

You can:
| Name | Somatostatin receptor type 3 |
|---|---|
| Species | Rattus norvegicus (Rat) |
| Gene | Sstr3 |
| Synonym | SSR-28 SS3R SS3-R SS-3-R SRIF1C [ Show all ] |
| Disease | N/A for non-human GPCRs |
| Length | 428 |
| Amino acid sequence | MAAVTYPSSVPTTLDPGNASSAWPLDTSLGNASAGTSLAGLAVSGILISLVYLVVCVVGLLGNSLVIYVVLRHTSSPSVTSVYILNLALADELFMLGLPFLAAQNALSYWPFGSLMCRLVMAVDGINQFTSIFCLTVMSVDRYLAVVHPTRSARWRTAPVARMVSAAVWVASAVVVLPVVVFSGVPRGMSTCHMQWPEPAAAWRTAFIIYTAALGFFGPLLVICLCYLLIVVKVRSTTRRVRAPSCQWVQAPACQRRRRSERRVTRMVVAVVALFVLCWMPFYLLNIVNVVCPLPEEPAFFGLYFLVVALPYANSCANPILYGFLSYRFKQGFRRILLRPSRRVRSQEPGSGPPEKTEEEEDEEEEERREEEERRMQRGQEMNGRLSQIAQPGPSGQQQRPCTGTAKEQQLLPQEATAGDKASTLSHL |
| UniProt | P30936 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL3340 |
| IUPHAR | 357 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 16295 | 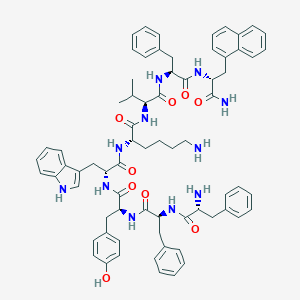 BDBM84629 BDBM84629 | C71H81N11O9 | 1232.5 | 11 / 12 | 8.1 | No |
| 39298 | 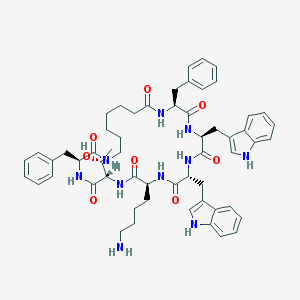 L-362855 L-362855 | C57H70N10O8 | 1023.25 | 9 / 11 | 5.7 | No |
| 40414 | 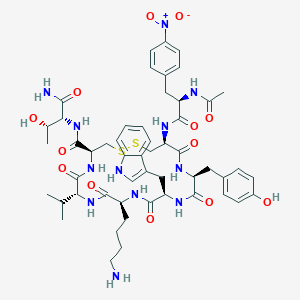 BDBM85012 BDBM85012 | C52H68N12O13S2 | 1133.31 | 16 / 13 | 1.3 | No |
| 40416 | 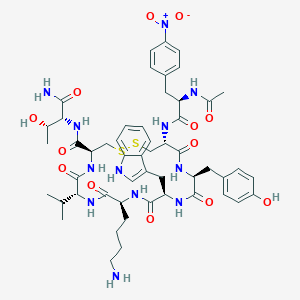 BDBM85011 BDBM85011 | C52H68N12O13S2 | 1133.31 | 16 / 13 | 1.3 | No |
| 58023 | 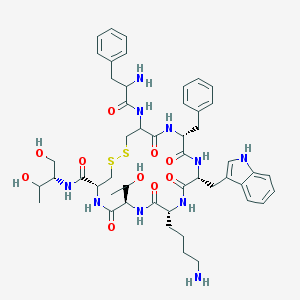 Octreotide Octreotide | C49H66N10O10S2 | 1019.25 | 14 / 13 | 1.0 | No |
| 553530 | 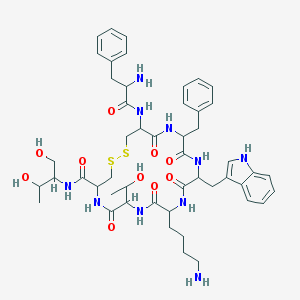 Octreotide Octreotide | C49H66N10O10S2 | 1019.25 | 14 / 13 | 1.0 | No |
| 96174 | 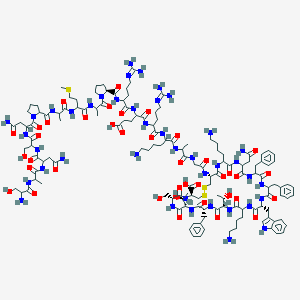 BDBM82460 BDBM82460 | C137H207N41O39S3 | 3148.59 | 48 / 44 | -16.0 | No |
| 553732 | 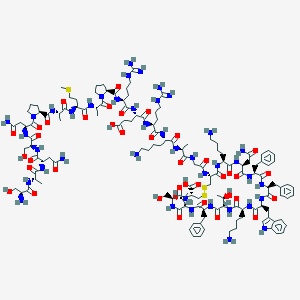 Somatostatin 28 Somatostatin 28 | C137H207N41O39S3 | 3148.59 | 48 / 46 | -15.2 | No |
| 118927 | 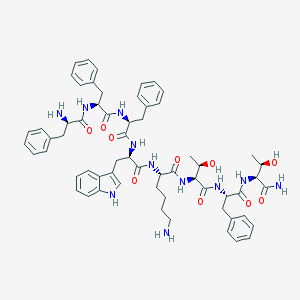 BIM 23052 BIM 23052 | C61H75N11O10 | 1122.34 | 12 / 13 | 2.7 | No |
| 131133 | 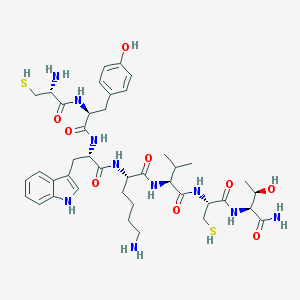 BDBM82254 BDBM82254 | C41H60N10O9S2 | 901.112 | 13 / 14 | 0.0 | No |
| 148748 | 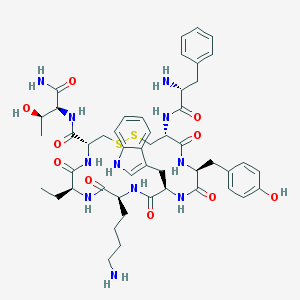 BIM-23023 BIM-23023 | C49H65N11O10S2 | 1032.25 | 14 / 13 | 0.8 | No |
| 155253 | 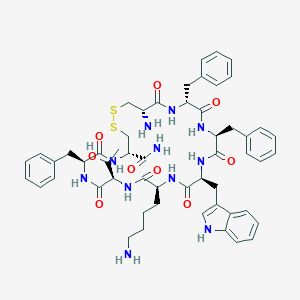 BIM 23268 BIM 23268 | C54H67N11O9S2 | 1078.32 | 13 / 12 | 2.8 | No |
| 183510 | 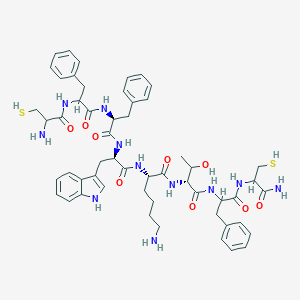 BDBM85230 BDBM85230 | C54H69N11O9S2 | 1080.33 | 13 / 14 | 2.3 | No |
| 213992 | 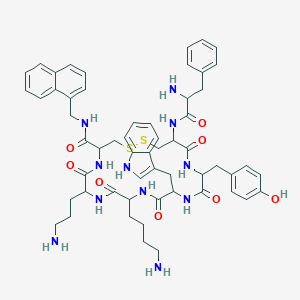 BDBM82255 BDBM82255 | C57H69N11O8S2 | 1100.37 | 13 / 12 | 3.7 | No |
| 219839 | 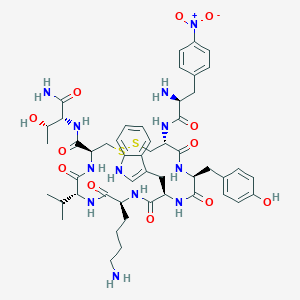 BDBM85009 BDBM85009 | C50H66N12O12S2 | 1091.27 | 16 / 13 | 1.1 | No |
| 219840 | 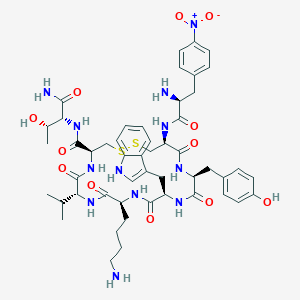 BDBM85013 BDBM85013 | C50H66N12O12S2 | 1091.27 | 16 / 13 | 1.1 | No |
| 223712 | 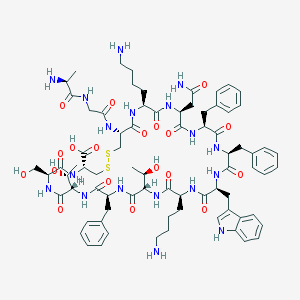 SOMATOSTATIN SOMATOSTATIN | C76H104N18O19S2 | 1637.9 | 24 / 22 | -3.1 | No |
| 223722 | 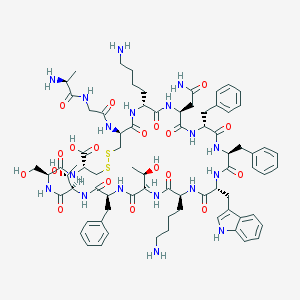 Somatostatin-14 Somatostatin-14 | C76H104N18O19S2 | 1637.9 | 24 / 22 | -3.1 | No |
| 554415 | 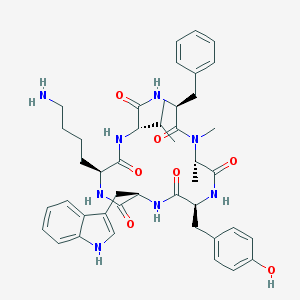 seglitide seglitide | C44H56N8O7 | 808.981 | 8 / 8 | 3.9 | No |
| 229008 | 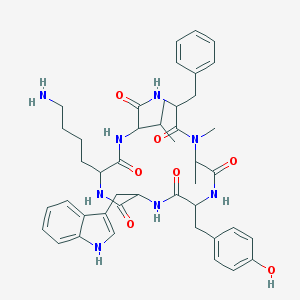 BDBM81766 BDBM81766 | C44H56N8O7 | 808.981 | 8 / 8 | 3.9 | No |
| 268945 | 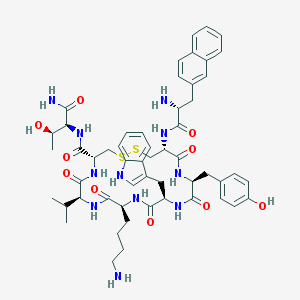 Lanreotide Lanreotide | C54H69N11O10S2 | 1096.33 | 14 / 13 | 2.5 | No |
| 554608 | 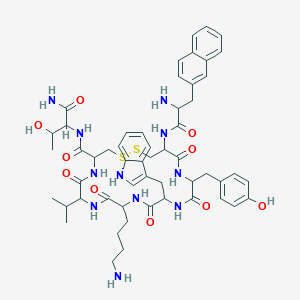 Lanreotide acetate Lanreotide acetate | C54H69N11O10S2 | 1096.33 | 14 / 13 | 2.5 | No |
| 275736 | 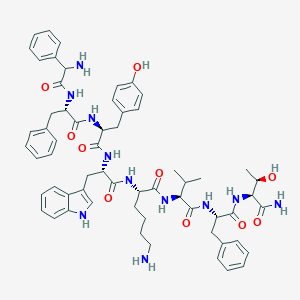 BDBM82257 BDBM82257 | C61H75N11O10 | 1122.34 | 12 / 13 | 4.4 | No |
| 499339 | 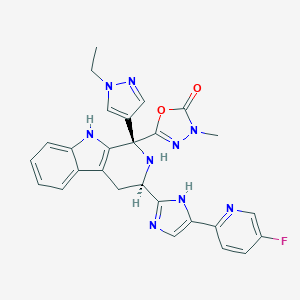 CHEMBL3582336 CHEMBL3582336 | C27H24FN9O2 | 525.548 | 8 / 3 | 2.0 | No |
| 325276 | 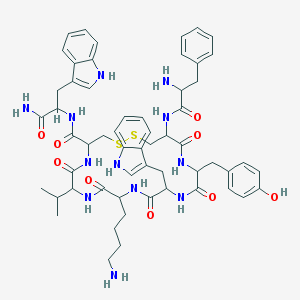 Vapreotide Vapreotide | C57H70N12O9S2 | 1131.38 | 13 / 13 | 3.6 | No |
| 326207 | 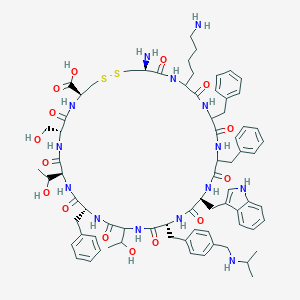 CH-275 CH-275 | C74H96N14O15S2 | 1485.79 | 20 / 18 | 1.6 | No |
| 361202 | 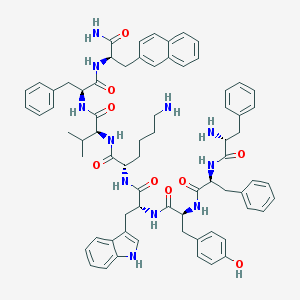 BIM 23056 BIM 23056 | C71H81N11O9 | 1232.5 | 11 / 12 | 8.1 | No |
| 555105 | 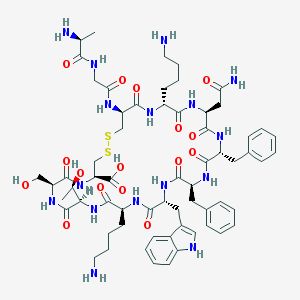 SS14 SS14 | C63H88N16O16S2 | 1389.61 | 21 / 19 | -4.1 | No |
| 433000 |  BDBM85229 BDBM85229 | C54H69N11O10S2 | 1096.33 | 14 / 15 | 2.0 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218