

You can:
| Name | Prostaglandin E2 receptor EP1 subtype |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | PTGER1 |
| Synonym | PGE receptor EP1 subtype EP1 receptor prostaglandin E receptor 1 (subtype EP1), 42kDa Prostanoid EP1 receptor EP1 prostanoid receptor [ Show all ] |
| Disease | Unspecified Thrombosis Pollakiuria Pain |
| Length | 402 |
| Amino acid sequence | MSPCGPLNLSLAGEATTCAAPWVPNTSAVPPSGASPALPIFSMTLGAVSNLLALALLAQAAGRLRRRRSAATFLLFVASLLATDLAGHVIPGALVLRLYTAGRAPAGGACHFLGGCMVFFGLCPLLLGCGMAVERCVGVTRPLLHAARVSVARARLALAAVAAVALAVALLPLARVGRYELQYPGTWCFIGLGPPGGWRQALLAGLFASLGLVALLAALVCNTLSGLALLRARWRRRSRRPPPASGPDSRRRWGAHGPRSASASSASSIASASTFFGGSRSSGSARRARAHDVEMVGQLVGIMVVSCICWSPMLVLVALAVGGWSSTSLQRPLFLAVRLASWNQILDPWVYILLRQAVLRQLLRLLPPRAGAKGGPAGLGLTPSAWEASSLRSSRHSGLSHF |
| UniProt | P34995 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | P34995 |
| 3D structure model | This predicted structure model is from GPCR-EXP P34995. |
| BioLiP | N/A |
| Therapeutic Target Database | T15497 |
| ChEMBL | CHEMBL1811 |
| IUPHAR | 340 |
| DrugBank | BE0000064 |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 121 | 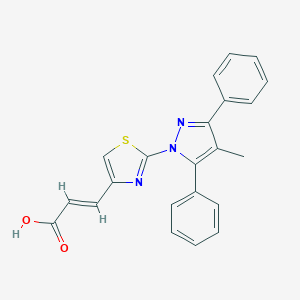 SCHEMBL2841361 SCHEMBL2841361 | C22H17N3O2S | 387.457 | 5 / 1 | 4.9 | Yes |
| 171 | 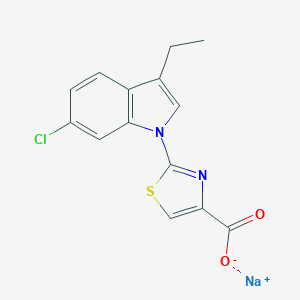 CHEMBL404176 CHEMBL404176 | C14H10ClN2NaO2S | 328.746 | 4 / 0 | N/A | N/A |
| 429 | 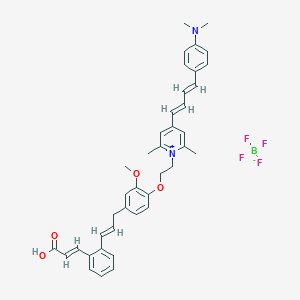 CHEMBL2164612 CHEMBL2164612 | C40H43BF4N2O4 | 702.598 | 10 / 1 | N/A | No |
| 855 | 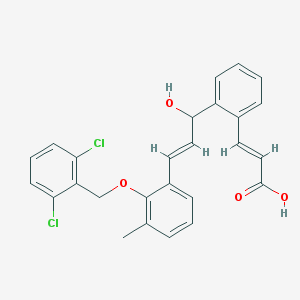 CHEMBL180742 CHEMBL180742 | C26H22Cl2O4 | 469.358 | 4 / 2 | 6.4 | No |
| 883 | 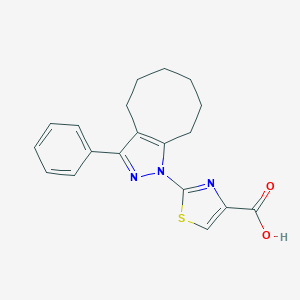 CHEMBL3127149 CHEMBL3127149 | C19H19N3O2S | 353.44 | 5 / 1 | 5.1 | No |
| 1260 | 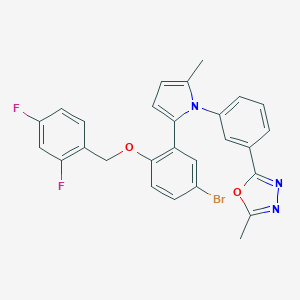 CHEMBL233160 CHEMBL233160 | C27H20BrF2N3O2 | 536.377 | 6 / 0 | 6.6 | No |
| 1596 | 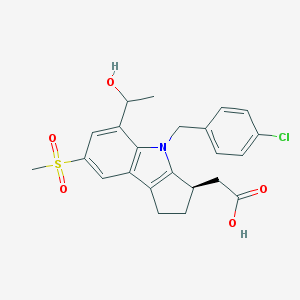 CHEMBL207203 CHEMBL207203 | C23H24ClNO5S | 461.957 | 5 / 2 | 3.0 | Yes |
| 1824 | 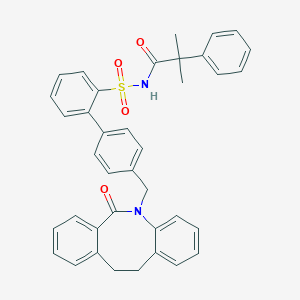 CHEMBL314533 CHEMBL314533 | C38H34N2O4S | 614.76 | 4 / 1 | 7.6 | No |
| 2425 | 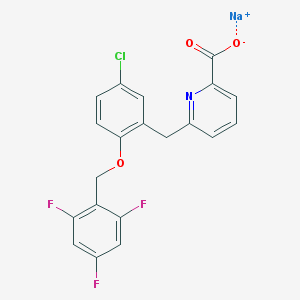 CHEMBL466102 CHEMBL466102 | C20H12ClF3NNaO3 | 429.755 | 7 / 0 | N/A | N/A |
| 3053 | 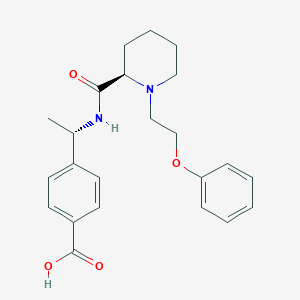 CHEMBL3115074 CHEMBL3115074 | C23H28N2O4 | 396.487 | 5 / 2 | 1.3 | Yes |
| 3369 | 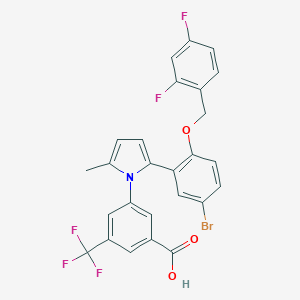 CHEMBL388243 CHEMBL388243 | C26H17BrF5NO3 | 566.322 | 8 / 1 | 7.2 | No |
| 4216 | 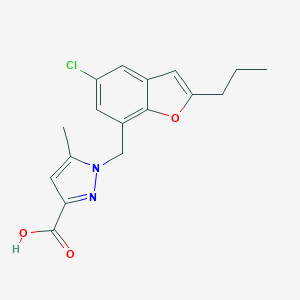 CHEMBL1915014 CHEMBL1915014 | C17H17ClN2O3 | 332.784 | 4 / 1 | 4.3 | Yes |
| 4314 | 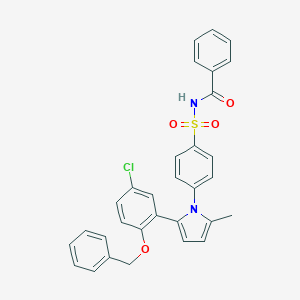 CHEMBL232543 CHEMBL232543 | C31H25ClN2O4S | 557.061 | 4 / 1 | 6.9 | No |
| 4552 | 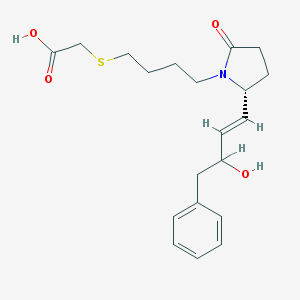 CHEMBL430124 CHEMBL430124 | C20H27NO4S | 377.499 | 5 / 2 | 2.4 | Yes |
| 4696 | 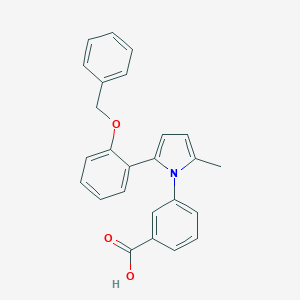 632621-53-5 632621-53-5 | C25H21NO3 | 383.447 | 3 / 1 | 5.4 | No |
| 521599 | 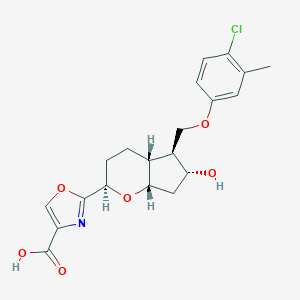 CHEMBL3793009 CHEMBL3793009 | C20H22ClNO6 | 407.847 | 7 / 2 | 3.2 | Yes |
| 5040 | 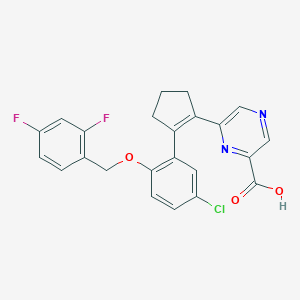 CHEMBL389996 CHEMBL389996 | C23H17ClF2N2O3 | 442.847 | 7 / 1 | 4.3 | Yes |
| 5460 | 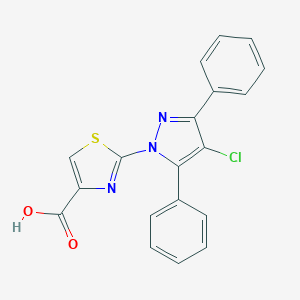 CHEMBL3092152 CHEMBL3092152 | C19H12ClN3O2S | 381.834 | 5 / 1 | 5.1 | No |
| 536082 | 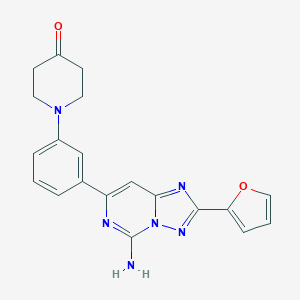 CHEMBL354419 CHEMBL354419 | C20H18N6O2 | 374.404 | 7 / 1 | 1.8 | Yes |
| 5827 | 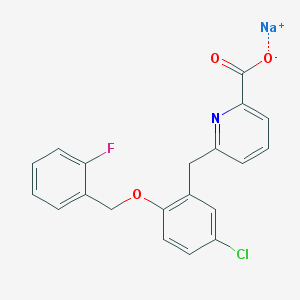 CHEMBL508276 CHEMBL508276 | C20H14ClFNNaO3 | 393.774 | 5 / 0 | N/A | N/A |
| 6059 | 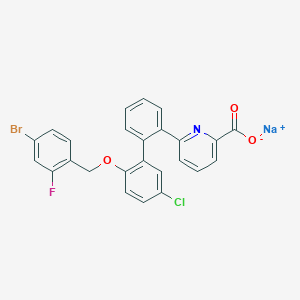 CHEMBL467578 CHEMBL467578 | C25H15BrClFNNaO3 | 534.741 | 5 / 0 | N/A | No |
| 6151 | 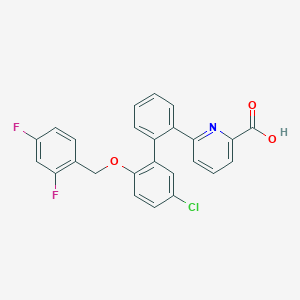 CHEMBL514574 CHEMBL514574 | C25H16ClF2NO3 | 451.854 | 6 / 1 | 6.3 | No |
| 6567 | 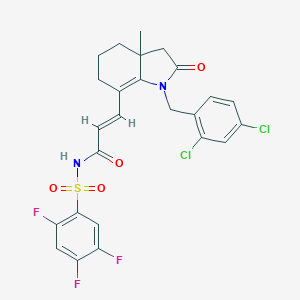 CHEMBL479439 CHEMBL479439 | C25H21Cl2F3N2O4S | 573.408 | 7 / 1 | 4.8 | No |
| 6659 | 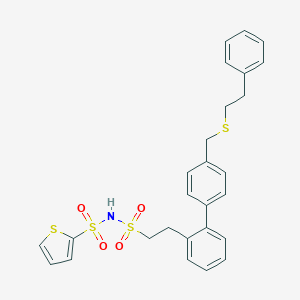 CHEMBL313266 CHEMBL313266 | C27H27NO4S4 | 557.756 | 7 / 1 | 6.5 | No |
| 6859 | 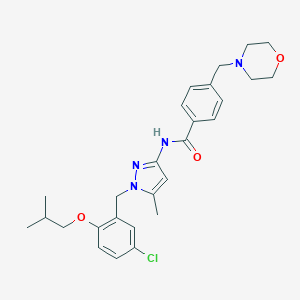 CHEMBL521988 CHEMBL521988 | C27H33ClN4O3 | 497.036 | 5 / 1 | 4.8 | Yes |
| 6892 | 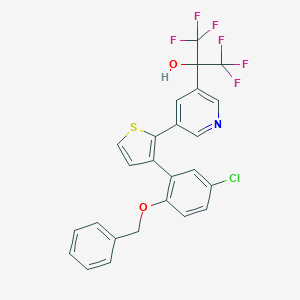 CHEMBL182367 CHEMBL182367 | C25H16ClF6NO2S | 543.908 | 10 / 1 | 7.1 | No |
| 7979 | 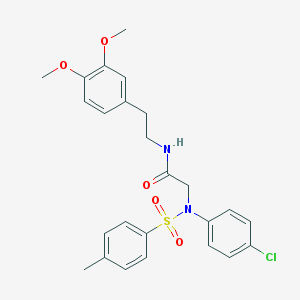 AC1MVS18 AC1MVS18 | C25H27ClN2O5S | 503.01 | 6 / 1 | 4.9 | No |
| 7999 | 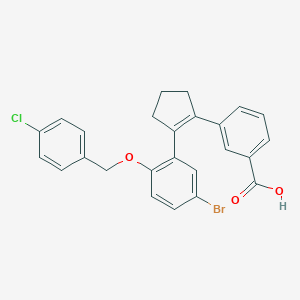 CHEMBL428370 CHEMBL428370 | C25H20BrClO3 | 483.786 | 3 / 1 | 6.6 | No |
| 8520 | 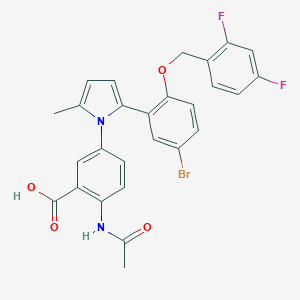 CHEMBL387969 CHEMBL387969 | C27H21BrF2N2O4 | 555.376 | 6 / 2 | 6.0 | No |
| 536211 | 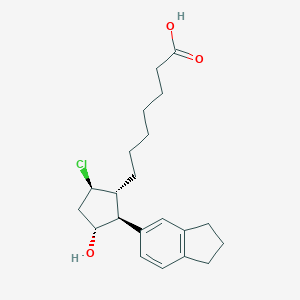 SCHEMBL5208850 SCHEMBL5208850 | C21H29ClO3 | 364.91 | 3 / 2 | 5.1 | No |
| 9113 | 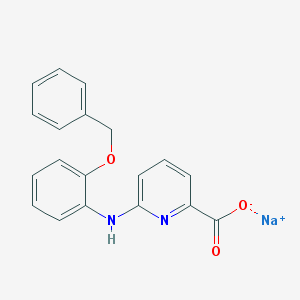 CHEMBL257792 CHEMBL257792 | C19H15N2NaO3 | 342.33 | 5 / 1 | N/A | N/A |
| 9198 | 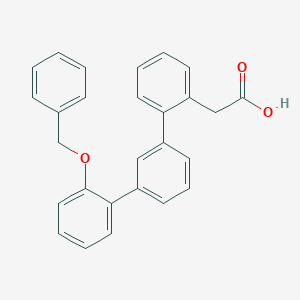 CHEMBL123844 CHEMBL123844 | C27H22O3 | 394.47 | 3 / 1 | 5.3 | No |
| 10345 | 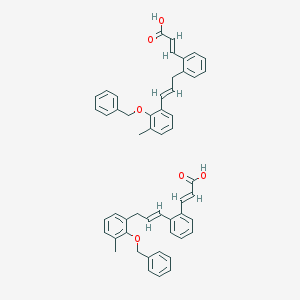 CID 52944193 CID 52944193 | C52H48O6 | 768.95 | 6 / 2 | N/A | No |
| 10868 | 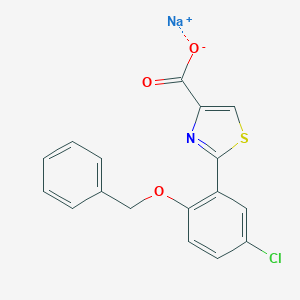 CHEMBL270162 CHEMBL270162 | C17H11ClNNaO3S | 367.779 | 5 / 0 | N/A | N/A |
| 11216 | 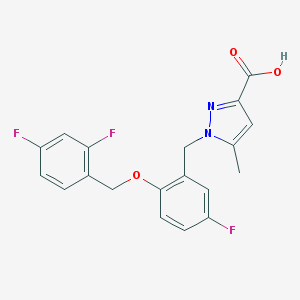 CHEMBL214914 CHEMBL214914 | C19H15F3N2O3 | 376.335 | 7 / 1 | 3.9 | Yes |
| 11427 | 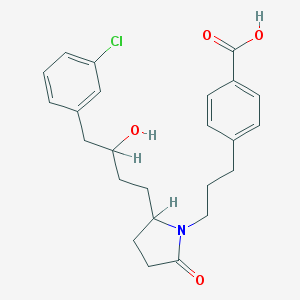 CHEMBL383515 CHEMBL383515 | C24H28ClNO4 | 429.941 | 4 / 2 | 4.4 | Yes |
| 11633 | 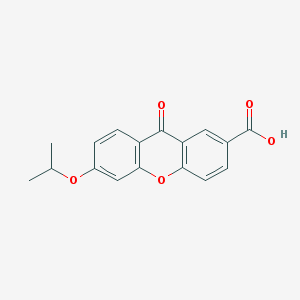 AH 6809 AH 6809 | C17H14O5 | 298.294 | 5 / 1 | 3.9 | Yes |
| 11959 | 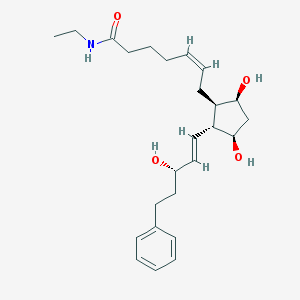 Bimatoprost Bimatoprost | C25H37NO4 | 415.574 | 4 / 4 | 2.8 | Yes |
| 12367 | 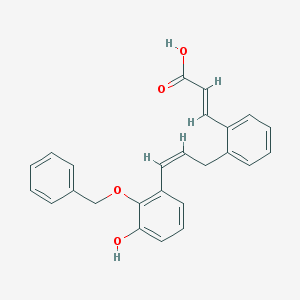 BDBM50370468 BDBM50370468 | C25H22O4 | 386.447 | 4 / 2 | 5.5 | No |
| 12574 | 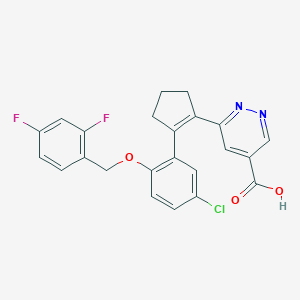 CHEMBL233245 CHEMBL233245 | C23H17ClF2N2O3 | 442.847 | 7 / 1 | 4.1 | Yes |
| 553315 | 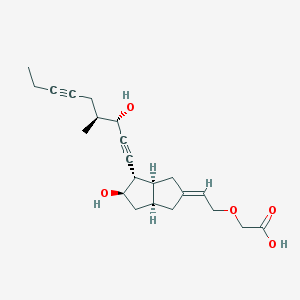 Cicaprost Cicaprost | C22H30O5 | 374.477 | 5 / 3 | 2.1 | Yes |
| 12776 | 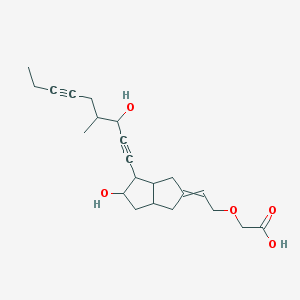 CID 53394056 CID 53394056 | C22H30O5 | 374.477 | 5 / 3 | 2.1 | Yes |
| 12830 | 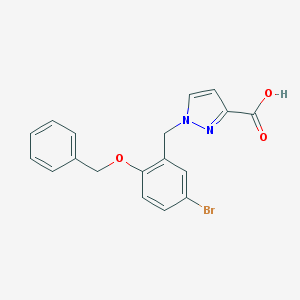 CHEMBL213412 CHEMBL213412 | C18H15BrN2O3 | 387.233 | 4 / 1 | 3.8 | Yes |
| 13139 | 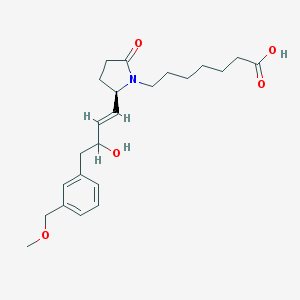 CHEMBL275245 CHEMBL275245 | C23H33NO5 | 403.519 | 5 / 2 | 2.2 | Yes |
| 13563 | 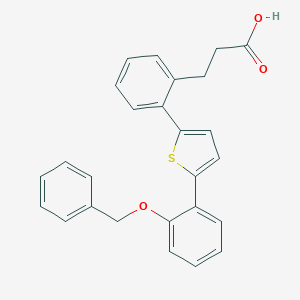 CHEMBL125110 CHEMBL125110 | C26H22O3S | 414.519 | 4 / 1 | 6.1 | No |
| 464550 | 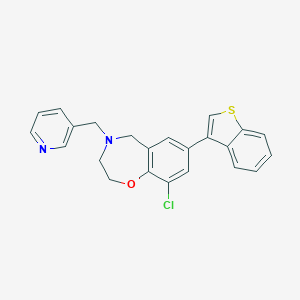 CHEMBL3586353 CHEMBL3586353 | C23H19ClN2OS | 406.928 | 4 / 0 | 5.2 | No |
| 13899 | 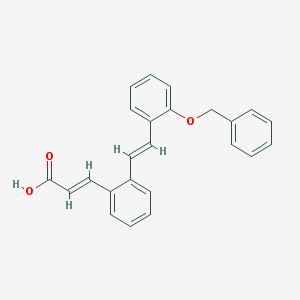 CHEMBL181035 CHEMBL181035 | C24H20O3 | 356.421 | 3 / 1 | 5.6 | No |
| 14159 | 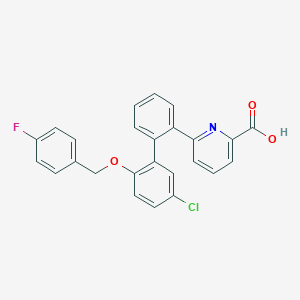 CHEMBL457142 CHEMBL457142 | C25H17ClFNO3 | 433.863 | 5 / 1 | 6.2 | No |
| 14343 | 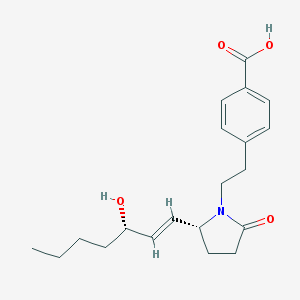 CHEMBL251294 CHEMBL251294 | C20H27NO4 | 345.439 | 4 / 2 | 2.8 | Yes |
| 14416 | 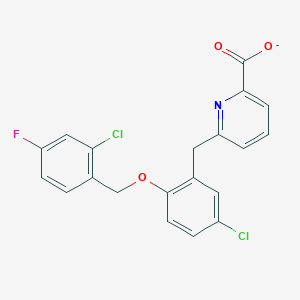 CHEMBL513491 CHEMBL513491 | C20H13Cl2FNO3- | 405.226 | 5 / 0 | 6.1 | No |
| 442243 | 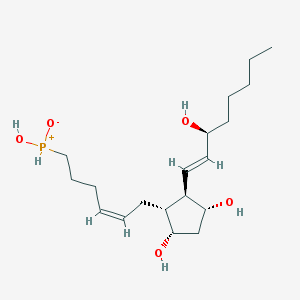 [(5Z,8R,9S,11R,12R,13E,15S)-9,11,15-Trihydroxy-1-norprosta-5,13-diene-2-yl]phosphinic acid [(5Z,8R,9S,11R,12R,13E,15S)-9,11,15-Trihydroxy-1-norprosta-5,13-diene-2-yl]phosphinic acid | C19H35O5P | 374.458 | 5 / 4 | 1.9 | Yes |
| 15306 | 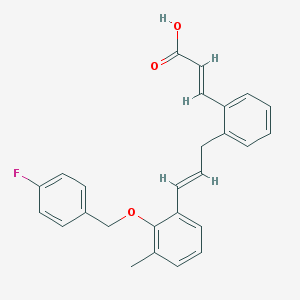 CHEMBL180089 CHEMBL180089 | C26H23FO3 | 402.465 | 4 / 1 | 6.4 | No |
| 15369 | 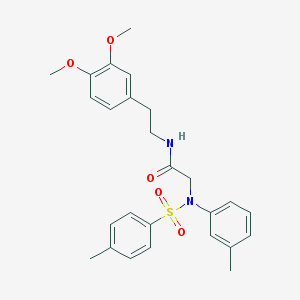 AC1MVUYC AC1MVUYC | C26H30N2O5S | 482.595 | 6 / 1 | 4.6 | Yes |
| 15659 | 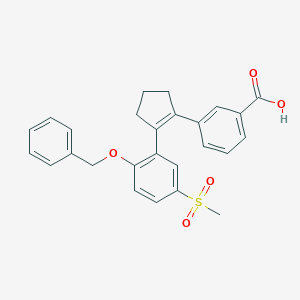 CHEMBL265086 CHEMBL265086 | C26H24O5S | 448.533 | 5 / 1 | 4.5 | Yes |
| 15827 | 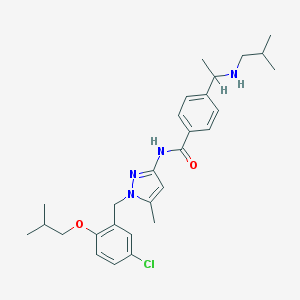 CHEMBL494268 CHEMBL494268 | C28H37ClN4O2 | 497.08 | 4 / 2 | 6.4 | No |
| 16164 | 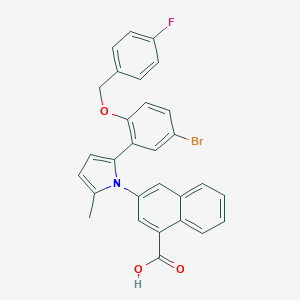 CHEMBL228746 CHEMBL228746 | C29H21BrFNO3 | 530.393 | 4 / 1 | 7.4 | No |
| 16331 | 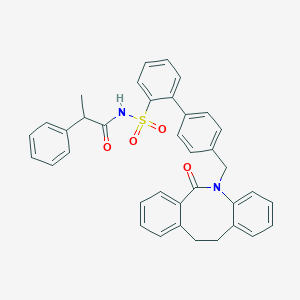 CHEMBL90491 CHEMBL90491 | C37H32N2O4S | 600.733 | 4 / 1 | 7.3 | No |
| 17049 | 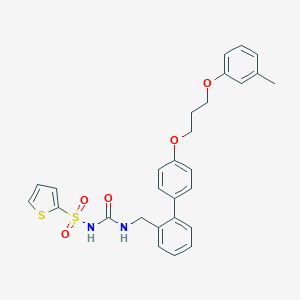 CHEMBL87797 CHEMBL87797 | C28H28N2O5S2 | 536.661 | 6 / 2 | 6.0 | No |
| 17508 | 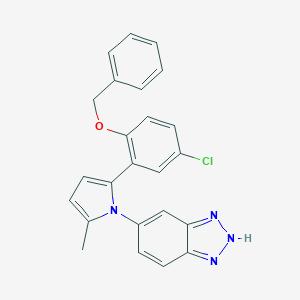 CHEMBL233975 CHEMBL233975 | C24H19ClN4O | 414.893 | 3 / 1 | 5.6 | No |
| 17843 | 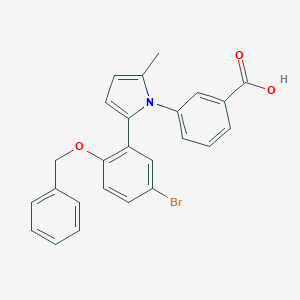 CHEMBL377649 CHEMBL377649 | C25H20BrNO3 | 462.343 | 3 / 1 | 6.1 | No |
| 18155 | 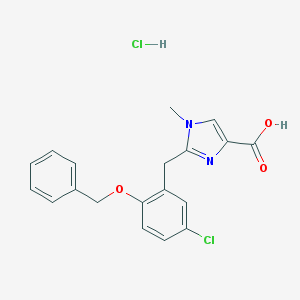 CHEMBL535640 CHEMBL535640 | C19H18Cl2N2O3 | 393.264 | 4 / 2 | N/A | N/A |
| 19180 | 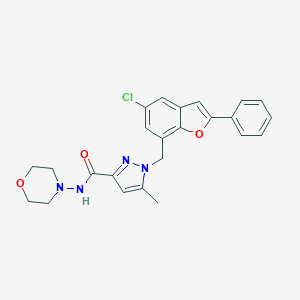 CHEMBL1915264 CHEMBL1915264 | C24H23ClN4O3 | 450.923 | 5 / 1 | 4.4 | Yes |
| 557866 | 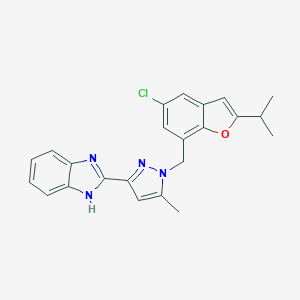 CHEMBL1915245 CHEMBL1915245 | C23H21ClN4O | 404.898 | 3 / 1 | 5.7 | No |
| 19721 | 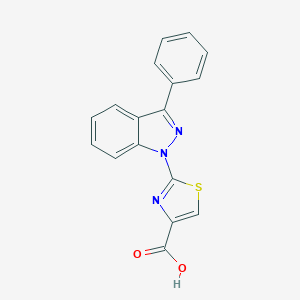 SCHEMBL1513030 SCHEMBL1513030 | C17H11N3O2S | 321.354 | 5 / 1 | 4.1 | Yes |
| 19963 | 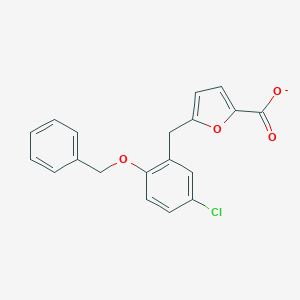 CHEMBL272793 CHEMBL272793 | C19H14ClO4- | 341.767 | 4 / 0 | 5.5 | No |
| 20027 | 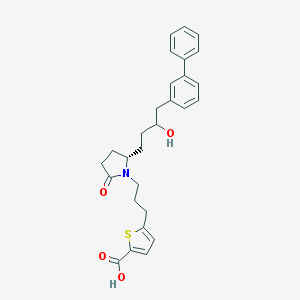 CHEMBL382197 CHEMBL382197 | C28H31NO4S | 477.619 | 5 / 2 | 5.4 | No |
| 21559 | 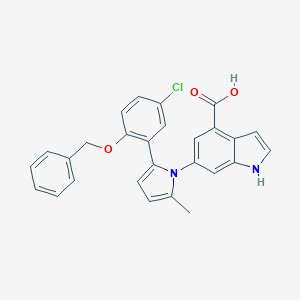 CHEMBL228963 CHEMBL228963 | C27H21ClN2O3 | 456.926 | 3 / 2 | 6.1 | No |
| 21797 | 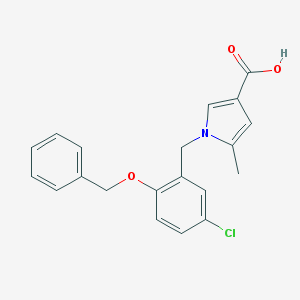 CHEMBL403459 CHEMBL403459 | C20H18ClNO3 | 355.818 | 3 / 1 | 4.3 | Yes |
| 22053 | 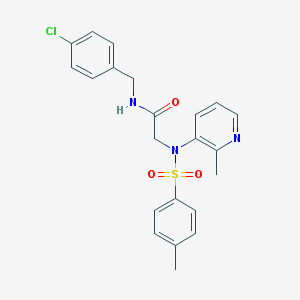 CHEMBL240866 CHEMBL240866 | C22H22ClN3O3S | 443.946 | 5 / 1 | 3.8 | Yes |
| 22148 | 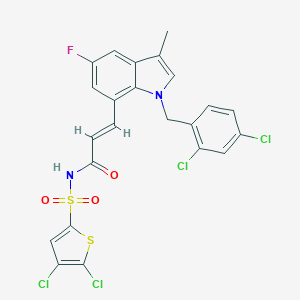 DG-041 DG-041 | C23H15Cl4FN2O3S2 | 592.302 | 5 / 1 | 7.8 | No |
| 536549 | 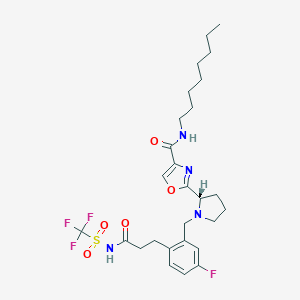 CHEMBL3896035 CHEMBL3896035 | C27H36F4N4O5S | 604.662 | 11 / 2 | 5.6 | No |
| 22283 | 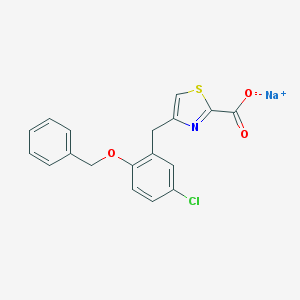 CHEMBL258057 CHEMBL258057 | C18H13ClNNaO3S | 381.806 | 5 / 0 | N/A | N/A |
| 22808 | 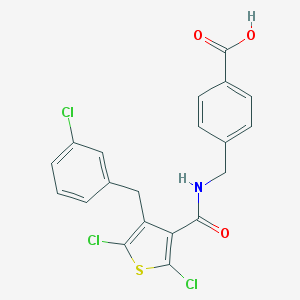 CHEMBL601299 CHEMBL601299 | C20H14Cl3NO3S | 454.746 | 4 / 2 | 6.4 | No |
| 23007 | 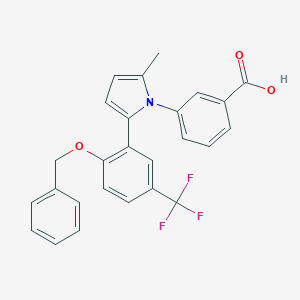 CHEMBL212688 CHEMBL212688 | C26H20F3NO3 | 451.445 | 6 / 1 | 6.3 | No |
| 23671 | 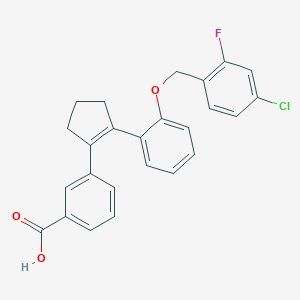 CHEMBL388655 CHEMBL388655 | C25H20ClFO3 | 422.88 | 4 / 1 | 6.0 | No |
| 459420 | 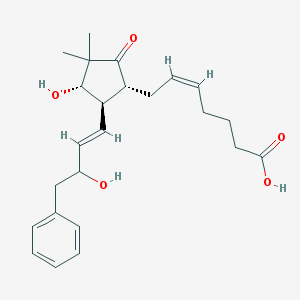 SCHEMBL2194130 SCHEMBL2194130 | C24H32O5 | 400.515 | 5 / 3 | 3.0 | Yes |
| 23779 | 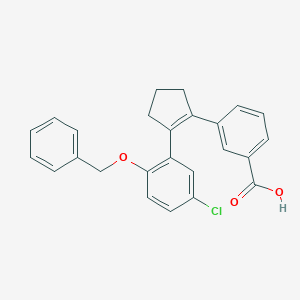 CHEMBL207577 CHEMBL207577 | C25H21ClO3 | 404.89 | 3 / 1 | 5.9 | No |
| 553377 | 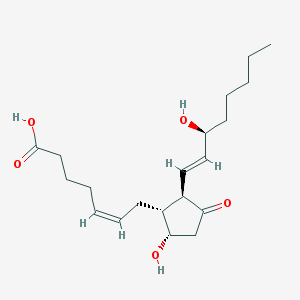 prostaglandin D2 prostaglandin D2 | C20H32O5 | 352.471 | 5 / 3 | 2.6 | Yes |
| 24342 | 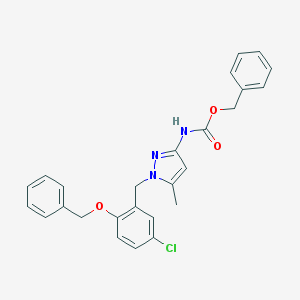 CHEMBL254963 CHEMBL254963 | C26H24ClN3O3 | 461.946 | 4 / 1 | 5.7 | No |
| 24511 | 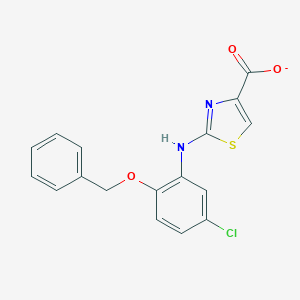 CHEMBL401768 CHEMBL401768 | C17H12ClN2O3S- | 359.804 | 6 / 1 | 5.4 | No |
| 465997 | 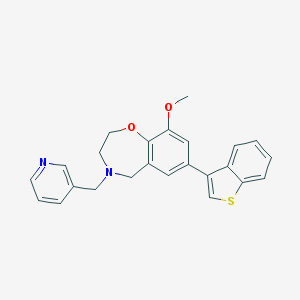 CHEMBL3586309 CHEMBL3586309 | C24H22N2O2S | 402.512 | 5 / 0 | 4.5 | Yes |
| 25216 | 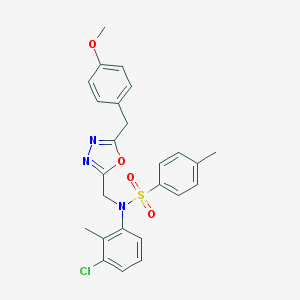 CHEMBL399613 CHEMBL399613 | C25H24ClN3O4S | 497.994 | 7 / 0 | 5.3 | No |
| 536648 | 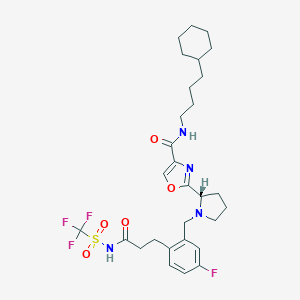 SCHEMBL671511 SCHEMBL671511 | C29H38F4N4O5S | 630.7 | 11 / 2 | 6.2 | No |
| 536673 | 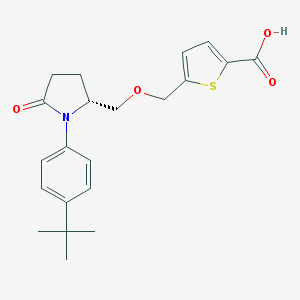 SCHEMBL1309617 SCHEMBL1309617 | C21H25NO4S | 387.494 | 5 / 1 | 4.0 | Yes |
| 26457 | 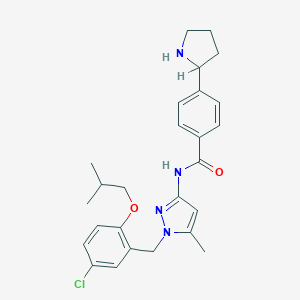 CHEMBL494466 CHEMBL494466 | C26H31ClN4O2 | 467.01 | 4 / 2 | 5.2 | No |
| 27147 | 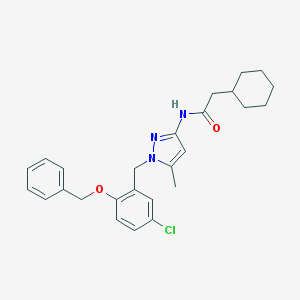 CHEMBL404297 CHEMBL404297 | C26H30ClN3O2 | 451.995 | 3 / 1 | 6.5 | No |
| 27458 | 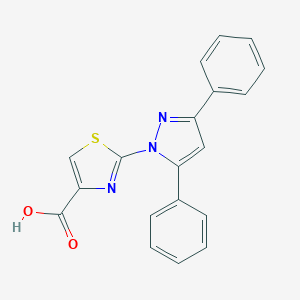 SCHEMBL1548505 SCHEMBL1548505 | C19H13N3O2S | 347.392 | 5 / 1 | 4.4 | Yes |
| 27564 | 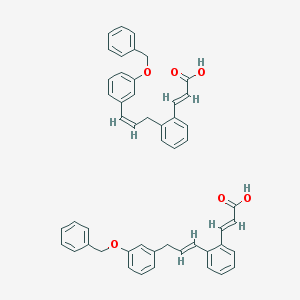 CID 52945421 CID 52945421 | C50H44O6 | 740.896 | 6 / 2 | N/A | No |
| 28098 | 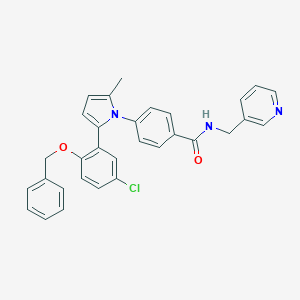 CHEMBL401023 CHEMBL401023 | C31H26ClN3O2 | 508.018 | 3 / 1 | 6.2 | No |
| 28504 | 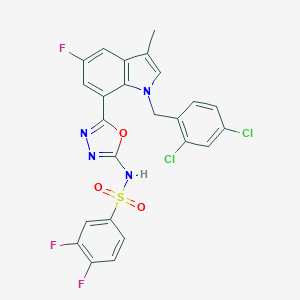 CHEMBL1080281 CHEMBL1080281 | C24H15Cl2F3N4O3S | 567.364 | 9 / 1 | 5.9 | No |
| 28690 | 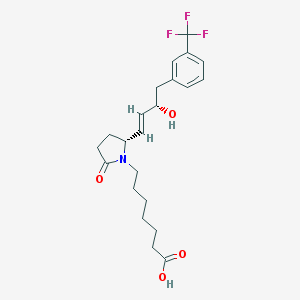 CHEMBL45008 CHEMBL45008 | C22H28F3NO4 | 427.464 | 7 / 2 | 3.4 | Yes |
| 28691 | 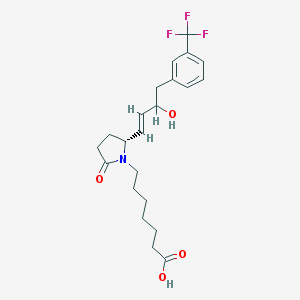 CHEMBL440474 CHEMBL440474 | C22H28F3NO4 | 427.464 | 7 / 2 | 3.4 | Yes |
| 522427 | 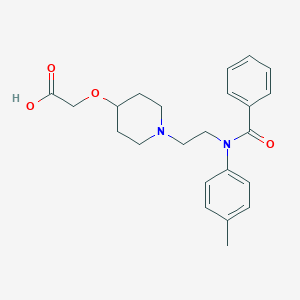 CHEMBL3793911 CHEMBL3793911 | C23H28N2O4 | 396.487 | 5 / 1 | 1.1 | Yes |
| 522431 | 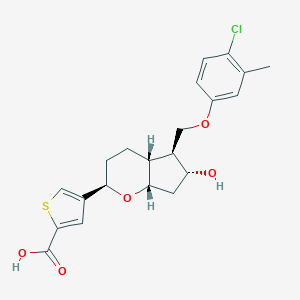 CHEMBL3794185 CHEMBL3794185 | C21H23ClO5S | 422.92 | 6 / 2 | 4.4 | Yes |
| 29952 | 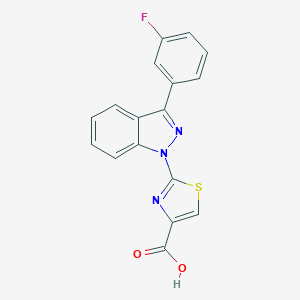 CHEMBL3127165 CHEMBL3127165 | C17H10FN3O2S | 339.344 | 6 / 1 | 4.2 | Yes |
| 31005 | 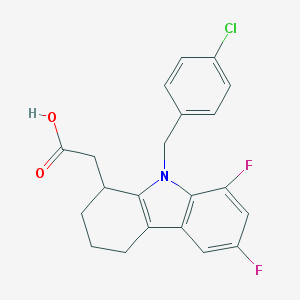 CHEMBL208260 CHEMBL208260 | C21H18ClF2NO2 | 389.827 | 4 / 1 | 4.9 | Yes |
| 31151 | 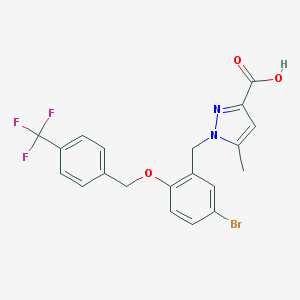 CHEMBL215956 CHEMBL215956 | C20H16BrF3N2O3 | 469.258 | 7 / 1 | 5.1 | No |
| 32575 | 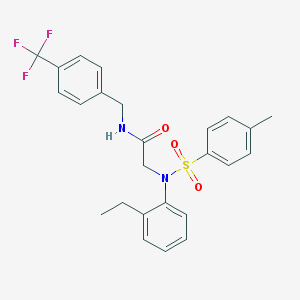 CHEMBL240457 CHEMBL240457 | C25H25F3N2O3S | 490.541 | 7 / 1 | 5.5 | No |
| 32682 | 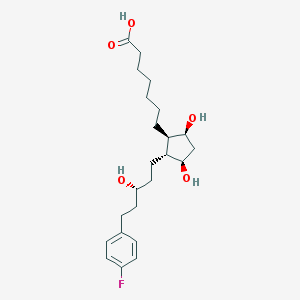 CHEMBL36911 CHEMBL36911 | C23H35FO5 | 410.526 | 6 / 4 | 4.0 | Yes |
| 32785 | 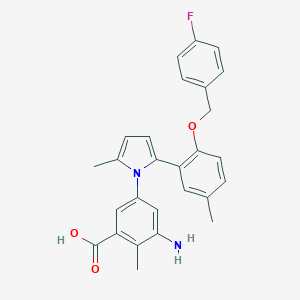 CHEMBL388510 CHEMBL388510 | C27H25FN2O3 | 444.506 | 5 / 2 | 5.5 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218