


Purpose:
Atomic structure prediction and fitting most times are independent procedures. This makes some predicted model impossible to fit into a density map when only rigid-body movements are employed. MVP-Fitting can flexibly move only one domain of the whole model during the fitting process and still keep all the geometric restraints of bond lengths, bond angles and steric clashes for the model.
Downloadable:
User Guide (examples of input files: flexible.txt, para.txt, modelnames.txt)
Illustration:
1. Fit the predicted multiple-domain model of beta subunit into the phosphorylase kinase EM data.
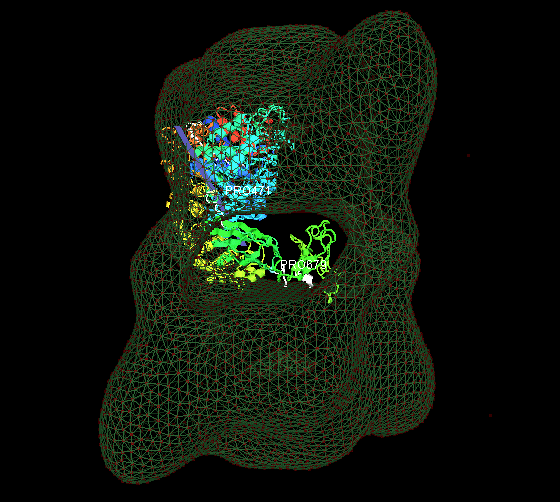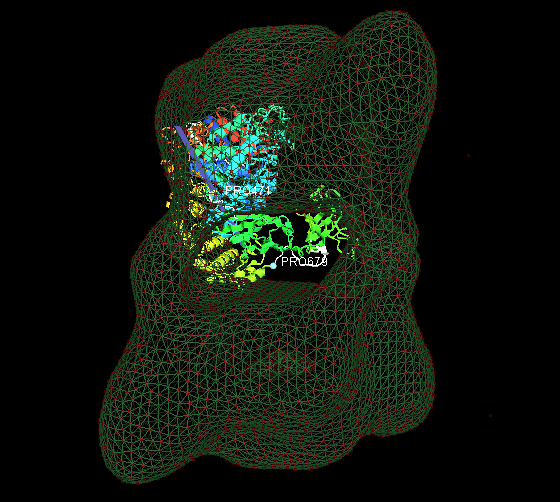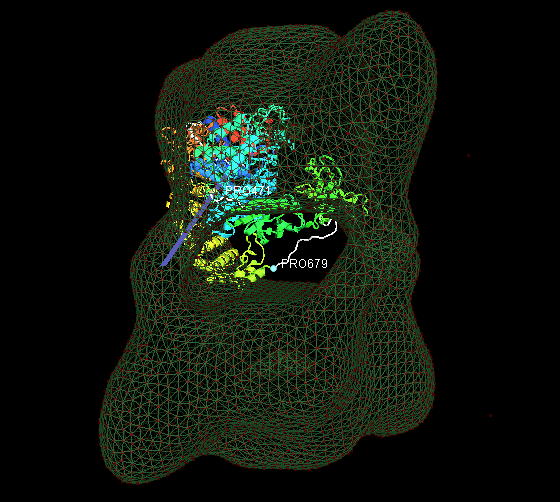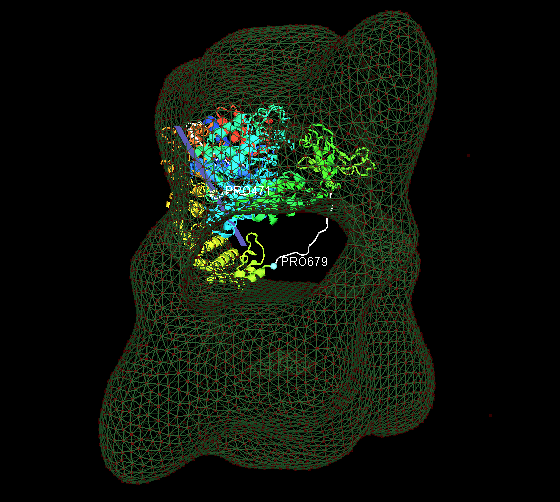
2. Fit 18 identical monomers(PDB ID: 1a6dA) into the chaperonin EM data.
3. Fit the multiple-chain crystal strcuture(PDB ID: 1aon) into the chaperonin EM data(EMDB ID: 1180).
4. Fit complex strcuture(PDB ID: 3iyd) which contains both protein and DNA chains with the intact activator-dependent transcription initiation EM data(EMDB ID: 5127). EM isosurfaces are shown with different density thresholds.
5. Fit 2 multiple-chain crystal strcutures(PDB ID: 1yjd) into the CD28-Fc fusion complexed with 5.11A1 monoclonal antibody EM data(EMDB ID: 1219).
yangzhanglab![]() umich.edu
| (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
umich.edu
| (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218