

You can:
| Name | Oxytocin receptor |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | OXTR |
| Synonym | OTR OT-R OT receptor |
| Disease | Threatened pre-term labour Postpartum haemorrhage Premature ejaculation Miscarriage Female sexual dysfunction [ Show all ] |
| Length | 389 |
| Amino acid sequence | MEGALAANWSAEAANASAAPPGAEGNRTAGPPRRNEALARVEVAVLCLILLLALSGNACVLLALRTTRQKHSRLFFFMKHLSIADLVVAVFQVLPQLLWDITFRFYGPDLLCRLVKYLQVVGMFASTYLLLLMSLDRCLAICQPLRSLRRRTDRLAVLATWLGCLVASAPQVHIFSLREVADGVFDCWAVFIQPWGPKAYITWITLAVYIVPVIVLAACYGLISFKIWQNLRLKTAAAAAAEAPEGAAAGDGGRVALARVSSVKLISKAKIRTVKMTFIIVLAFIVCWTPFFFVQMWSVWDANAPKEASAFIIVMLLASLNSCCNPWIYMLFTGHLFHELVQRFLCCSASYLKGRRLGETSASKKSNSSSFVLSHRSSSQRSCSQPSTA |
| UniProt | P30559 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | P30559 |
| 3D structure model | This predicted structure model is from GPCR-EXP P30559. |
| BioLiP | N/A |
| Therapeutic Target Database | T84486 |
| ChEMBL | CHEMBL2049 |
| IUPHAR | 369 |
| DrugBank | BE0000844 |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 468 | 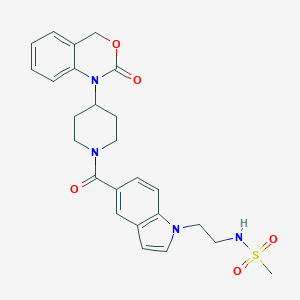 CHEMBL285013 CHEMBL285013 | C25H28N4O5S | 496.582 | 6 / 1 | 2.1 | Yes |
| 521500 | 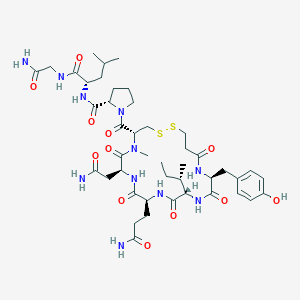 CHEMBL3813771 CHEMBL3813771 | C44H67N11O12S2 | 1006.21 | 14 / 10 | -1.6 | No |
| 2420 | 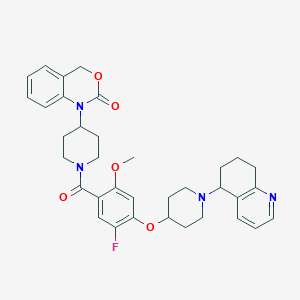 CHEMBL302029 CHEMBL302029 | C35H39FN4O5 | 614.718 | 8 / 0 | 4.8 | No |
| 2997 | 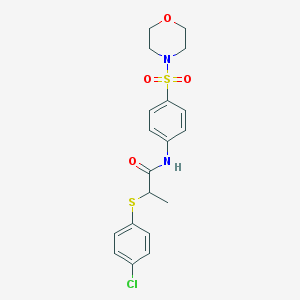 MLS000530432 MLS000530432 | C19H21ClN2O4S2 | 440.957 | 6 / 1 | 3.1 | Yes |
| 3130 | 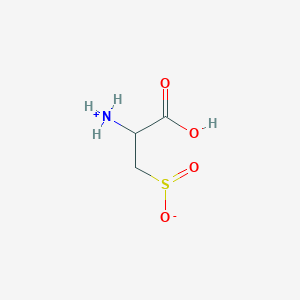 AC1N0YGA AC1N0YGA | C3H7NO4S | 153.152 | 5 / 2 | -4.7 | Yes |
| 3445 | 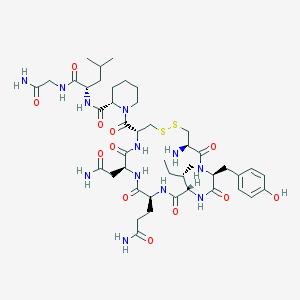 [Pip7]OT [Pip7]OT | C44H68N12O12S2 | 1021.22 | 15 / 12 | -2.2 | No |
| 5511 | 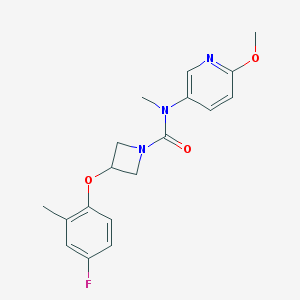 CHEMBL1092499 CHEMBL1092499 | C18H20FN3O3 | 345.374 | 5 / 0 | 2.5 | Yes |
| 5787 | 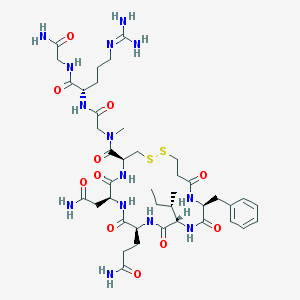 CHEMBL87636 CHEMBL87636 | C41H64N14O11S2 | 993.17 | 14 / 12 | -4.5 | No |
| 5791 | 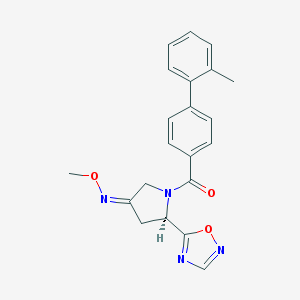 CHEMBL467153 CHEMBL467153 | C21H20N4O3 | 376.416 | 6 / 0 | 3.3 | Yes |
| 5999 | 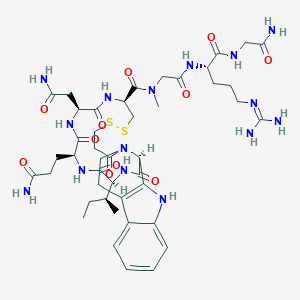 CHEMBL2371240 CHEMBL2371240 | C44H65N15O11S2 | 1044.22 | 14 / 12 | -4.4 | No |
| 6243 | 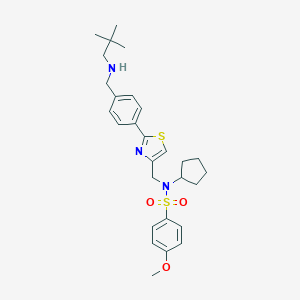 CHEMBL1643587 CHEMBL1643587 | C28H37N3O3S2 | 527.742 | 7 / 1 | 5.6 | No |
| 7031 | 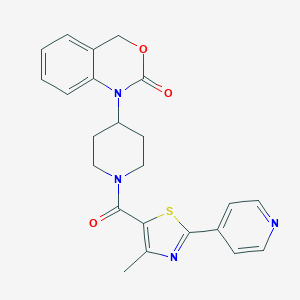 CHEMBL31986 CHEMBL31986 | C23H22N4O3S | 434.514 | 6 / 0 | 3.3 | Yes |
| 7090 | 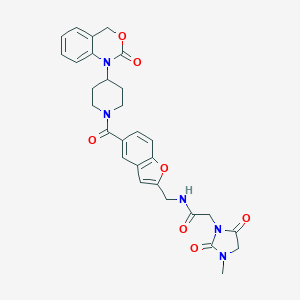 CHEMBL262384 CHEMBL262384 | C29H29N5O7 | 559.579 | 7 / 1 | 1.6 | No |
| 7673 | 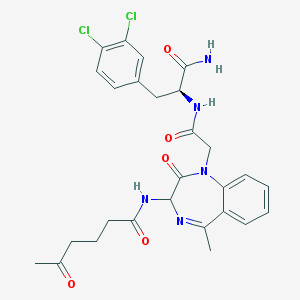 CHEMBL24353 CHEMBL24353 | C27H29Cl2N5O5 | 574.459 | 6 / 3 | 1.9 | No |
| 7733 | 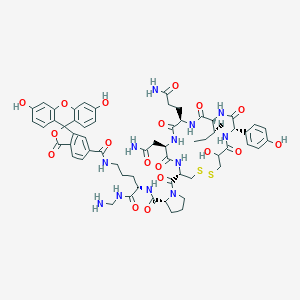 CHEMBL1790720 CHEMBL1790720 | C61H72N12O18S2 | 1325.43 | 21 / 15 | -0.9 | No |
| 9381 | 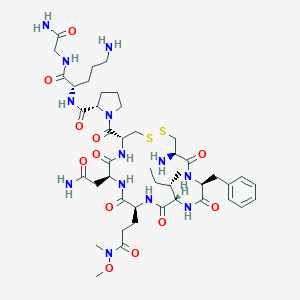 CHEMBL1817690 CHEMBL1817690 | C44H69N13O12S2 | 1036.23 | 16 / 11 | -3.5 | No |
| 10062 | 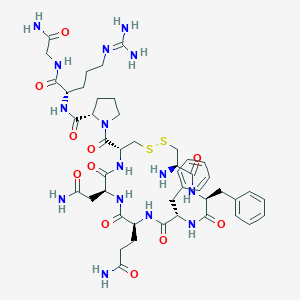 Phenypressin Phenypressin | C46H65N15O11S2 | 1068.24 | 15 / 13 | -4.5 | No |
| 11657 | 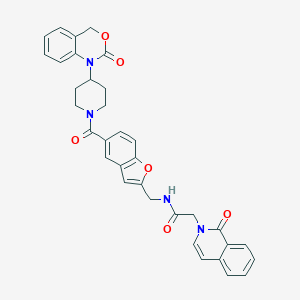 CHEMBL281044 CHEMBL281044 | C34H30N4O6 | 590.636 | 6 / 1 | 3.8 | No |
| 12152 | 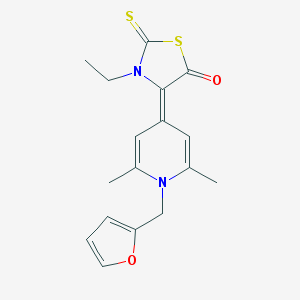 CHEMBL473106 CHEMBL473106 | C17H18N2O2S2 | 346.463 | 5 / 0 | 3.4 | Yes |
| 12881 | 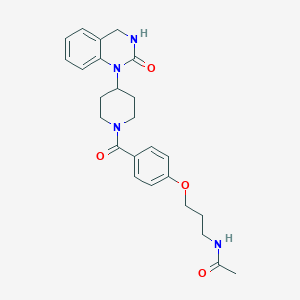 CHEMBL143341 CHEMBL143341 | C25H30N4O4 | 450.539 | 4 / 2 | 2.0 | Yes |
| 13805 | 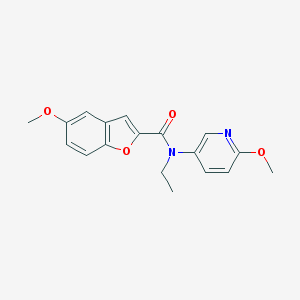 CHEMBL1093341 CHEMBL1093341 | C18H18N2O4 | 326.352 | 5 / 0 | 3.2 | Yes |
| 13912 | 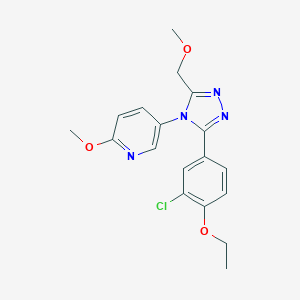 CHEMBL489386 CHEMBL489386 | C18H19ClN4O3 | 374.825 | 6 / 0 | 2.6 | Yes |
| 13940 | 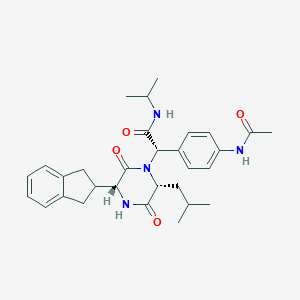 CHEMBL372619 CHEMBL372619 | C30H38N4O4 | 518.658 | 4 / 3 | 3.9 | No |
| 13941 | 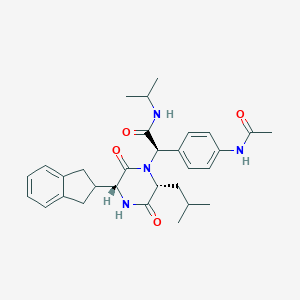 CHEMBL371008 CHEMBL371008 | C30H38N4O4 | 518.658 | 4 / 3 | 3.9 | No |
| 14502 | 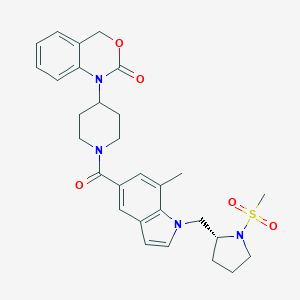 CHEMBL32745 CHEMBL32745 | C29H34N4O5S | 550.674 | 6 / 0 | 3.2 | No |
| 15032 | 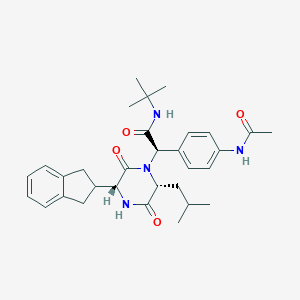 CHEMBL372554 CHEMBL372554 | C31H40N4O4 | 532.685 | 4 / 3 | 4.1 | No |
| 15573 | 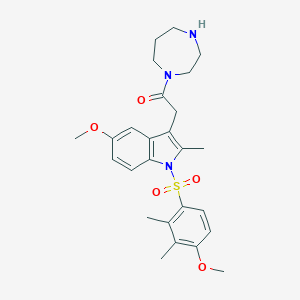 CHEMBL1779235 CHEMBL1779235 | C26H33N3O5S | 499.626 | 6 / 1 | 3.4 | Yes |
| 16163 | 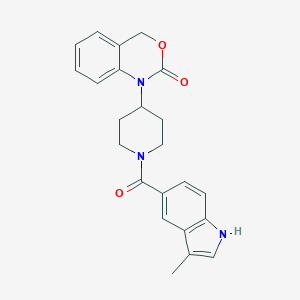 CHEMBL287067 CHEMBL287067 | C23H23N3O3 | 389.455 | 3 / 1 | 3.5 | Yes |
| 17415 | 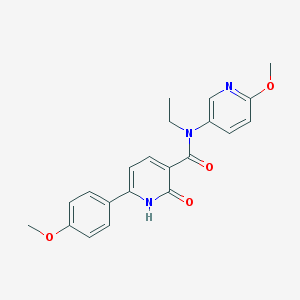 CHEMBL1093709 CHEMBL1093709 | C21H21N3O4 | 379.416 | 5 / 1 | 2.6 | Yes |
| 18528 | 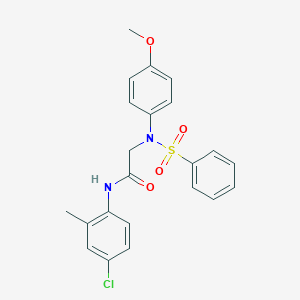 CHEMBL200682 CHEMBL200682 | C22H21ClN2O4S | 444.93 | 5 / 1 | 4.5 | Yes |
| 18759 | 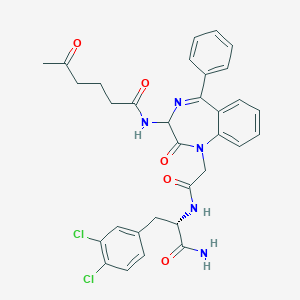 CHEMBL284120 CHEMBL284120 | C32H31Cl2N5O5 | 636.53 | 6 / 3 | 3.5 | No |
| 19074 | 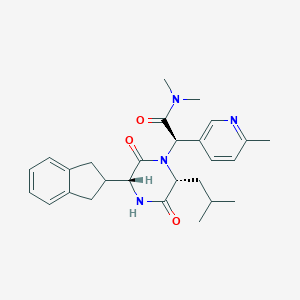 CHEMBL2037491 CHEMBL2037491 | C27H34N4O3 | 462.594 | 4 / 1 | 3.4 | Yes |
| 19334 | 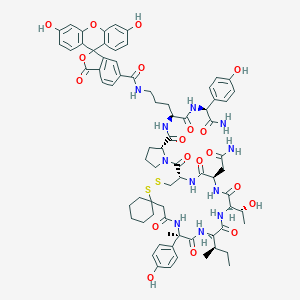 CHEMBL1790713 CHEMBL1790713 | C73H85N11O19S2 | 1484.66 | 21 / 15 | 3.7 | No |
| 20238 | 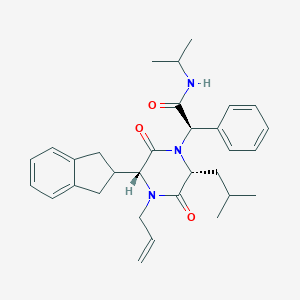 CHEMBL194432 CHEMBL194432 | C31H39N3O3 | 501.671 | 3 / 1 | 5.5 | No |
| 20585 | 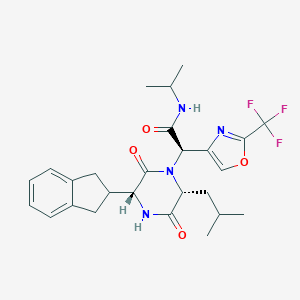 CHEMBL257916 CHEMBL257916 | C26H31F3N4O4 | 520.553 | 8 / 2 | 4.1 | No |
| 442491 | 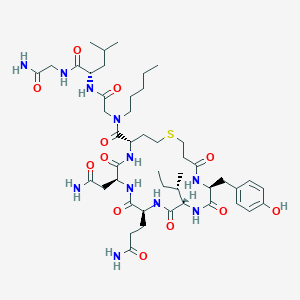 CHEMBL3354586 CHEMBL3354586 | C46H73N11O12S | 1004.22 | 13 / 11 | -0.2 | No |
| 20927 | 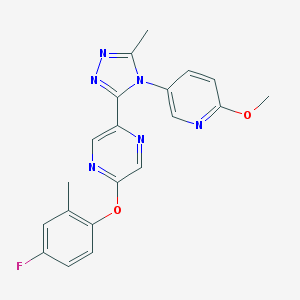 CHEMBL563771 CHEMBL563771 | C20H17FN6O2 | 392.394 | 8 / 0 | 2.6 | Yes |
| 21635 | 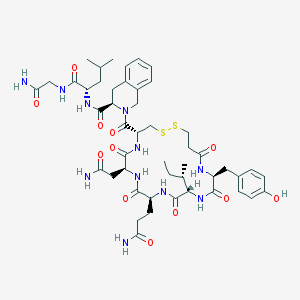 [Mpa1, D-Tic7]OT [Mpa1, D-Tic7]OT | C48H67N11O12S2 | 1054.25 | 14 / 11 | -0.7 | No |
| 21636 | 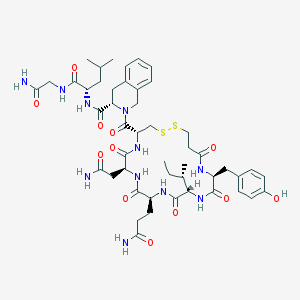 [Mpa1, L-Tic7]OT [Mpa1, L-Tic7]OT | C48H67N11O12S2 | 1054.25 | 14 / 11 | -0.7 | No |
| 442516 | 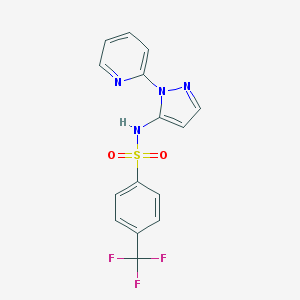 CHEMBL3342790 CHEMBL3342790 | C15H11F3N4O2S | 368.334 | 8 / 1 | 2.9 | Yes |
| 22000 | 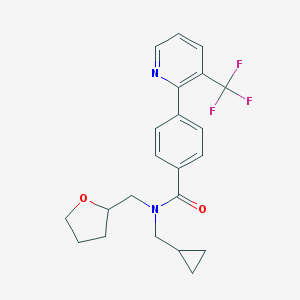 CHEMBL481730 CHEMBL481730 | C22H23F3N2O2 | 404.433 | 6 / 0 | 4.0 | Yes |
| 22500 | 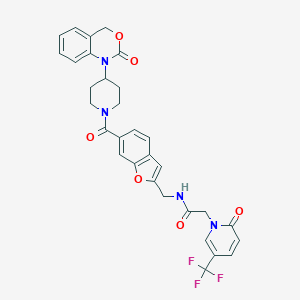 CHEMBL30550 CHEMBL30550 | C31H27F3N4O6 | 608.574 | 9 / 1 | 3.1 | No |
| 23435 | 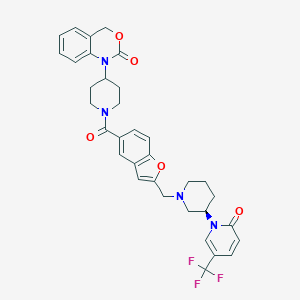 CHEMBL281988 CHEMBL281988 | C34H33F3N4O5 | 634.656 | 9 / 0 | 4.4 | No |
| 23730 | 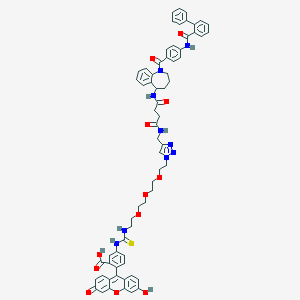 CHEMBL2172288 CHEMBL2172288 | C66H63N9O12S | 1206.34 | 15 / 7 | 4.8 | No |
| 24029 | 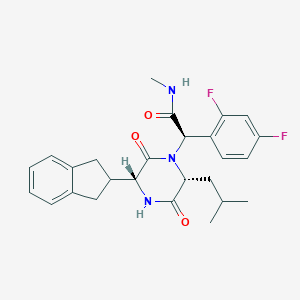 CHEMBL379192 CHEMBL379192 | C26H29F2N3O3 | 469.533 | 5 / 2 | 4.1 | Yes |
| 442622 | 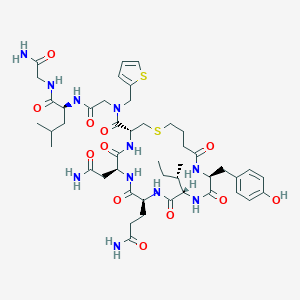 CHEMBL3354600 CHEMBL3354600 | C46H67N11O12S2 | 1030.23 | 14 / 11 | -0.8 | No |
| 24904 | 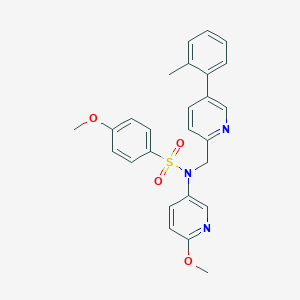 CHEMBL458635 CHEMBL458635 | C26H25N3O4S | 475.563 | 7 / 0 | 4.3 | Yes |
| 25319 | 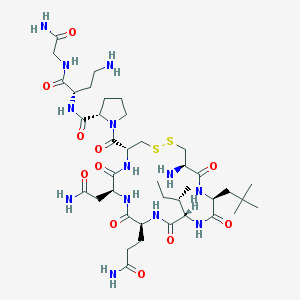 CHEMBL1819552 CHEMBL1819552 | C39H67N13O11S2 | 958.165 | 15 / 12 | -4.4 | No |
| 25349 | 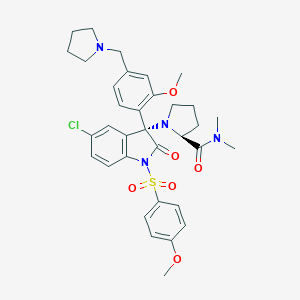 CHEMBL1779406 CHEMBL1779406 | C34H39ClN4O6S | 667.218 | 8 / 0 | 4.7 | No |
| 25988 | 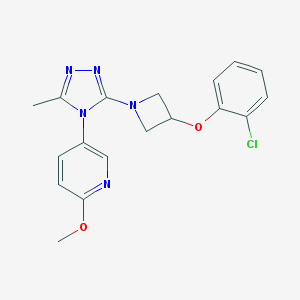 CHEMBL590099 CHEMBL590099 | C18H18ClN5O2 | 371.825 | 6 / 0 | 3.3 | Yes |
| 26885 | 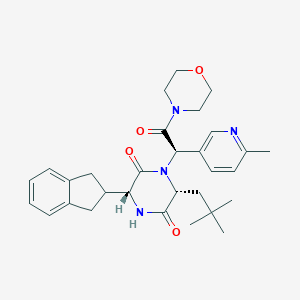 CHEMBL2037497 CHEMBL2037497 | C30H38N4O4 | 518.658 | 5 / 1 | 3.5 | No |
| 558107 | 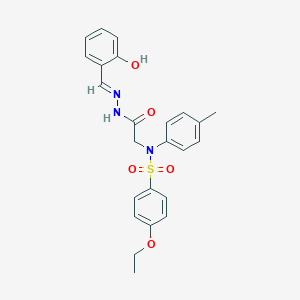 CHEMBL200329 CHEMBL200329 | C24H25N3O5S | 467.54 | 7 / 2 | 4.0 | Yes |
| 558110 | 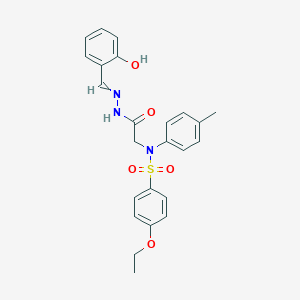 CID 136494531 CID 136494531 | C24H25N3O5S | 467.54 | 7 / 2 | 4.0 | Yes |
| 28852 | 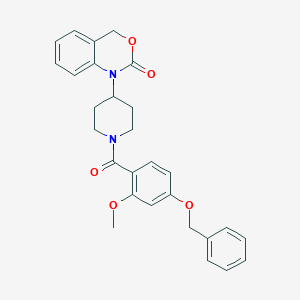 CHEMBL319646 CHEMBL319646 | C28H28N2O5 | 472.541 | 5 / 0 | 4.4 | Yes |
| 29644 | 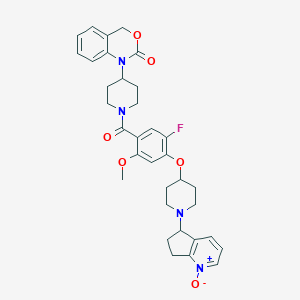 CHEMBL68816 CHEMBL68816 | C34H37FN4O6 | 616.69 | 8 / 0 | 3.5 | No |
| 29986 | 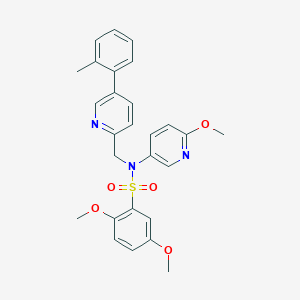 CHEMBL456254 CHEMBL456254 | C27H27N3O5S | 505.589 | 8 / 0 | 4.3 | No |
| 30002 | 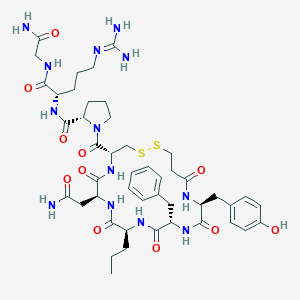 CHEMBL2369845 CHEMBL2369845 | C46H65N13O11S2 | 1040.23 | 14 / 12 | -2.0 | No |
| 30007 | 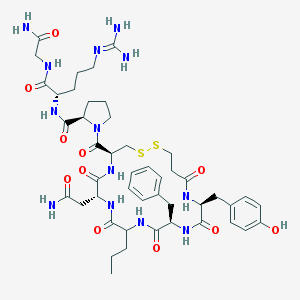 CHEMBL411013 CHEMBL411013 | C46H65N13O11S2 | 1040.23 | 14 / 12 | -2.0 | No |
| 30144 | 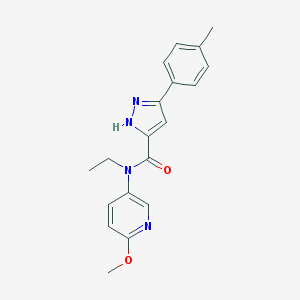 CHEMBL1093776 CHEMBL1093776 | C19H20N4O2 | 336.395 | 4 / 1 | 3.2 | Yes |
| 30821 | 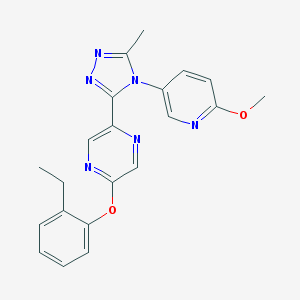 CHEMBL551395 CHEMBL551395 | C21H20N6O2 | 388.431 | 7 / 0 | 2.9 | Yes |
| 31353 | 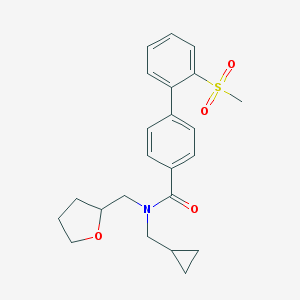 CHEMBL474219 CHEMBL474219 | C23H27NO4S | 413.532 | 4 / 0 | 3.4 | Yes |
| 31819 | 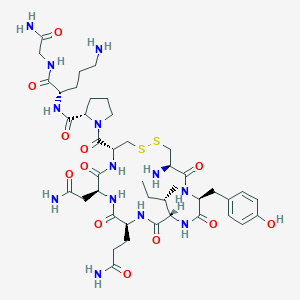 CHEMBL1819541 CHEMBL1819541 | C42H65N13O12S2 | 1008.18 | 16 / 13 | -4.5 | No |
| 32200 | 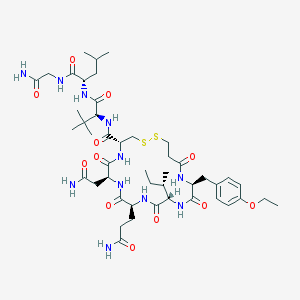 CHEMBL441128 CHEMBL441128 | C46H73N11O12S2 | 1036.28 | 14 / 11 | 0.0 | No |
| 32440 | 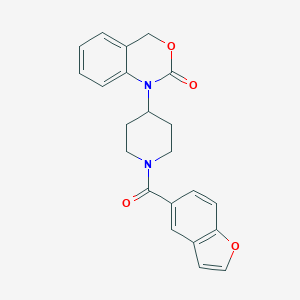 CHEMBL286084 CHEMBL286084 | C22H20N2O4 | 376.412 | 4 / 0 | 3.4 | Yes |
| 32723 | 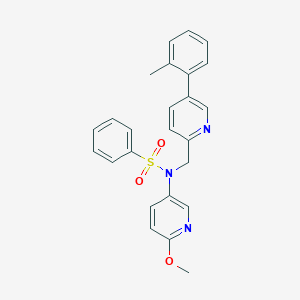 CHEMBL457998 CHEMBL457998 | C25H23N3O3S | 445.537 | 6 / 0 | 4.3 | Yes |
| 522544 | 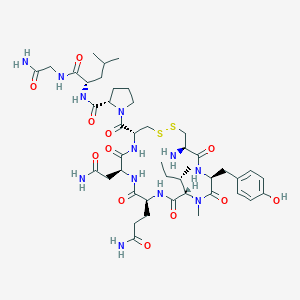 CHEMBL3814126 CHEMBL3814126 | C44H68N12O12S2 | 1021.22 | 15 / 11 | -2.4 | No |
| 34298 | 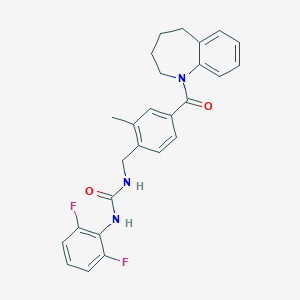 CHEMBL455699 CHEMBL455699 | C26H25F2N3O2 | 449.502 | 4 / 2 | 4.8 | Yes |
| 443033 | 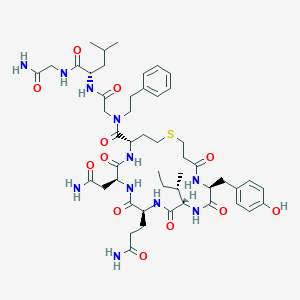 CHEMBL3354598 CHEMBL3354598 | C49H71N11O12S | 1038.23 | 13 / 11 | 0.0 | No |
| 35883 | 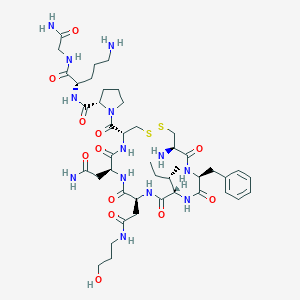 CHEMBL1817698 CHEMBL1817698 | C44H69N13O12S2 | 1036.23 | 16 / 13 | -4.4 | No |
| 36166 | 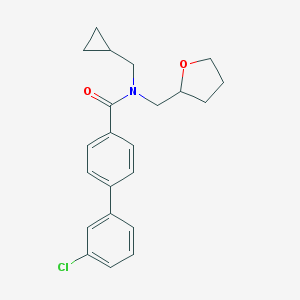 CHEMBL473442 CHEMBL473442 | C22H24ClNO2 | 369.889 | 2 / 0 | 4.8 | Yes |
| 36438 | 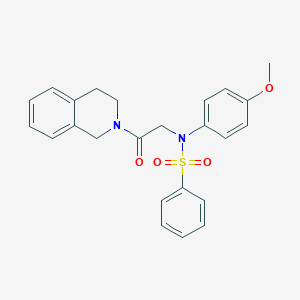 CHEMBL373167 CHEMBL373167 | C24H24N2O4S | 436.526 | 5 / 0 | 3.7 | Yes |
| 37057 | 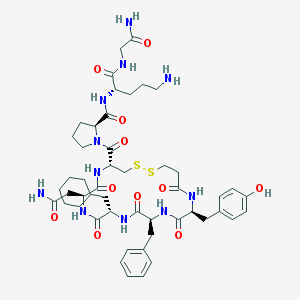 d[Cha4,Orn8]VP d[Cha4,Orn8]VP | C49H69N11O11S2 | 1052.28 | 14 / 11 | 0.5 | No |
| 37829 | 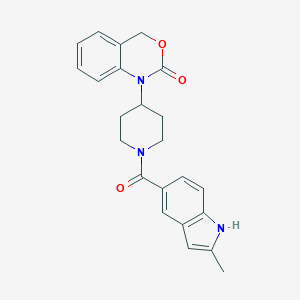 CHEMBL32740 CHEMBL32740 | C23H23N3O3 | 389.455 | 3 / 1 | 3.5 | Yes |
| 38023 | 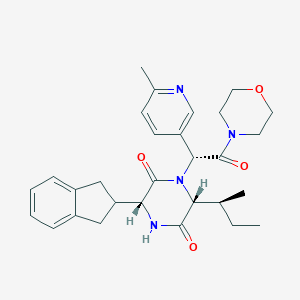 CHEMBL2037501 CHEMBL2037501 | C29H36N4O4 | 504.631 | 5 / 1 | 3.1 | No |
| 38024 | 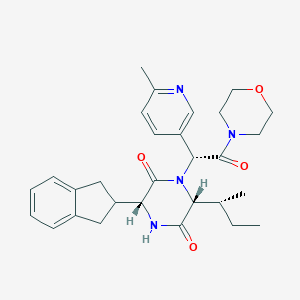 CHEMBL2037500 CHEMBL2037500 | C29H36N4O4 | 504.631 | 5 / 1 | 3.1 | No |
| 38473 | 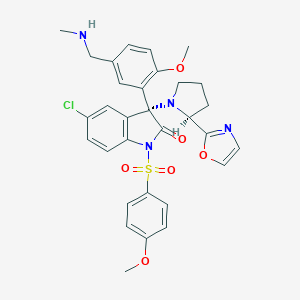 CHEMBL1779419 CHEMBL1779419 | C31H31ClN4O6S | 623.121 | 9 / 1 | 4.1 | No |
| 39146 | 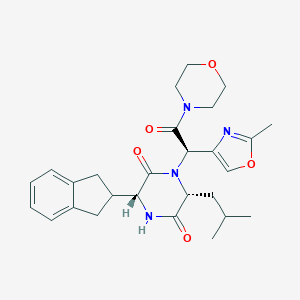 CHEMBL271039 CHEMBL271039 | C27H34N4O5 | 494.592 | 6 / 1 | 2.6 | Yes |
| 39147 | 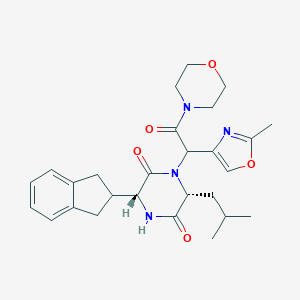 CHEMBL513488 CHEMBL513488 | C27H34N4O5 | 494.592 | 6 / 1 | 2.6 | Yes |
| 39220 | 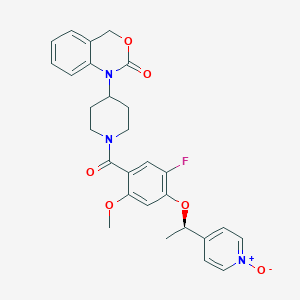 CHEMBL105514 CHEMBL105514 | C28H28FN3O6 | 521.545 | 7 / 0 | 2.8 | No |
| 443245 | 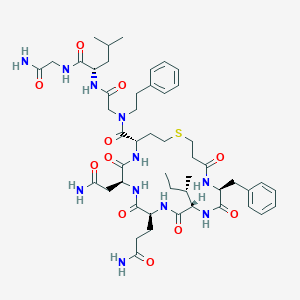 CHEMBL3354599 CHEMBL3354599 | C49H71N11O11S | 1022.23 | 12 / 10 | 0.3 | No |
| 39751 | 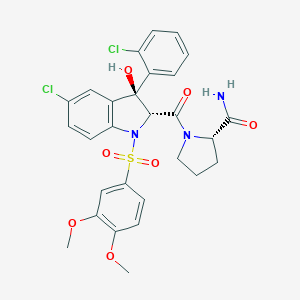 Relcovaptan Relcovaptan | C28H27Cl2N3O7S | 620.498 | 8 / 2 | 3.8 | No |
| 558495 | 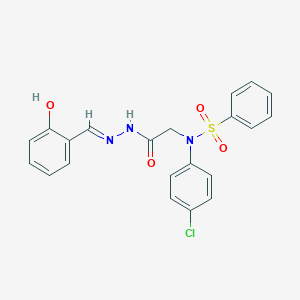 CHEMBL200792 CHEMBL200792 | C21H18ClN3O4S | 443.902 | 6 / 2 | 3.9 | Yes |
| 39865 | 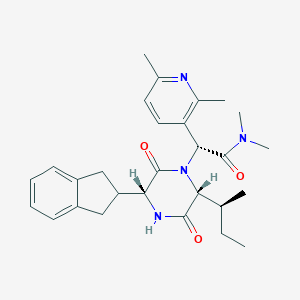 CHEMBL2037510 CHEMBL2037510 | C28H36N4O3 | 476.621 | 4 / 1 | 3.8 | Yes |
| 522736 | 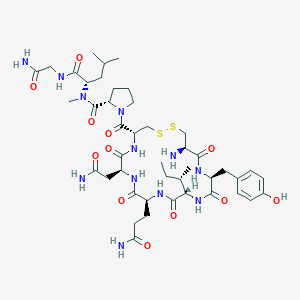 CHEMBL3814744 CHEMBL3814744 | C44H68N12O12S2 | 1021.22 | 15 / 11 | -2.4 | No |
| 39984 | 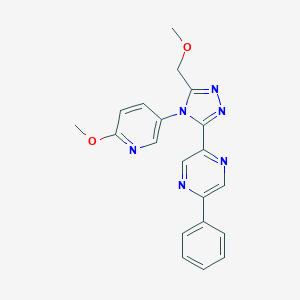 CHEMBL512257 CHEMBL512257 | C20H18N6O2 | 374.404 | 7 / 0 | 1.2 | Yes |
| 40573 | 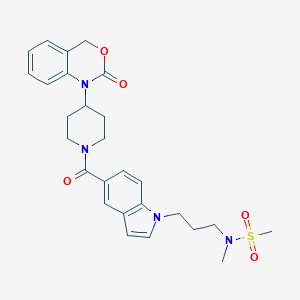 CHEMBL285012 CHEMBL285012 | C27H32N4O5S | 524.636 | 6 / 0 | 2.6 | No |
| 41113 | 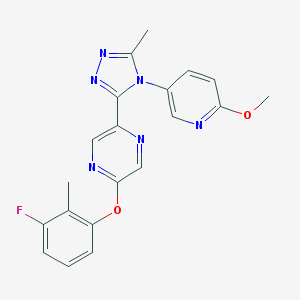 CHEMBL549494 CHEMBL549494 | C20H17FN6O2 | 392.394 | 8 / 0 | 2.6 | Yes |
| 41471 | 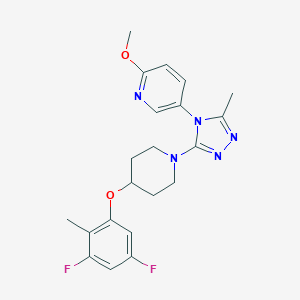 CHEMBL606976 CHEMBL606976 | C21H23F2N5O2 | 415.445 | 8 / 0 | 4.0 | Yes |
| 41965 | 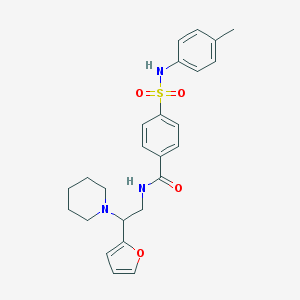 MLS001079319 MLS001079319 | C25H29N3O4S | 467.584 | 6 / 2 | 3.6 | Yes |
| 42318 | 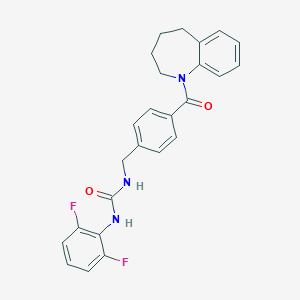 CHEMBL455718 CHEMBL455718 | C25H23F2N3O2 | 435.475 | 4 / 2 | 4.4 | Yes |
| 44244 | 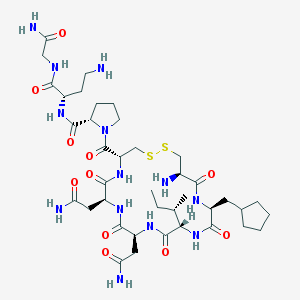 CHEMBL1817769 CHEMBL1817769 | C39H65N13O11S2 | 956.149 | 15 / 12 | -4.2 | No |
| 44756 | 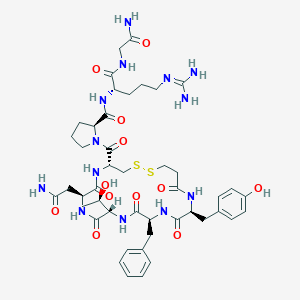 CHEMBL2369831 CHEMBL2369831 | C45H63N13O12S2 | 1042.2 | 15 / 13 | -2.9 | No |
| 44762 | 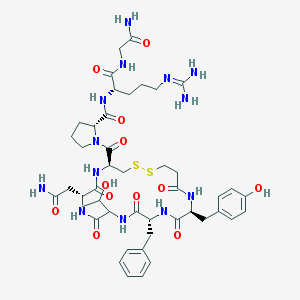 CHEMBL405783 CHEMBL405783 | C45H63N13O12S2 | 1042.2 | 15 / 13 | -2.9 | No |
| 45753 | 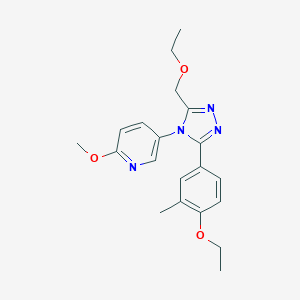 CHEMBL492029 CHEMBL492029 | C20H24N4O3 | 368.437 | 6 / 0 | 2.7 | Yes |
| 443523 | 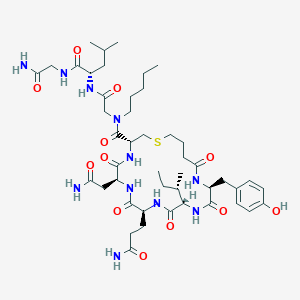 CHEMBL3354585 CHEMBL3354585 | C46H73N11O12S | 1004.22 | 13 / 11 | -0.2 | No |
| 47395 | 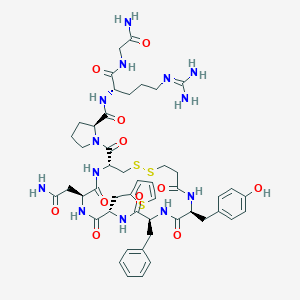 CHEMBL2369848 CHEMBL2369848 | C48H63N13O11S3 | 1094.29 | 15 / 12 | -1.5 | No |
| 47396 | 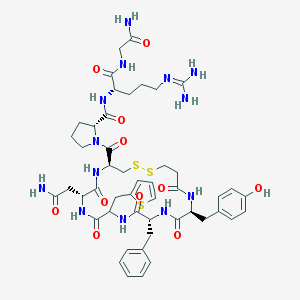 CHEMBL411485 CHEMBL411485 | C48H63N13O11S3 | 1094.29 | 15 / 12 | -1.5 | No |
| 48326 | 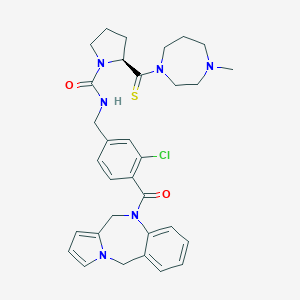 CHEMBL601929 CHEMBL601929 | C32H37ClN6O2S | 605.198 | 4 / 1 | 3.5 | No |
| 48813 | 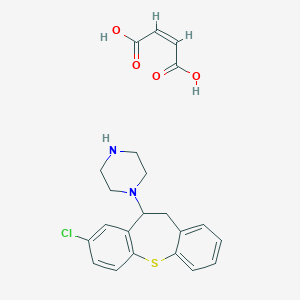 MLS000547068 MLS000547068 | C22H23ClN2O4S | 446.946 | 7 / 3 | N/A | N/A |
| 49000 | 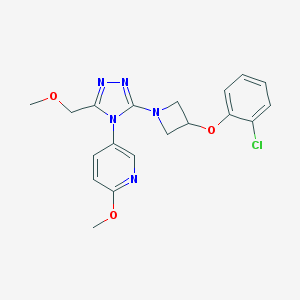 CHEMBL590349 CHEMBL590349 | C19H20ClN5O3 | 401.851 | 7 / 0 | 2.6 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218