

| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 340 | |
| | | | | | | | | | | | | | | | | | | |
| MAGNCSWEAHPGNRNKMCPGLSEAPELYSRGFLTIEQIAMLPPPAVMNYIFLLLCLCGLVGNGLVLWFFGFSIKRNPFSIYFLHLASADVGYLFSKAVFSILNTGGFLGTFADYIRSVCRVLGLCMFLTGVSLLPAVSAERCASVIFPAWYWRRRPKRLSAVVCALLWVLSLLVTCLHNYFCVFLGRGAPGAACRHMDIFLGILLFLLCCPLMVLPCLALILHVECRARRRQRSAKLNHVILAMVSVFLVSSIYLGIDWFLFWVFQIPAPFPEYVTDLCICINSSAKPIVYFLAGRDKSQRLWEPLRVVFQRALRDGAELGEAGGSTPNTVTMEMQCPPGNAS | |
| CCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCHHHCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHSSSSSCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCHHHSSSSSSSSSHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCCCCCCCCCCCCCCCCCC | |
| 9999988789998777899998886556789776433024416799999999999999999999899988457786879999999999999999879999999955899637699999999999999999999999999999999989875467872378999999999999999889988245306899625899999999999999999999999999999997057888786303303567887636556899999999998621668999999999999999999998157988799999999999985758888788998899988677779998999 |
| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 340 | |
| | | | | | | | | | | | | | | | | | | |
| MAGNCSWEAHPGNRNKMCPGLSEAPELYSRGFLTIEQIAMLPPPAVMNYIFLLLCLCGLVGNGLVLWFFGFSIKRNPFSIYFLHLASADVGYLFSKAVFSILNTGGFLGTFADYIRSVCRVLGLCMFLTGVSLLPAVSAERCASVIFPAWYWRRRPKRLSAVVCALLWVLSLLVTCLHNYFCVFLGRGAPGAACRHMDIFLGILLFLLCCPLMVLPCLALILHVECRARRRQRSAKLNHVILAMVSVFLVSSIYLGIDWFLFWVFQIPAPFPEYVTDLCICINSSAKPIVYFLAGRDKSQRLWEPLRVVFQRALRDGAELGEAGGSTPNTVTMEMQCPPGNAS | |
| 8664331423343323323333333332333323243243322301112202200231232102000001023421000000000010000000001010001023321110100021002113301310010000000000000000010232133110000001002100100000000000124544210011001122322323231011001000000010334635443000000000020001012221100001001313131012001000123002301010000340152024002200330043444456446434445444354466668 |
| Rank | PDB Hit | Iden1 | Iden2 | Cov. | Norm. Z-score | Download Align. | 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 340 | | | | | | | | | | | | | | | | | | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sec.Str Seq | CCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCHHHCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHSSSSSCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCHHHSSSSSSSSSHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCCCCCCCCCCCCCCCCCC MAGNCSWEAHPGNRNKMCPGLSEAPELYSRGFLTIEQIAMLPPPAVMNYIFLLLCLCGLVGNGLVLWFFGFSIKRNPFSIYFLHLASADVGYLFSKAVFSILNTGGFLGTFADYIRSVCRVLGLCMFLTGVSLLPAVSAERCASVIFPAWYWRRRPKRLSAVVCALLWVLSLLVTCLHNYFCVFLGRGAPGAACRHMDIFLGILLFLLCCPLMVLPCLALILHVECRARRRQRSAKLNHVILAMVSVFLVSSIYLGIDWFLFWVFQIPAPFPEYVTDLCICINSSAKPIVYFLAGRDKSQRLWEPLRVVFQRALRDGAELGEAGGSTPNTVTMEMQCPPGNAS | |||||||||||||||||||||||||
| 1 | 5o9hA | 0.25 | 0.24 | 0.82 | 2.84 | Download | -------------------------------------NTLRVPDILALVIFAVVFLVGVLGNALVVWVTAFEAKRTINAIWFLNLAVADFLACLALPALFTSIVQHHHWPFGGAACSILPSLILLNMYASILLLATISADRFLLVFKPAWCQRFRGAGLAWILCAVAWGLALLLTIPSALYRVVREEYFPPDKRRERAVAIVRLVLGFLWPLLTLTICYTFILLRTWSARETRSTKTLKVVVAVVASFFIFWLPYQVTGIMMSFLEPSLKKLDSLCVSFAYINCCINPIIYVVAGQGFQKSLPELLREVLTE----ESVVR---------------------- | |||||||||||||||||||
| 2 | 4xnwA | 0.19 | 0.22 | 0.80 | 3.91 | Download | ------------------------------SSFKCALTKTGFQFYYLPAVYILVFIIGFLGNSVAIWMFVFHMKWSGISVYMFNLALADFLYVLTLPALIFYYFNKTDWIFGDAMCKLQRFIFHVNLYGSILFLTCISAHRYSGVVYPLKSLGRLKKKNAICISVLVWLIVVVAISPILFYSGTGVRKNKTITCSYFIYSMCTTVAMFCVPLVLILGCYGLIVRALIYKEEPLRRKSIYLVIIVLTVFAVSYIPFHVMKTMNLRARLDVYATYQVTRGLASLNSCVNPILYFLAGDTFRRRLSR--------------------------------------- | |||||||||||||||||||
| 3 | 4n6hA | 0.17 | 0.22 | 0.81 | 3.43 | Download | -------------------------------SPGARSASSLALAIAITALYSAVCAVGLLGNVLVMFGIVRYTKKTATNIYIFNLALADALATSTLPFQSAKYLMET-WPFGELLCKAVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKALDFRTPAKAKLINICIWVLASGVGVPIMVMAVTRPRD-GAVVCMDTVTKICVFLFAFVVPILIITVCYGLMLLRLRSVRLLSLRRITRMVLVVVGAFVVCWAPIHIFVIVWTLVDIDVVAALHLCIALGYANSSLNPVLYAFLDENFKRCFRQLCRKPCG-------------------------------- | |||||||||||||||||||
| 4 | 2ziy | 0.13 | 0.22 | 0.94 | 1.53 | Download | LRDNETWWYNPSI--------IV---HPHWRE--FD-QVPDAVYYSLGIFIGICGIIGCGGNGIVIYLFTKTKSQTPANMFIINLAFSDFTFSLVFPLMTISCFLKKWIFGFAACKVYGFIGGIFGFM-SIMTMAMISIDRYNVIGRPMAASKKMSHRRAFIMIIFVWLWSVLWAIGPIFGWGAYTLEGVLCNCSTRSNILCMFILGFFGPILIIFFCYFNIVMSVSNHEKNAEMRLAKISIVIVSQFLLS-------WSPYAVVALLAQFGAQLPVMFAKASAIHNPMIYSVSHPKFREAISQTFPWVLTCCQFDDKETDDKDAETEIPAGESSDAAPSADA | |||||||||||||||||||
| 5 | 4djh | 0.18 | 0.21 | 0.78 | 1.15 | Download | ---------------------------------------SPAIPVIITAVYSVVFVVGLVGNSLVMFVIIRYTMKTATNIYIFNLALADALVTTTMPFQSTVYLM-NSWPFGDVLCKIVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKALDFRTPLKAKIINICIWLLSSSVGISAIVLGGTKVREVDVIECSLLFMKICVFIFAFVIPVLIIIVCYTLMILRLKSVRDRNLRRITRLVLVVVAVFVVCWTPIHIFILVEALGS-AALSSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCFP----------------------------------- | |||||||||||||||||||
| 6 | 5o9hA | 0.25 | 0.24 | 0.81 | 3.03 | Download | --------------------------------------TLRVPDILALVIFAVVFLVGVLGNALVVWVTAFEAKRTINAIWFLNLAVADFLACLALPALFTSIVQHHHWPFGGAACSILPSLILLNMYASILLLATISADRFLLVFKPAWCQRFRGAGLAWILCAVAWGLALLLTIPSALYRVVREEYVDHDKRRERAVAIVRLVLGFLWPLLTLTICYTFILLRTWSARETRSTKTLKVVVAVVASFFIFWLPYQVTGIMMSFLEPLLKKLDSLCVSFAYINCCINPIIYVVAGQGFQKSLPELLREVLTESVVR--------------------------- | |||||||||||||||||||
| 7 | 3uon | 0.17 | 0.22 | 0.77 | 1.69 | Download | -------------------------------------------VVFIVLVAGSLSLVTIIGNILVMVSIKVRHLQTVNNYFLFSLACADLIIGVFSMNLYTLYTVIGYWPLGPVVCDLWLALDYVVSNASVMNLLIISFDRYFCVTKPLTYPVKRTTKMAGMMIAAAWVLSFILWAPAILFWQFIVDGECYIQFFNAAVTFGTAIAAFYLPVIIMTVLYWHISRASKYPPPSREKKVTRTILAILLAFIITWAPYNVMVLINTFCAPIPNTVWTIGYWLCYINSTINPACYALCNATFKKTFKHLL------------------------------------- | |||||||||||||||||||
| 8 | 4n6hA | 0.17 | 0.22 | 0.82 | 3.37 | Download | ------------------------------GSPGARSASSLALAIAITALYSAVCAVGLLGNVLVMFGIVRYTKKTATNIYIFNLALADALATSTLPFQSAKYLMET-WPFGELLCKAVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKALDFRTPAKAKLINICIWVLASGVGVPIMVMAVTRPRDGSPSWYWDTVTKICVFLFAFVVPILIITVCYGLMLLRLRSVRLLSGRRITRMVLVVVGAFVVCWAPIHIFVIVWTLVDPLVVAALHLCIALGYANSSLNPVLYAFLD----ENFKRCFRQLCRKPCG---------------------------- | |||||||||||||||||||
| 9 | 5o9hA | 0.26 | 0.24 | 0.81 | 2.87 | Download | -------------------------------------NTLRVPDILALVIFAVVFLVGVLGNALVVWVTAFEAKRTINAIWFLNLAVADFLACLALPALFTSIVQHHHWPFGGAACSILPSLILLNMYASILLLATISADRFLLVFKPAWCQRFRGAGLAWILCAVAWGLALLLTIPSALYRVVREEYFPPDKRRERAVAIVRLVLGFLWPLLTLTICYTFILLRTWSARETRSTKTLKVVVAVVASFFVTGIMMSFLE-PSSPTFLLLKKLDSLCVSFAYINCCINPIIYVVAGQGFQKSLPELLREVLTE----ESVVR---------------------- | |||||||||||||||||||
| 10 | 5o9hA | 0.24 | 0.24 | 0.82 | 3.54 | Download | -------------------------------------NTLRVPDILALVIFAVVFLVGVLGNALVVWVTAFEAKRTINAIWFLNLAVADFLACLALPALFTSIVQHHHWPFGGAACSILPSLILLNMYASILLLATISADRFLLVFKPAWCQRFRGAGLAWILCAVAWGLALLLTIPSALYRVVREEYFPHDKRRERAVAIVRLVLGFLWPLLTLTICYTFILLRTWSARETRSTKTLKVVVAVVASFFIFWLPYQVTGIMMSPTFLLLKKLDSLCVSFAYINCCINPIIYVVAGQGFQK----SLPELLREVLTEESVVR---------------------- | |||||||||||||||||||
| ||||||||||||||||||||||||||
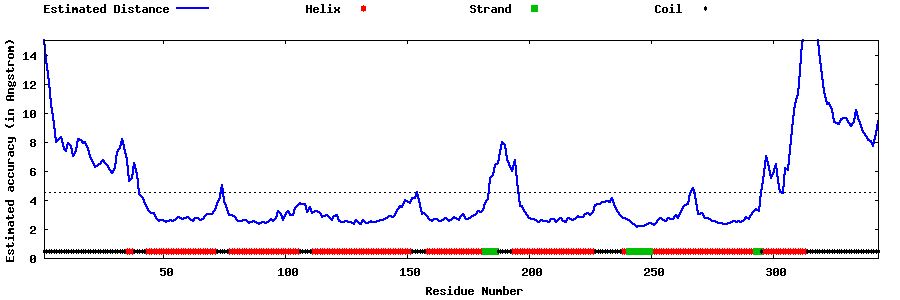
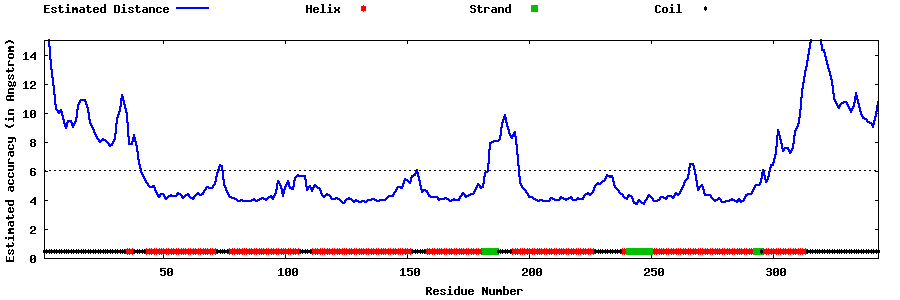
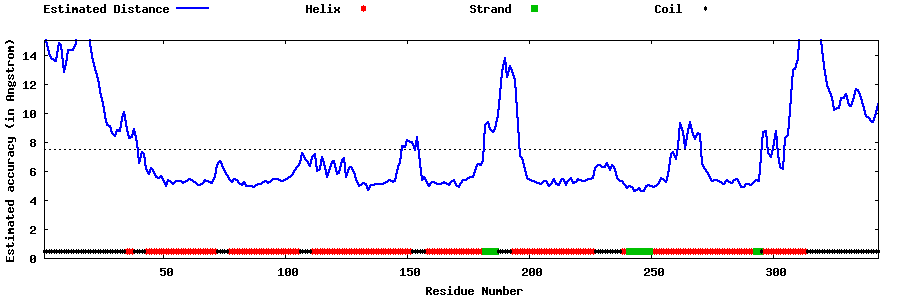
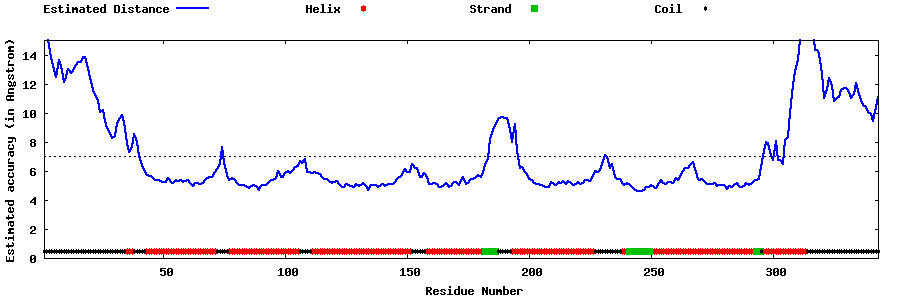
| Generated 3D models | Estimated local accuracy of models | ||
 | |||
|
|
|||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||