


|
This page contains 3D structural models (Version 3, built on Aug 2014) of 1,026 putative G protein-coupled receptors (GPCRs) in the human genome generated by the GPCR-I-TASSER pipeline. The most recent (Version 4, built on June 2018) is now available as part of the GPCR-EXP database. In GPCR-I-TASSER, the GPCR sequences are first threaded through the GPCR template library to identify muliple structure templates by the LOMETS programs. When close homolgous templates are identified, full-length models will be constructed by the I-TASSER based fragment assembly simulations, assisted by a GPCR and membrane specific force field and spatial restraints collected from mutagenesis experiments in GPCR-RD. In case that homologous templates are not available, an ab initio folding procedure is used to assemble the 7-TM-helix bundle from scratch, followed by the GPCR-I-TASSER fragment reassembly simulations. For multiple domain GPCRs, structural models are built by GPCR-I-TASSER for each domain separately which are then reassembly by the I-TASSER approach. All the models are finally subjected to FG-MD for fragment-guided molecular dynamic simulation refinements. Note:
|
Other GPCR-related resources
|
| HG ID | UniProt ID | Length | C-score | Estimated TM-score |
Estimated RMSD |
Top 5 models |
| HG0030 | P0C628 | 307 | -0.13 | 0.7 ± 0.12 | 6.5 ± 3.9 | 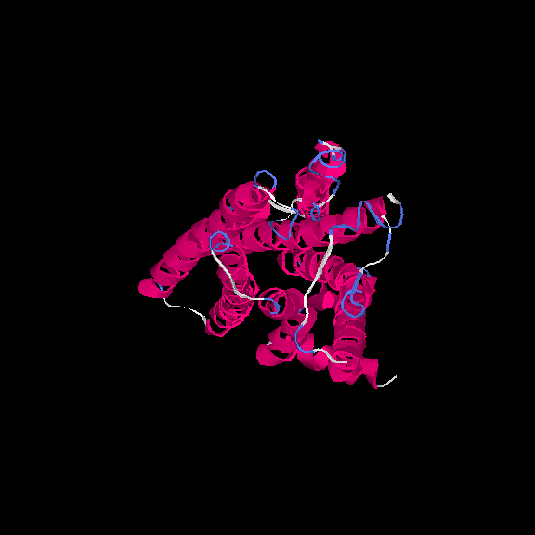   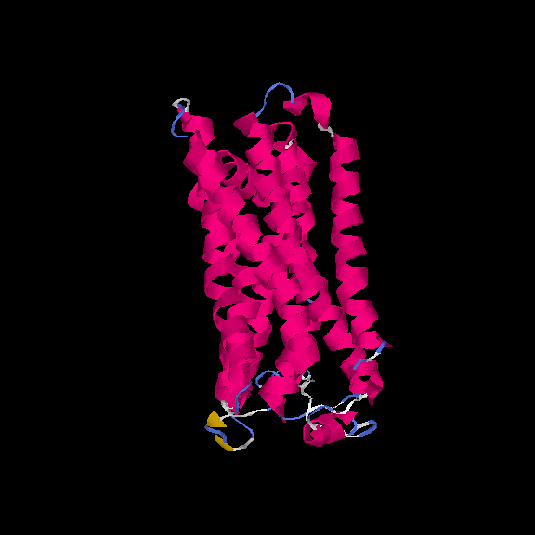 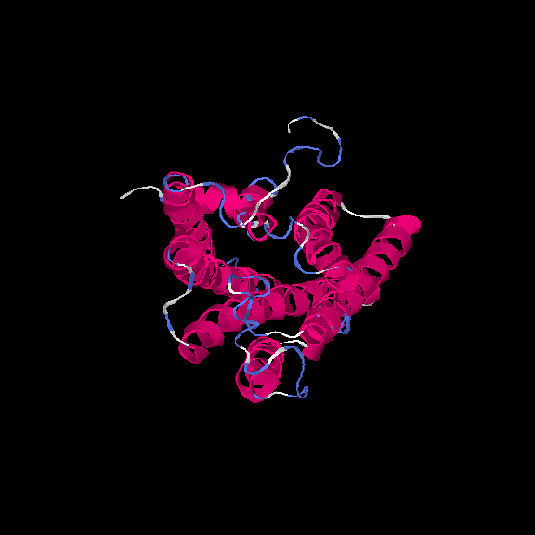 |
| HG0031 | Q8N127 | 324 | -0.3 | 0.67 ± 0.12 | 7 ± 4.1 |   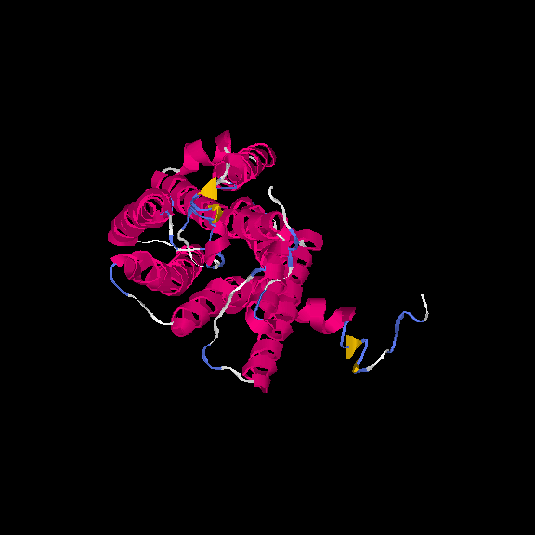  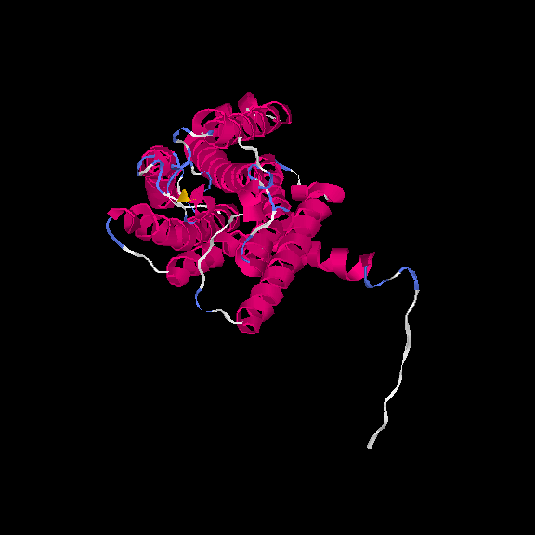 |
| HG0032 | Q9NYV7 | 291 | 0.04 | 0.72 ± 0.11 | 6 ± 3.7 | 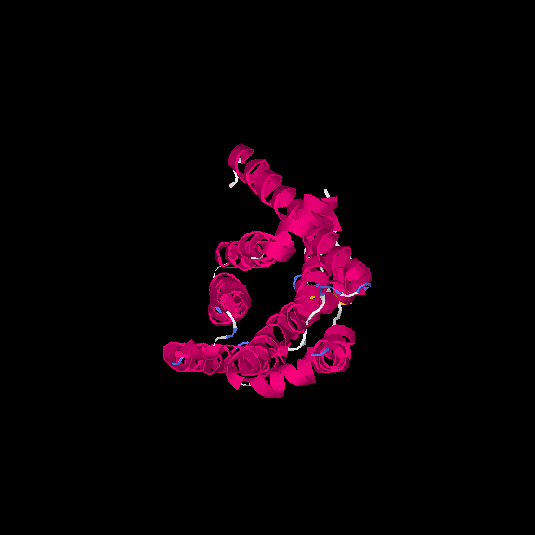 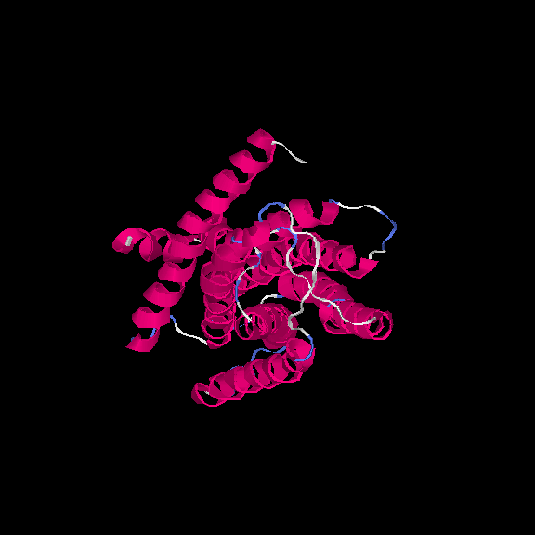 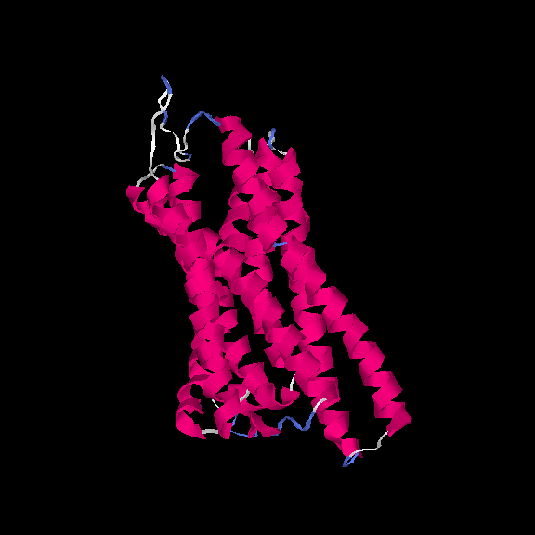   |
| HG0033 | P46093 | 362 | -1.14 | 0.57 ± 0.15 | 9.2 ± 4.6 |      |
| HG0034 | P18825 | 462 | -1.41 | 0.54 ± 0.15 | 9.99 ± 4.6 |      |
| HG0035 | B9EIS6 | 323 | -0.34 | 0.67 ± 0.13 | 7.1 ± 4.1 |      |
| HG0036 | P46663 | 353 | -0.16 | 0.69 ± 0.12 | 6.9 ± 4.1 |      |
| HG0037 | Q8NH74 | 314 | -0.24 | 0.68 ± 0.12 | 6.8 ± 4 |   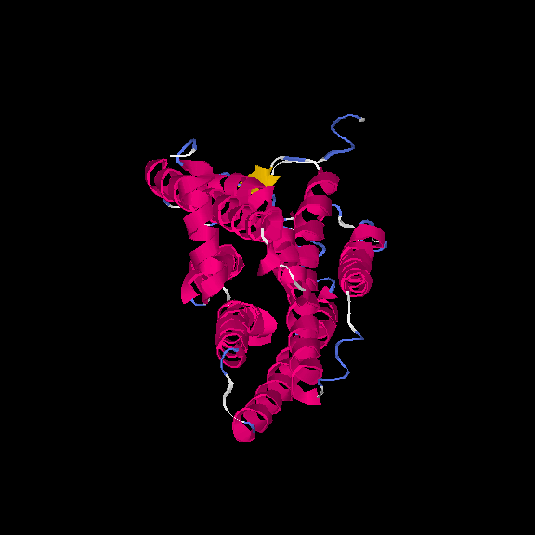 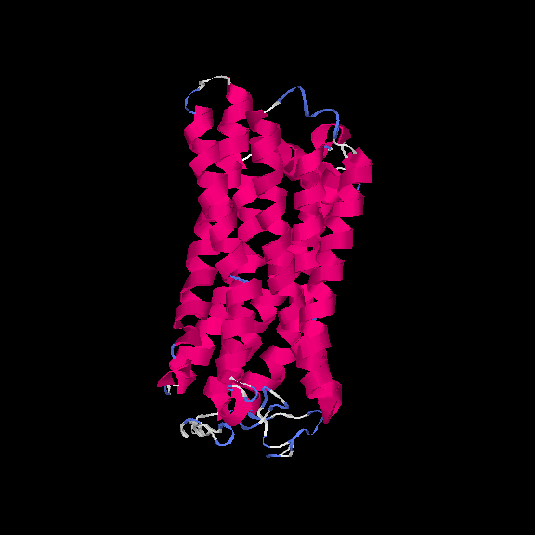 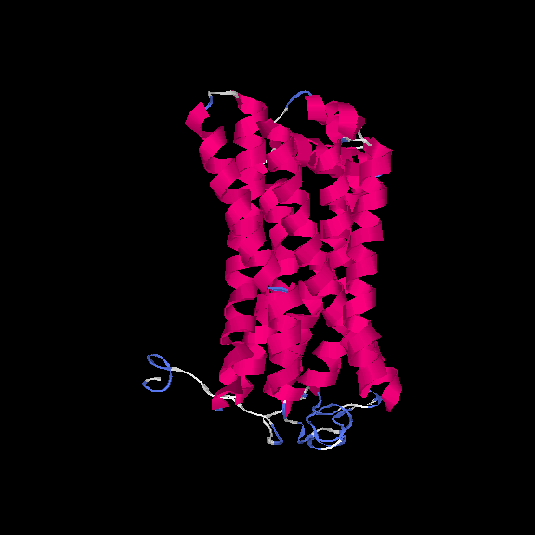 |
| HG0038 | O60755 | 368 | -0.69 | 0.63 ± 0.14 | 8.2 ± 4.5 | 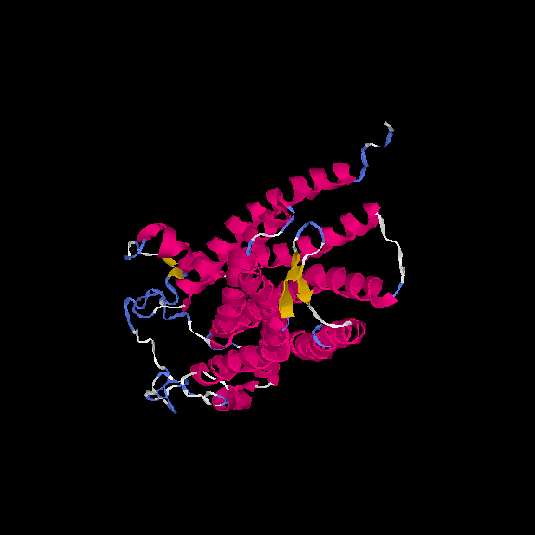 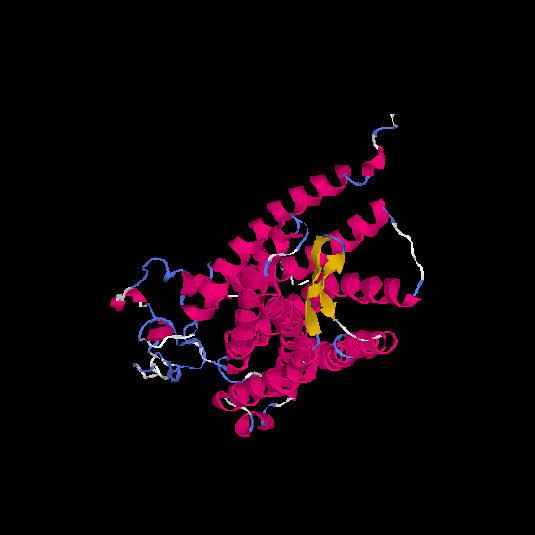 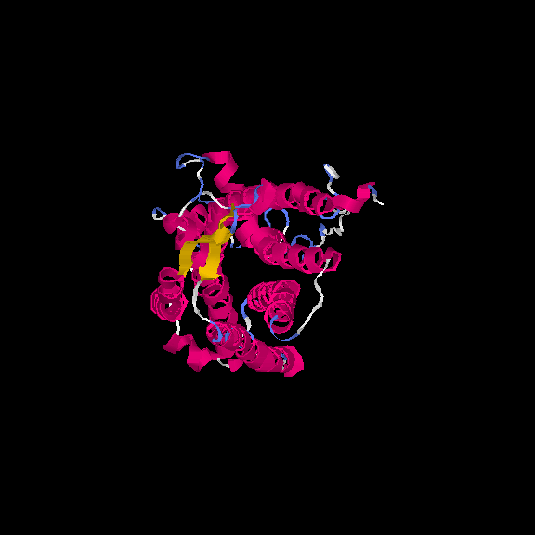 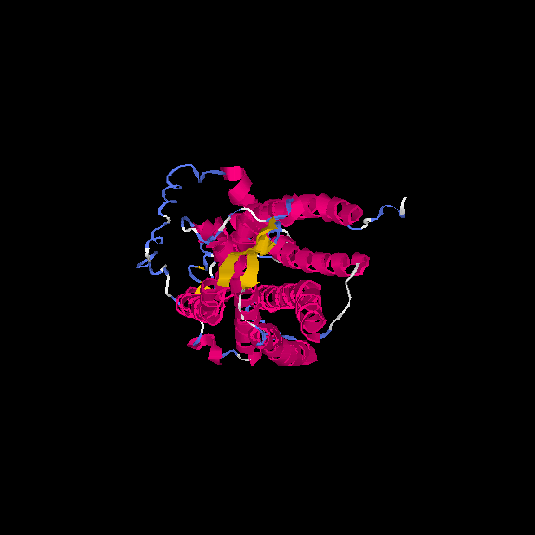 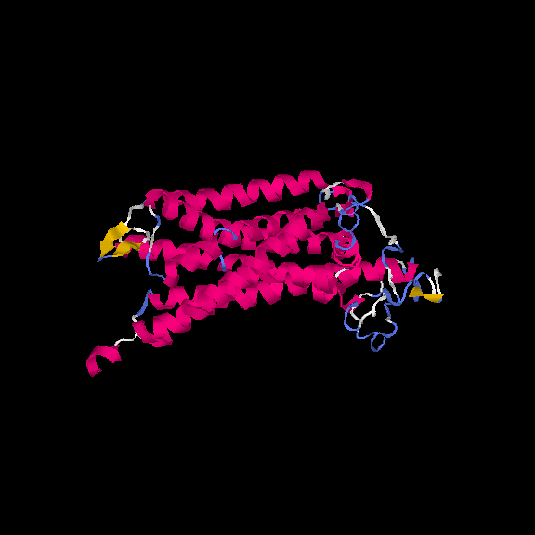 |
| HG0039 | P0C623 | 313 | -0.14 | 0.7 ± 0.12 | 6.6 ± 4 |      |
| HG0040 | Q8NH09 | 359 | -1.11 | 0.58 ± 0.14 | 9.1 ± 4.6 |      |
yangzhanglab![]() umich.edu
| (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
umich.edu
| (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218