

You can:
| Name | Corticotropin-releasing factor receptor 1 |
|---|---|
| Species | Mus musculus (Mouse) |
| Gene | Crhr1 |
| Synonym | CRH-R1 CRH-R-1 CRFR1 CRFR-1 CRF1 receptor [ Show all ] |
| Disease | N/A for non-human GPCRs |
| Length | 415 |
| Amino acid sequence | MGQRPQLRLVKALLLLGLNPVSTSLQDQQCESLSLASNVSGLQCNASVDLIGTCWPRSPAGQLVVRPCPAFFYGVRYNTTNNGYRECLANGSWAARVNYSECQEILNEEKKSKVHYHIAVIINYLGHCISLVALLVAFVLFLRLRSIRCLRNIIHWNLISAFILRNATWFVVQLTVSPEVHQSNVAWCRLVTAAYNYFHVTNFFWMFGEGCYLHTAIVLTYSTDRLRKWMFVCIGWGVPFPIIVAWAIGKLYYDNEKCWFGKRPGVYTDYIYQGPMILVLLINFIFLFNIVRILMTKLRASTTSETIQYRKAVKATLVLLPLLGITYMLFFVNPGEDEVSRVVFIYFNSFLESFQGFFVSVFYCFLNSEVRSAIRKRWRRWQDKHSIRARVARAMSIPTSPTRVSFHSIKQSTAV |
| UniProt | P35347 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL2446 |
| IUPHAR | N/A |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 1269 |  CHEMBL430078 CHEMBL430078 | C213H348N56O61S | 4701.52 | 68 / 60 | -16.1 | No |
| 2133 |  CID 77232001 CID 77232001 | C208H345N57O61S | 4652.45 | 69 / 60 | -18.0 | No |
| 8146 |  CHEMBL429069 CHEMBL429069 | C209H347N57O62S | 4682.47 | 69 / 61 | -18.8 | No |
| 10094 |  CID 44388678 CID 44388678 | C204H349N55O61S | 4580.42 | 68 / 59 | -17.1 | No |
| 11459 |  CID 44388704 CID 44388704 | C208H349N55O62S | 4644.46 | 69 / 60 | -16.8 | No |
| 16843 |  CHEMBL411590 CHEMBL411590 | C217H350N56O61S | 4751.58 | 68 / 60 | -14.8 | No |
| 19066 |  BDBM50158991 BDBM50158991 | C212H351N55O61S | 4678.52 | 68 / 61 | -14.5 | No |
| 20089 |  BDBM50158995 BDBM50158995 | C207H347N55O62S | 4630.44 | 69 / 62 | -16.4 | No |
| 22592 |  CID 44388705 CID 44388705 | C201H343N55O61S | 4538.34 | 68 / 59 | -18.5 | No |
| 39684 |  CID 77231998 CID 77231998 | C210H346N58O61S | 4691.48 | 69 / 61 | -17.9 | No |
| 53117 |  CID 44388647 CID 44388647 | C203H345N57O63S | 4624.39 | 70 / 61 | -21.7 | No |
| 55732 |  CID 44388723 CID 44388723 | C206H345N55O63S | 4632.41 | 70 / 61 | -18.8 | No |
| 65405 |  CID 44388688 CID 44388688 | C205H351N55O61S | 4594.45 | 68 / 59 | -16.8 | No |
| 70728 |  CID 77231997 CID 77231997 | C208H345N57O62S | 4668.44 | 70 / 61 | -18.4 | No |
| 78786 |  BDBM50158965 BDBM50158965 | C207H346N56O62S | 4643.44 | 69 / 62 | -18.2 | No |
| 89874 |  CID 44388677 CID 44388677 | C204H348N56O62S | 4609.42 | 69 / 60 | -19.2 | No |
| 100686 | 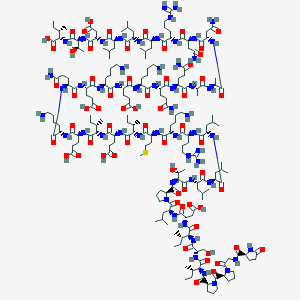 BDBM50159003 BDBM50159003 | C203H347N55O62S | 4582.39 | 69 / 62 | -17.9 | No |
| 108276 | 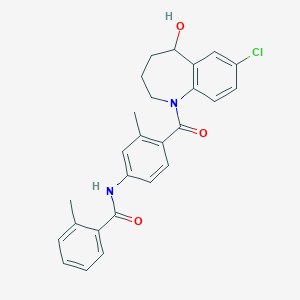 Tolvaptan Tolvaptan | C26H25ClN2O3 | 448.947 | 3 / 2 | 4.8 | Yes |
| 111626 | 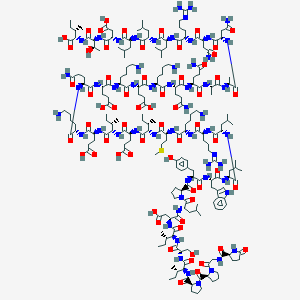 CHEMBL438220 CHEMBL438220 | C213H348N56O62S | 4717.52 | 69 / 61 | -16.4 | No |
| 112471 | 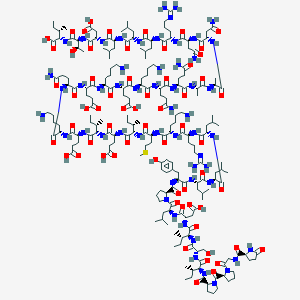 CID 44388711 CID 44388711 | C208H349N55O62S | 4644.46 | 69 / 60 | -16.8 | No |
| 525037 |  CHEMBL3775924 CHEMBL3775924 | C161H269N49O42 | 3563.22 | 49 / 51 | -8.0 | No |
| 141249 |  BDBM50158970 BDBM50158970 | C205H351N55O61S | 4594.45 | 68 / 61 | -16.0 | No |
| 142266 |  CID 44388680 CID 44388680 | C215H349N55O61S | 4712.54 | 68 / 59 | -15.0 | No |
| 145406 |  CID 44388679 CID 44388679 | C211H347N55O61S | 4662.48 | 68 / 59 | -16.2 | No |
| 152167 |  CID 44388608 CID 44388608 | C203H347N55O62S | 4582.39 | 69 / 60 | -18.7 | No |
| 158457 |  BDBM50158961 BDBM50158961 | C208H349N55O61S | 4628.46 | 68 / 61 | -15.7 | No |
| 159038 |  CID 44388623 CID 44388623 | C208H349N55O61S | 4628.46 | 68 / 59 | -16.5 | No |
| 162052 |  BDBM50158969 BDBM50158969 | C205H351N55O61S | 4594.45 | 68 / 61 | -16.0 | No |
| 167961 |  BDBM50158974 BDBM50158974 | C205H343N55O61S | 4586.38 | 68 / 61 | -17.1 | No |
| 168113 |  BDBM50158983 BDBM50158983 | C202H345N55O64S | 4600.36 | 71 / 64 | -19.2 | No |
| 182978 |  BDBM50158998 BDBM50158998 | C204H347N55O63S | 4610.4 | 70 / 62 | -17.8 | No |
| 184215 |  BDBM50158968 BDBM50158968 | C211H348N56O62S | 4693.5 | 69 / 62 | -17.0 | No |
| 186171 |  CID 44388638 CID 44388638 | C205H343N55O62S | 4602.38 | 69 / 60 | -18.2 | No |
| 193525 |  BDBM50158962 BDBM50158962 | C204H348N56O62S | 4609.42 | 69 / 62 | -18.5 | No |
| 205091 |  CID 44388710 CID 44388710 | C211H347N55O62S | 4678.48 | 69 / 60 | -16.6 | No |
| 206389 |  CID 44388681 CID 44388681 | C207H347N55O61S | 4614.44 | 68 / 59 | -16.8 | No |
| 206761 |  BDBM50158986 BDBM50158986 | C206H345N55O63S | 4632.41 | 70 / 63 | -18.0 | No |
| 209051 |  CID 44388685 CID 44388685 | C206H344N56O63S | 4645.41 | 70 / 61 | -19.7 | No |
| 233343 |  CHEMBL429547 CHEMBL429547 | C210H350N56O61S | 4667.5 | 68 / 60 | -16.4 | No |
| 257117 |  BDBM50158979 BDBM50158979 | C215H349N55O62S | 4728.54 | 69 / 62 | -14.6 | No |
| 259191 |  CID 77231999 CID 77231999 | C212H347N57O61S | 4702.51 | 69 / 60 | -16.8 | No |
| 260870 |  CID 44388659 CID 44388659 | C183H323N57O52 | 4153.94 | 58 / 58 | -10.2 | No |
| 264149 |  UNII-D8690V3244 UNII-D8690V3244 | C208H343N59O64S2 | 4758.5 | 73 / 67 | -13.6 | No |
| 277839 |  CID 44388687 CID 44388687 | C205H351N55O61S | 4594.45 | 68 / 59 | -16.8 | No |
| 311222 | 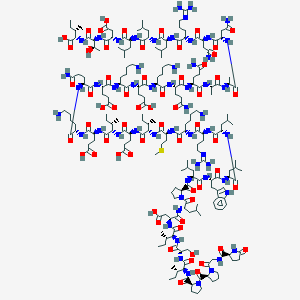 CHEMBL410347 CHEMBL410347 | C210H350N56O61S | 4667.5 | 68 / 60 | -16.4 | No |
| 530591 | 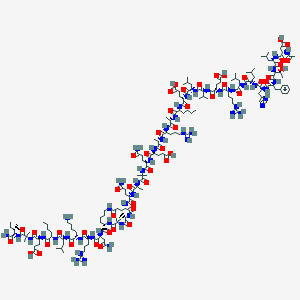 CHEMBL3774939 CHEMBL3774939 | C171H286N50O49 | 3826.47 | 54 / 55 | -8.2 | No |
| 336730 | 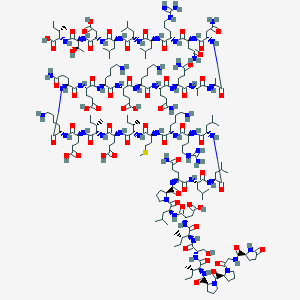 BDBM50158996 BDBM50158996 | C204H348N56O62S | 4609.42 | 69 / 62 | -18.5 | No |
| 531149 | 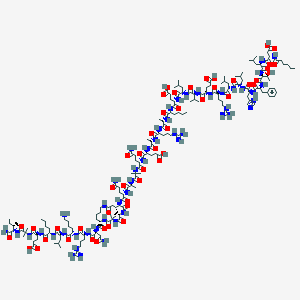 CHEMBL3774395 CHEMBL3774395 | C175H294N50O49 | 3882.58 | 54 / 55 | -6.3 | No |
| 356488 |  CID 44388696 CID 44388696 | C207H346N56O63S | 4659.43 | 70 / 61 | -19.3 | No |
| 364140 |  BDBM50158957 BDBM50158957 | C208H349N55O62S | 4644.46 | 69 / 62 | -16.1 | No |
| 367113 |  CHEMBL414568 CHEMBL414568 | C208H346N56O62S | 4655.45 | 69 / 61 | -18.3 | No |
| 369106 |  BDBM50159004 BDBM50159004 | C206H345N55O62S | 4616.41 | 69 / 62 | -17.7 | No |
| 384928 |  BDBM50158992 BDBM50158992 | C207H346N56O62S | 4643.44 | 69 / 62 | -18.2 | No |
| 387948 |  CID 77232002 CID 77232002 | C205H347N57O61S | 4618.43 | 69 / 60 | -18.3 | No |
| 388741 |  CID 77232000 CID 77232000 | C204H344N58O62S | 4633.4 | 70 / 61 | -20.8 | No |
| 389180 |  BDBM50158993 BDBM50158993 | C204H348N56O62S | 4609.42 | 69 / 62 | -18.5 | No |
| 403822 |  BDBM50158975 BDBM50158975 | C209H345N55O61S | 4636.44 | 68 / 61 | -15.8 | No |
| 404191 |  CID 44388720 CID 44388720 | C211H347N55O63S | 4694.48 | 70 / 61 | -16.9 | No |
| 533038 |  CHEMBL3775796 CHEMBL3775796 | C179H298N52O49 | 3962.67 | 55 / 56 | -6.0 | No |
| 434226 |  BDBM50158966 BDBM50158966 | C207H346N56O63S | 4659.43 | 70 / 63 | -18.6 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218