

You can:
| Name | Somatostatin receptor type 4 |
|---|---|
| Species | Rattus norvegicus (Rat) |
| Gene | Sstr4 |
| Synonym | SRIF2B SS-4-R SS4-R SS4R SST4 receptor |
| Disease | N/A for non-human GPCRs |
| Length | 384 |
| Amino acid sequence | MNTPATLPLGGEDTTWTPGINASWAPDEEEDAVRSDGTGTAGMVTIQCIYALVCLVGLVGNALVIFVILRYAKMKTATNIYLLNLAVADELFMLSVPFVASAAALRHWPFGAVLCRAVLSVDGLNMFTSVFCLTVLSVDRYVAVVHPLRAATYRRPSVAKLINLGVWLASLLVTLPIAVFADTRPARGGEAVACNLHWPHPAWSAVFVIYTFLLGFLLPVLAIGLCYLLIVGKMRAVALRAGWQQRRRSEKKITRLVLMVVTVFVLCWMPFYVVQLLNLFVTSLDATVNHVSLILSYANSCANPILYGFLSDNFRRSFQRVLCLRCCLLETTGGAEEEPLDYYATALKSRGGPGCICPPLPCQQEPMQAEPACKRVPFTKTTTF |
| UniProt | P30937 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | N/A |
| IUPHAR | 358 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 16300 | 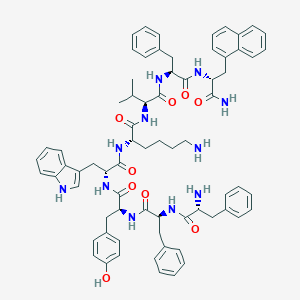 BDBM84629 BDBM84629 | C71H81N11O9 | 1232.5 | 11 / 12 | 8.1 | No |
| 19010 | 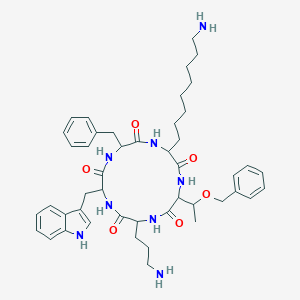 SA V SA V | C46H62N8O6 | 823.052 | 8 / 8 | 4.6 | No |
| 28612 | 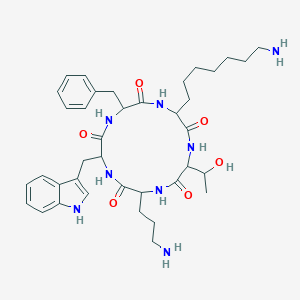 SA II SA II | C38H54N8O6 | 718.9 | 8 / 9 | 2.6 | No |
| 39297 | 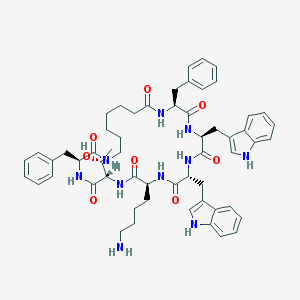 L-362855 L-362855 | C57H70N10O8 | 1023.25 | 9 / 11 | 5.7 | No |
| 46043 | 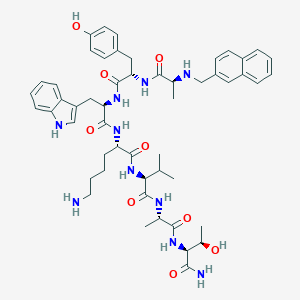 BDBM82458 BDBM82458 | C52H68N10O9 | 977.177 | 11 / 12 | 3.4 | No |
| 50027 | 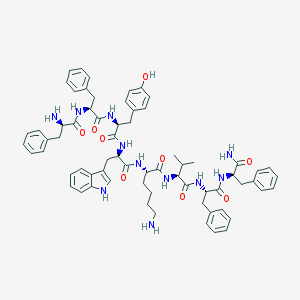 CHEMBL268112 CHEMBL268112 | C67H79N11O9 | 1182.44 | 11 / 12 | 5.6 | No |
| 52240 | 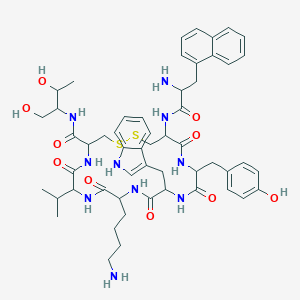 BDBM84620 BDBM84620 | C54H70N10O10S2 | 1083.33 | 14 / 13 | 2.9 | No |
| 58024 | 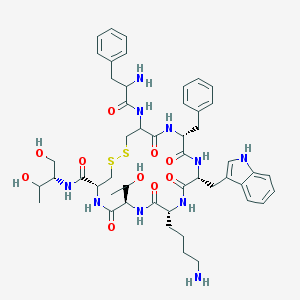 Octreotide Octreotide | C49H66N10O10S2 | 1019.25 | 14 / 13 | 1.0 | No |
| 59019 | 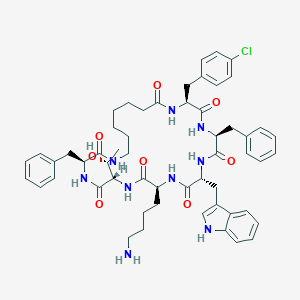 BDBM82466 BDBM82466 | C55H68ClN9O8 | 1018.65 | 9 / 10 | 6.2 | No |
| 61297 | 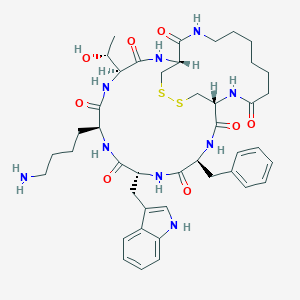 CHEMBL2369699 CHEMBL2369699 | C43H59N9O8S2 | 894.12 | 11 / 10 | 2.1 | No |
| 65484 | 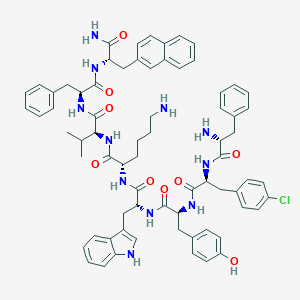 BDBM84640 BDBM84640 | C71H80ClN11O9 | 1266.94 | 11 / 12 | 8.8 | No |
| 78637 | 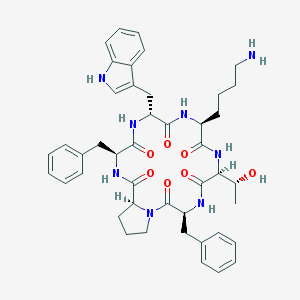 CHEMBL421362 CHEMBL421362 | C44H54N8O7 | 806.965 | 8 / 8 | 3.4 | No |
| 96178 | 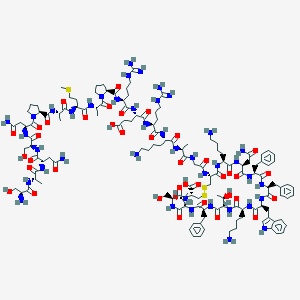 Somatostatin 28 Somatostatin 28 | C137H207N41O39S3 | 3148.59 | 48 / 46 | -15.2 | No |
| 106548 | 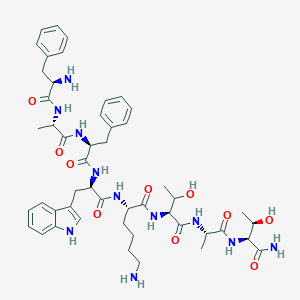 BDBM84625 BDBM84625 | C49H67N11O10 | 970.142 | 12 / 13 | -0.5 | No |
| 106955 | 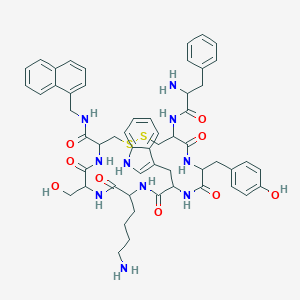 BDBM84623 BDBM84623 | C55H64N10O9S2 | 1073.3 | 13 / 12 | 3.8 | No |
| 118923 | 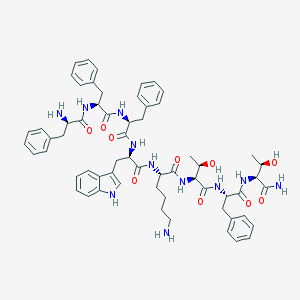 BIM 23052 BIM 23052 | C61H75N11O10 | 1122.34 | 12 / 13 | 2.7 | No |
| 128023 | 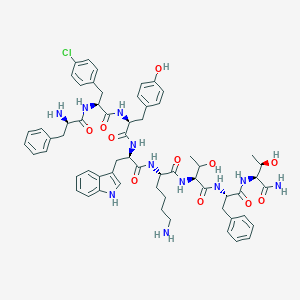 BDBM84624 BDBM84624 | C61H74ClN11O11 | 1172.78 | 13 / 14 | 3.7 | No |
| 131135 | 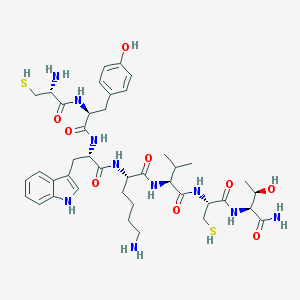 BDBM82254 BDBM82254 | C41H60N10O9S2 | 901.112 | 13 / 14 | 0.0 | No |
| 148744 | 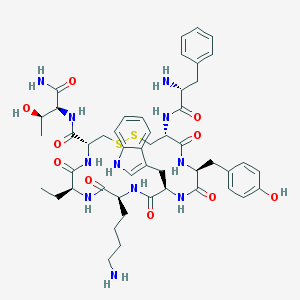 BIM-23023 BIM-23023 | C49H65N11O10S2 | 1032.25 | 14 / 13 | 0.8 | No |
| 160211 | 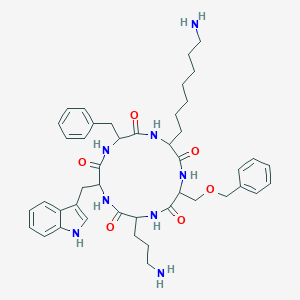 BDBM84644 BDBM84644 | C44H58N8O6 | 794.998 | 8 / 8 | 3.6 | No |
| 174021 | 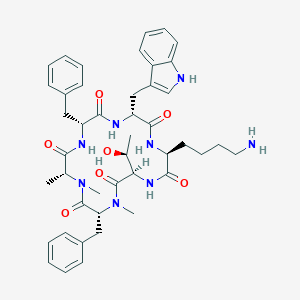 BDBM82454 BDBM82454 | C44H56N8O7 | 808.981 | 8 / 7 | 3.4 | No |
| 174352 | 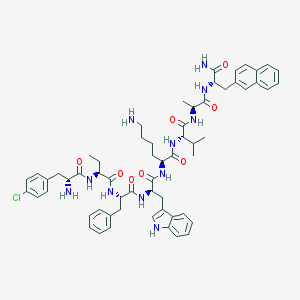 BDBM84617 BDBM84617 | C60H74ClN11O8 | 1112.77 | 10 / 11 | 6.5 | No |
| 183482 | 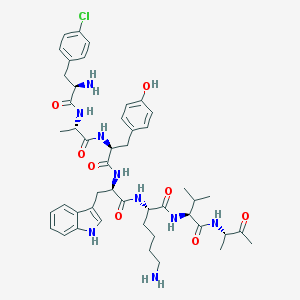 BDBM84616 BDBM84616 | C47H62ClN9O8 | 916.518 | 10 / 10 | 2.5 | No |
| 200928 | 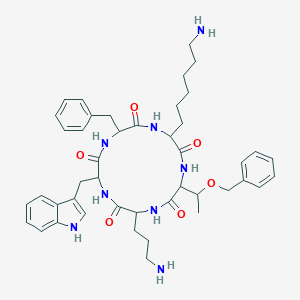 SA IV SA IV | C44H58N8O6 | 794.998 | 8 / 8 | 3.5 | No |
| 213987 | 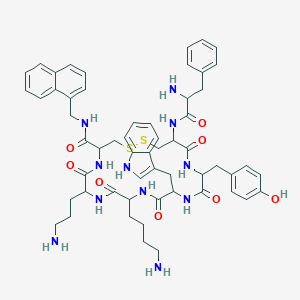 BDBM82255 BDBM82255 | C57H69N11O8S2 | 1100.37 | 13 / 12 | 3.7 | No |
| 223710 | 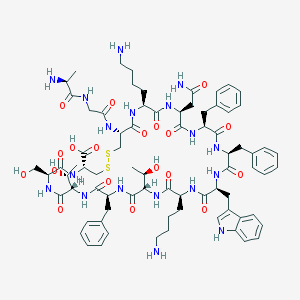 SOMATOSTATIN SOMATOSTATIN | C76H104N18O19S2 | 1637.9 | 24 / 22 | -3.1 | No |
| 223721 | 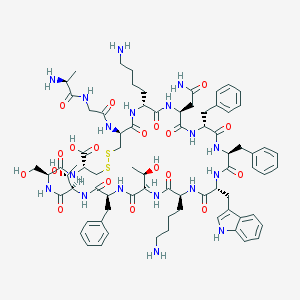 Somatostatin-14 Somatostatin-14 | C76H104N18O19S2 | 1637.9 | 24 / 22 | -3.1 | No |
| 223729 | 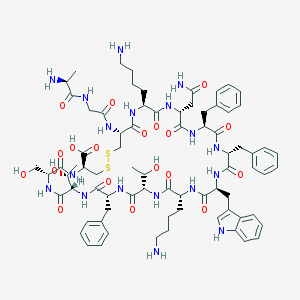 SOMATOSTATIN SOMATOSTATIN | C76H104N18O19S2 | 1637.9 | 24 / 22 | -3.1 | No |
| 228999 | 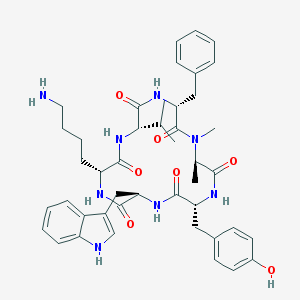 CHEMBL413535 CHEMBL413535 | C44H56N8O7 | 808.981 | 8 / 8 | 3.9 | No |
| 229007 | 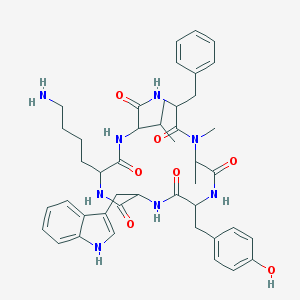 BDBM81766 BDBM81766 | C44H56N8O7 | 808.981 | 8 / 8 | 3.9 | No |
| 261035 | 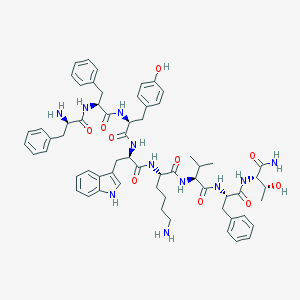 BIM-23058 BIM-23058 | C62H77N11O10 | 1136.36 | 12 / 13 | 4.7 | No |
| 268944 | 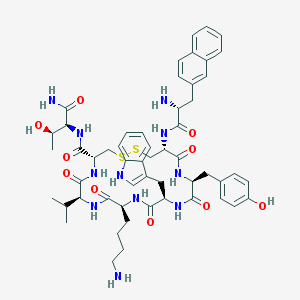 Lanreotide Lanreotide | C54H69N11O10S2 | 1096.33 | 14 / 13 | 2.5 | No |
| 275733 | 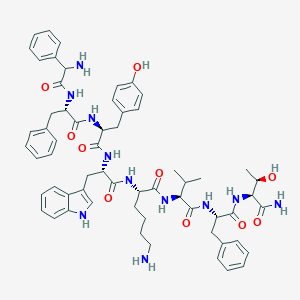 BDBM82257 BDBM82257 | C61H75N11O10 | 1122.34 | 12 / 13 | 4.4 | No |
| 276240 | 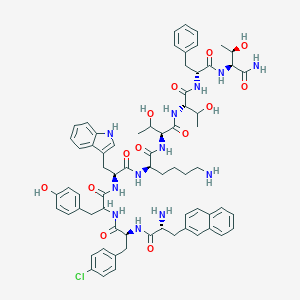 BDBM84619 BDBM84619 | C69H83ClN12O13 | 1323.94 | 15 / 16 | 4.1 | No |
| 293364 | 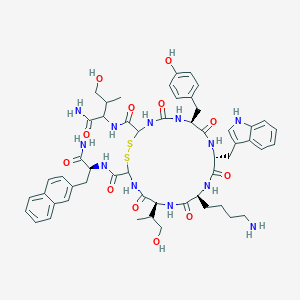 BDBM82462 BDBM82462 | C54H68N12O12S2 | 1141.33 | 15 / 15 | 1.5 | No |
| 316656 | 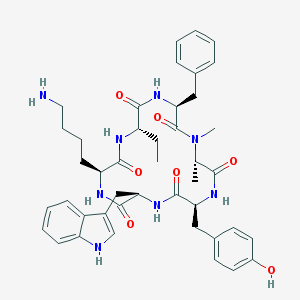 BIM-23027 BIM-23027 | C43H54N8O7 | 794.954 | 8 / 8 | 3.5 | No |
| 326209 | 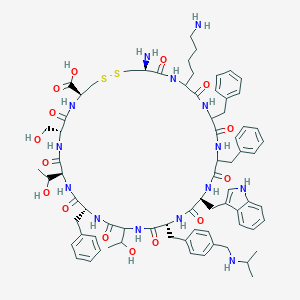 CH-275 CH-275 | C74H96N14O15S2 | 1485.79 | 20 / 18 | 1.6 | No |
| 332967 | 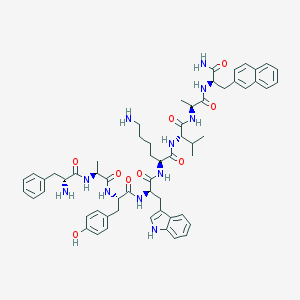 BDBM82461 BDBM82461 | C59H73N11O9 | 1080.3 | 11 / 12 | 5.0 | No |
| 345535 | 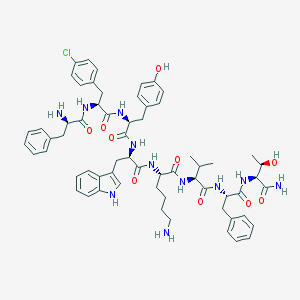 SCHEMBL1579410 SCHEMBL1579410 | C62H76ClN11O10 | 1170.81 | 12 / 13 | 5.3 | No |
| 345922 | 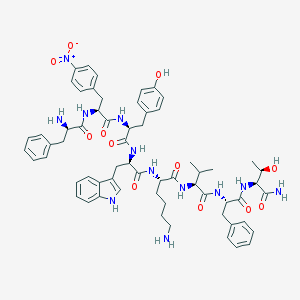 CHEMBL3350185 CHEMBL3350185 | C62H76N12O12 | 1181.36 | 14 / 13 | 4.5 | No |
| 348765 | 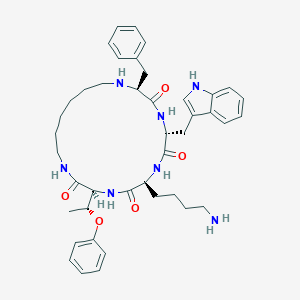 cycloantagonist SA cycloantagonist SA | C42H55N7O5 | 737.946 | 7 / 7 | 4.8 | No |
| 357457 | 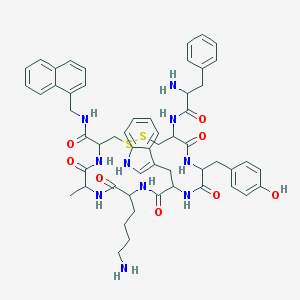 BDBM84626 BDBM84626 | C55H64N10O8S2 | 1057.3 | 12 / 11 | 4.3 | No |
| 361203 | 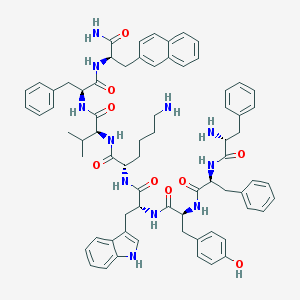 BIM 23056 BIM 23056 | C71H81N11O9 | 1232.5 | 11 / 12 | 8.1 | No |
| 361648 | 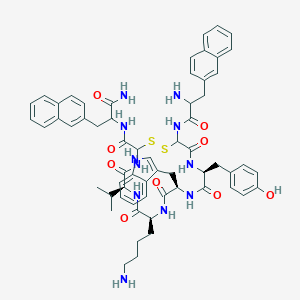 BDBM84621 BDBM84621 | C61H69N11O9S2 | 1164.41 | 13 / 12 | 6.1 | No |
| 555101 | 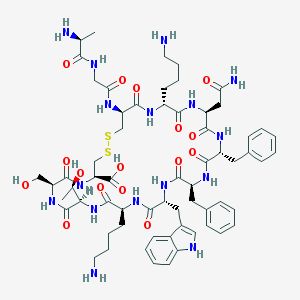 SS14 SS14 | C63H88N16O16S2 | 1389.61 | 21 / 19 | -4.1 | No |
| 383352 | 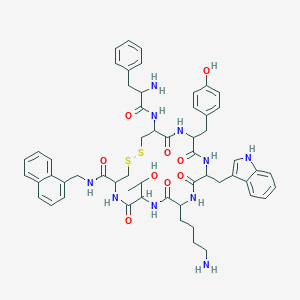 BDBM84630 BDBM84630 | C56H66N10O9S2 | 1087.32 | 13 / 12 | 4.2 | No |
| 390652 | 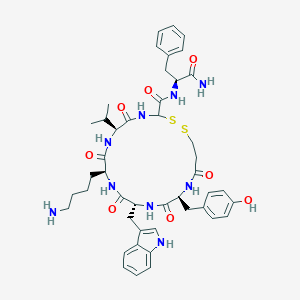 BDBM82456 BDBM82456 | C45H57N9O8S2 | 916.126 | 11 / 10 | 3.0 | No |
| 394983 | 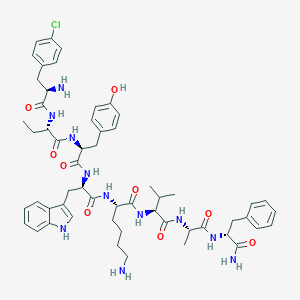 BDBM84627 BDBM84627 | C56H72ClN11O9 | 1078.71 | 11 / 12 | 4.2 | No |
| 400290 | 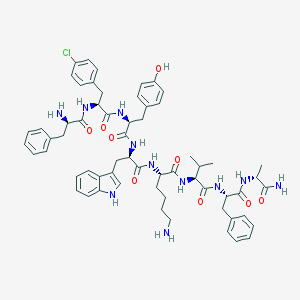 CHEMBL264562 CHEMBL264562 | C61H74ClN11O9 | 1140.78 | 11 / 12 | 5.1 | No |
| 416684 | 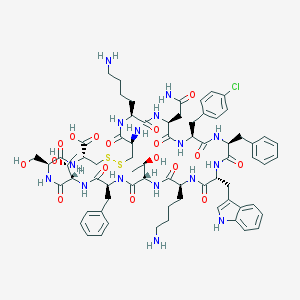 Bim-23003 Bim-23003 | C71H95ClN16O17S2 | 1544.21 | 22 / 20 | -1.6 | No |
| 435785 | 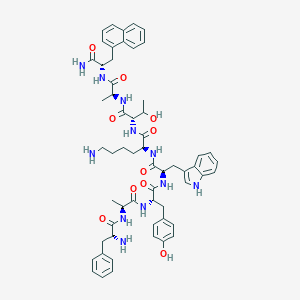 BDBM84638 BDBM84638 | C58H71N11O10 | 1082.27 | 12 / 13 | 3.4 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218