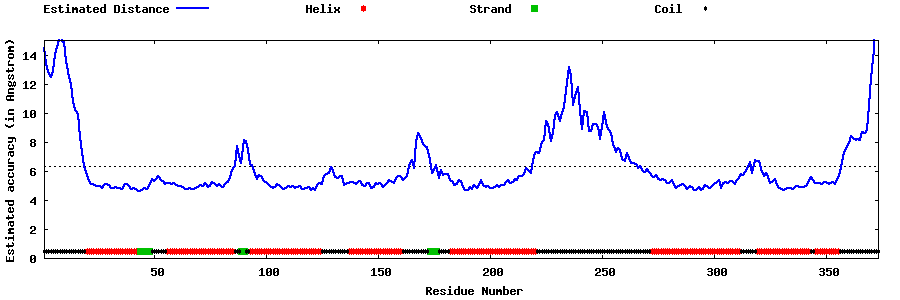
| Generated 3D models | Estimated local accuracy of models | ||
 | |||
|
|
|||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||