

| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| MESALPSIFTLVIIAEFIIGNLSNGFIVLINCIDWVSKRELSSVDKLLIILAISRIGLIWEILVSWFLALHYLAIFVSGTGLRIMIFSWIVSNHFNLWLATIFSIFYLLKIASFSSPAFLYLKWRVNKVILMILLGTLVFLFLNLIQINMHIKDWLDRYERNTTWNFSMSDFETFSVSVKFTMTMFSLTPFTVAFISFLLLIFSLQKHLQKMQLNYKGHRDPRTKVHTNALKIVISFLLFYASFFLCVLISWISELYQNTVIYMLCETIGVFSPSSHSFLLILGNAKLRQAFLLVAAKVWAKR | |
| CCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCHHHCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSSCCCCCCHHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCSSSSSSSSCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHCCCHHHHHHHHHHHHHCCCCC | |
| 986899999999999999999999999999999999088898478999999998688211799989999948325068643442588999986899999999999982785189980899999874284599999999999999999985510003677775136678741221059999999999999999999999999999999998466779979999967999999999999999999999999999998666969999999998776998489996197089999999998710179 |
| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| MESALPSIFTLVIIAEFIIGNLSNGFIVLINCIDWVSKRELSSVDKLLIILAISRIGLIWEILVSWFLALHYLAIFVSGTGLRIMIFSWIVSNHFNLWLATIFSIFYLLKIASFSSPAFLYLKWRVNKVILMILLGTLVFLFLNLIQINMHIKDWLDRYERNTTWNFSMSDFETFSVSVKFTMTMFSLTPFTVAFISFLLLIFSLQKHLQKMQLNYKGHRDPRTKVHTNALKIVISFLLFYASFFLCVLISWISELYQNTVIYMLCETIGVFSPSSHSFLLILGNAKLRQAFLLVAAKVWAKR | |
| 743323331132133133313313220221011000453303000000112140333112113332000000031134331001000101221030002002310000000022311100101330431001101212233232313131303132443431212312033133000111111133333233313323330220022023202333444412314102200200020223233223113310112134432110211031100100103143240430130012002302138 |
| Rank | PDB Hit | Iden1 | Iden2 | Cov. | Norm. Z-score | Download Align. | 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | | | | | | | | | | | | | | | | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sec.Str Seq | CCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCHHHCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSSCCCCCCHHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCSSSSSSSSCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHCCCHHHHHHHHHHHHHCCCCC MESALPSIFTLVIIAEFIIGNLSNGFIVLINCIDWVSKRELSSVDKLLIILAISRIGLIWEILVSWFLALHYLAIFVSGTGLRIMIFSWIVSNHFNLWLATIFSIFYLLKIASFSSPAFLYLKWRVNKVILMILLGTLVFLFLNLIQINMHIKDWLDRYERNTTWNFSMSDFETFSVSVKFTMTMFSLTPFTVAFISFLLLIFSLQKHLQKMQLNYKGHRDPRTKVHTNALKIVISFLLFYASFFLCVLISWISELYQNTVIYMLCETIGVFSPSSHSFLLILGNAKLRQAFLLVAAKVWAKR | |||||||||||||||||||||||||
| 1 | 4djhA | 0.12 | 0.15 | 0.91 | 1.41 | Download | -SPAIPVIITAVYSVVFVVGLVGNSLVMFVIIR----TKMKTATNIYIFNLALADALVTTTMPFQSTVYLMN-SWPFGDVLCKIVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKA----LDFRTPLKAKIINICIWLLSSSVGISAIVLGGTKVREDVDVIECSLQFPDYSWWDLFMKICVFIFAVIPVLIIIVCYTLMILRLKSVRLL------------DRNLRRITRLVLVVVAVFVVCWTPIHIFILVEALGALSSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCFP----- | |||||||||||||||||||
| 2 | 5tjvA | 0.09 | 0.14 | 0.90 | 2.19 | Download | LNPSQQLAIAVLSLTLGTFTVLENLLVLCVILHS--RSLRCRPSYHFIGSLAVADLLG-SVIFVYSFIDFHVFHRKDSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLA----YKRIVTRPKAVVAFCLMWTIAIVIAVLPLLGWNCEKLQSVCSDI-------FPHIDETYLMFWIGVTSVLLLFIVYAYMYILWKAGKRAMSFSD--------QARMDIRLAKTLVLILVVLIICWGPLLAIMVYDVFGKMTVFAFCSMLCLLNSTVNPIIYALRSKDLRHAFRSMF------- | |||||||||||||||||||
| 3 | 4djhA | 0.11 | 0.15 | 0.91 | 2.51 | Download | -SPAIPVIITAVYSVVFVVGLVGNSLVMFVIIRY--TKMK-TATNIYIFNLALADALVTTT--MPFQSTVYLMNSWFGDVLCKIVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKAL----DFRTPLKAKIINICIWLLSSSVGISAIVLGGTKVREDVDVIECSLQFDYSWWDLFMKICVFIFAFVIPVLIIIVCYTLMILRLKSVRLL-------------RNLRRITRLVLVVVAVFVVCWTPIHIFILVEALGALSSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCFP----- | |||||||||||||||||||
| 4 | 4djh | 0.12 | 0.20 | 0.94 | 1.56 | Download | -SPAIPVIITAVYSVVFVVGLVGNSLVMFVIIR---YTKMKTATNIYIFNLALADALVTTTMPFQST-VYLMNSWPF-GDVLCIVLSIDYYNMFTSIFTLTMMSVDRYIAVC---HPVKAL-DFRTPLKAKIINICIWLLSSVGISAIVLGGKVDDVECSLQFPDDDYSWDLFMKI----CVFIFAFVIPVLIIIVCYTLMILRLKSVRLLSGNIFTDAYREKDRNLRRITRLVLVVVAVFVVCWTPIHIFILVALGAALSSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCFP----- | |||||||||||||||||||
| 5 | 5glh | 0.14 | 0.20 | 0.90 | 1.20 | Download | IKETFKYINTVVSCLVFVLGIIGNSTLLYIIYK--------NGPNILIASLALGDLLHIVIAIPINVYKLLAEDWPFGAEMCKLVPFIQKASVGITVLSLCALSIDRYRAVASW--------------WTAVEIVLIWVVSVVLAVPEAIGFDIITMDYKGSYLRICLLHPVQAFMQFDWWLFSFYFCLPLAITAFFYTLMTCEMLRKNEGLRLTWDAYLNDHLKQRREVAKTVFCLVLVFALCWLPLHLARILKLLYNNVLDYIGINMASLNSCANPIALYLVSKRFKNAFKSAL------- | |||||||||||||||||||
| 6 | 4djhA | 0.11 | 0.15 | 0.91 | 1.60 | Download | ---AIPVIITAVYSVVFVVGLVGNSLVMFVIIRY---TKMKTATNIYIFNLALADALVTTTMPFQSTVYLMN-SWPFGDVLCKIVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKAL----DFRTPLKAKIINICIWLLSSSVGISAIVLGGTKVREDVDVIECSLQFPYSWWDLFMKICVFIFAFVIPVLIIIVCYTLMILRLKSVRLLSD-----------RNLRRITRLVLVVVAVFVVCWTPIHIFILVEALGALSSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCFP----- | |||||||||||||||||||
| 7 | 3uon | 0.09 | 0.20 | 0.93 | 1.70 | Download | -----VVFIVLVAGSLSLVTIIGNILVMVSIKV---NRHLQTVNNYFLFSLACADLIIGVFSMNLYTLYTVIGYWPLGPVVCDLWLALDYVVSNASVMNLLIISFDRYFCVTKPLTYPVKRTTKMAGMMIAAAWVLSFILWAPAILFWQFIVGVRTV-EDGECYIQ--FF---SNAAVTFGTAIAAFYLPVIIMTVLYWHISRASKSRINIFEMGATWPPSREKKVTRTILAILLAFIITWAPYNVMVLINTFCAPCIPNTVWTIGYWLCYINSTINPACYALCNATFKKTFKHLLM------ | |||||||||||||||||||
| 8 | 4buoA | 0.17 | 0.20 | 0.91 | 2.64 | Download | TDIYSKVLVTAIYLALFVVGTVGNSVTLFTLAR----KKSLSTVDYYLGSLALSDLLILLLAMPVELYNFIWVHWAFGDAGCRGYYFLRDACTYATALNVVSLSVELYLAICH-------PFKAKTRSRTKKFISAIWLASALLAIPMLFTMGLQNLSGDGTHLVCTPIVDTATLKVVIQVNTFMSFLFPMLVASILNTVIANKLTVMVHQ---------PGRVQALRRGVLVLRAVVIAFVVCWLPYHVRRLMFCYISDYFYMLTNALVYVSAAINPILYNLVSANFRQVFLSTL------- | |||||||||||||||||||
| 9 | 4djhA | 0.12 | 0.15 | 0.91 | 1.52 | Download | -SPAIPVIITAVYSVVFVVGLVGNSLVMFVIIR----TKMKTATNIYIFNLALADALVTTTMPFQSTVYLMNS--PFGDVLCKIVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKA----LDFRTPLKAKIINICIWLLSSSVGISAIVLGGTKVREDVDVIECSLQFPDDDYSLFMKICVFIFAFVIPVLIIIVCYTLMILRLKSVRLL-----------SDRNLRRITRLVLVVVAVFVVCWTPIHIFILVEALGSASSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCFP----- | |||||||||||||||||||
| 10 | 4ea3A | 0.10 | 0.20 | 0.89 | 1.70 | Download | -PLGLKVTIVGLYLAVCVGGLLGNCLVMYVILR---HTKMKTATNIYIFNLALADTLVLLTLPFQGTDILLGFWPF-GNALCKTVIAIDYYNMFTSTFTLTAMSVDRYVAICHP-----------TSSKAQAVNVAIWALASVVGVPVAIMGSAQVEDEEIECLVEIPTPQDYWGPVFAICIFLFSFIVPVLVISVCYSLMIRRLRG------VRLLSGSREKDRNLRRITRLVLVVVAVFVGCWTPVQVFVLAQGLGVQAILRFCTALGYVNSCLNPILYAFLDENFKACFR---------- | |||||||||||||||||||
| ||||||||||||||||||||||||||
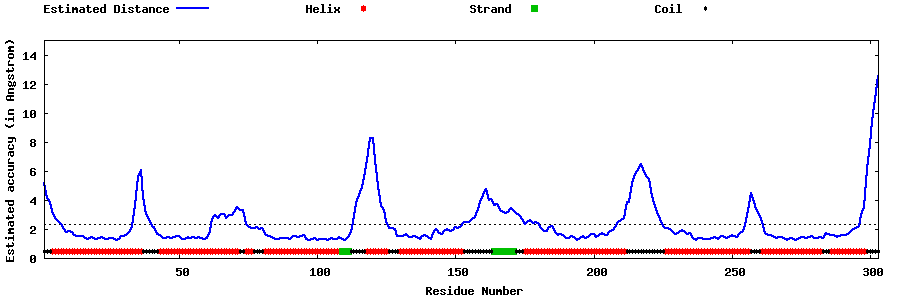
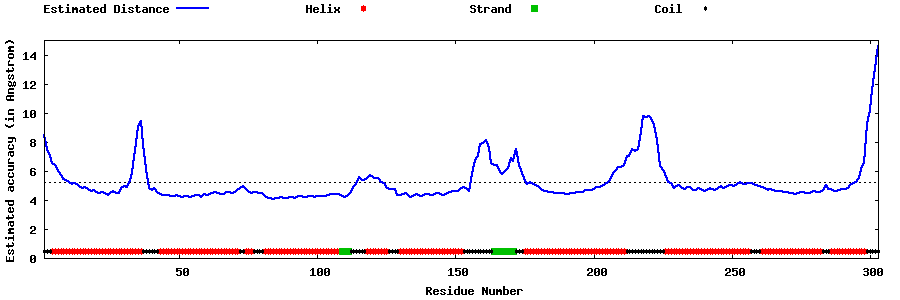
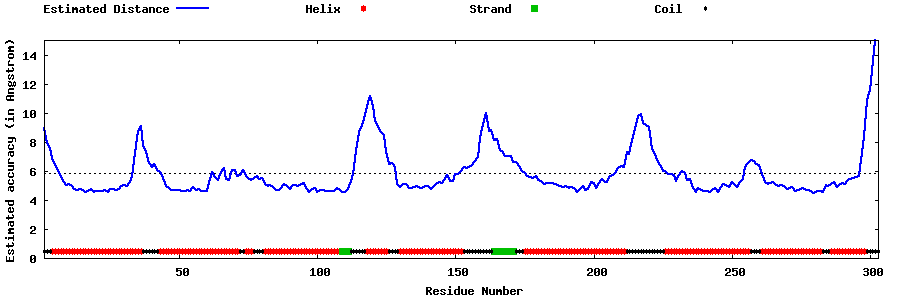
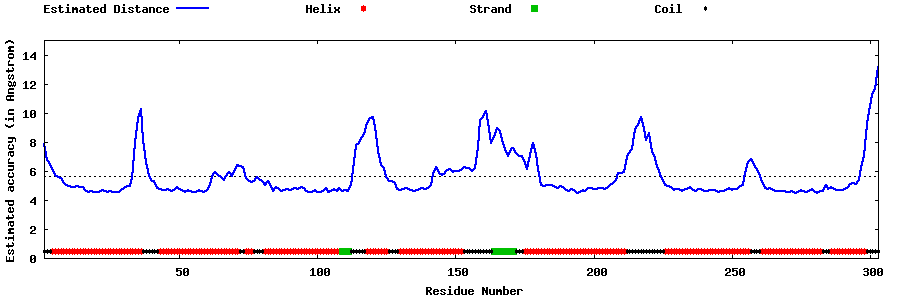
| Generated 3D models | Estimated local accuracy of models | ||
 | |||
|
|
|||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||