

You can:
| Name | P2Y purinoceptor 1 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | P2RY1 |
| Synonym | ATP receptor Purinergic receptor P2Y1 purinergic receptor P2Y Purinergic receptor platelet ADP receptor [ Show all ] |
| Disease | Thrombosis |
| Length | 373 |
| Amino acid sequence | MTEVLWPAVPNGTDAAFLAGPGSSWGNSTVASTAAVSSSFKCALTKTGFQFYYLPAVYILVFIIGFLGNSVAIWMFVFHMKPWSGISVYMFNLALADFLYVLTLPALIFYYFNKTDWIFGDAMCKLQRFIFHVNLYGSILFLTCISAHRYSGVVYPLKSLGRLKKKNAICISVLVWLIVVVAISPILFYSGTGVRKNKTITCYDTTSDEYLRSYFIYSMCTTVAMFCVPLVLILGCYGLIVRALIYKDLDNSPLRRKSIYLVIIVLTVFAVSYIPFHVMKTMNLRARLDFQTPAMCAFNDRVYATYQVTRGLASLNSCVDPILYFLAGDTFRRRLSRATRKASRRSEANLQSKSEDMTLNILPEFKQNGDTSL |
| UniProt | P47900 |
| Protein Data Bank | 4xnv, 4xnw |
| GPCR-HGmod model | P47900 |
| 3D structure model | This structure is from PDB ID 4xnv. |
| BioLiP | BL0311594,BL0311596, BL0311593, BL0311590,BL0311591,BL0311592, BL0311589, BL0311588, BL0311595,BL0311597 |
| Therapeutic Target Database | T67818 |
| ChEMBL | CHEMBL4315 |
| IUPHAR | 323 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 239 | 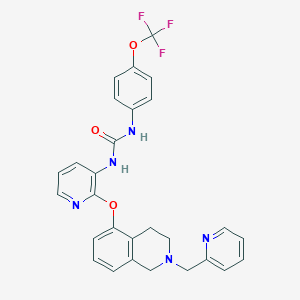 CHEMBL3092622 CHEMBL3092622 | C28H24F3N5O3 | 535.527 | 9 / 2 | 4.8 | No |
| 459238 | 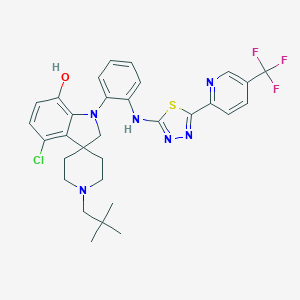 CHEMBL3896467 CHEMBL3896467 | C31H32ClF3N6OS | 629.143 | 11 / 2 | 7.8 | No |
| 636 | 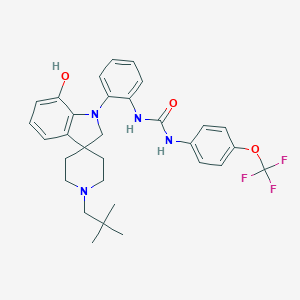 CHEMBL3287047 CHEMBL3287047 | C31H35F3N4O3 | 568.641 | 8 / 3 | 7.1 | No |
| 557332 | 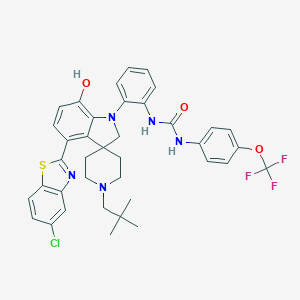 CHEMBL3901297 CHEMBL3901297 | C38H37ClF3N5O3S | 736.251 | 10 / 3 | 9.8 | No |
| 557343 | 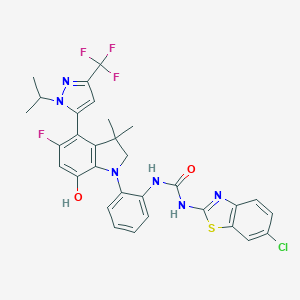 SCHEMBL15597014 SCHEMBL15597014 | C31H27ClF4N6O2S | 659.101 | 10 / 3 | 7.8 | No |
| 1978 | 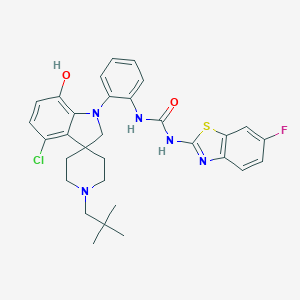 CHEMBL3287049 CHEMBL3287049 | C31H33ClFN5O2S | 594.146 | 7 / 3 | 7.4 | No |
| 533921 | 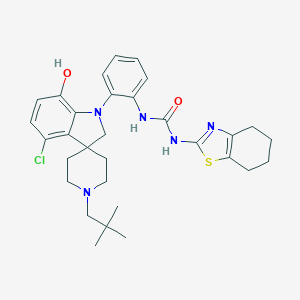 CHEMBL3977624 CHEMBL3977624 | C31H38ClN5O2S | 580.188 | 6 / 3 | 7.0 | No |
| 3031 | 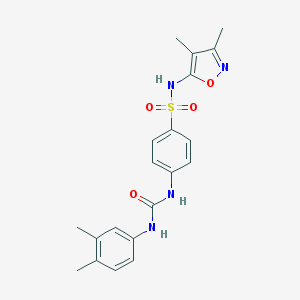 CHEMBL2071539 CHEMBL2071539 | C20H22N4O4S | 414.48 | 6 / 3 | 3.0 | Yes |
| 3211 | 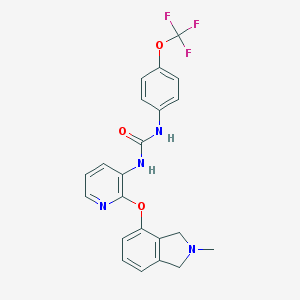 CHEMBL3092630 CHEMBL3092630 | C22H19F3N4O3 | 444.414 | 8 / 2 | 3.9 | Yes |
| 553259 | 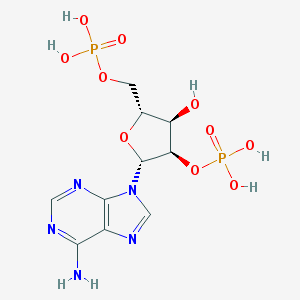 adenosine-2'-5'-diphosphate adenosine-2'-5'-diphosphate | C10H15N5O10P2 | 427.203 | 14 / 6 | -4.6 | No |
| 533924 | 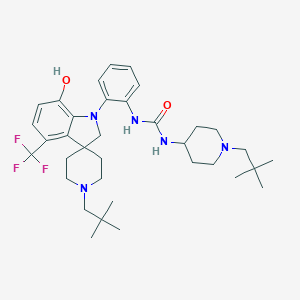 CHEMBL3930814 CHEMBL3930814 | C35H50F3N5O2 | 629.813 | 8 / 3 | 7.5 | No |
| 557406 | 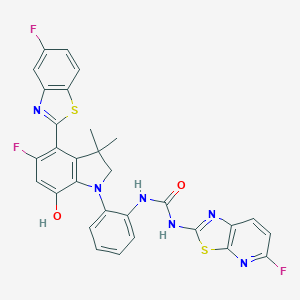 SCHEMBL17077170 SCHEMBL17077170 | C30H21F3N6O2S2 | 618.653 | 11 / 3 | 7.5 | No |
| 5499 | 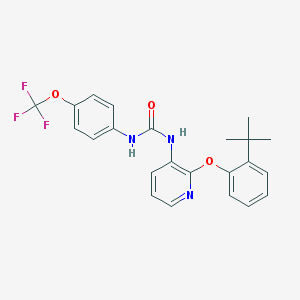 BPTU BPTU | C23H22F3N3O3 | 445.442 | 7 / 2 | 6.1 | No |
| 5634 | 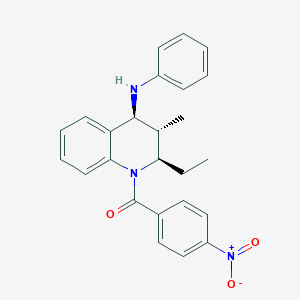 CHEMBL453637 CHEMBL453637 | C25H25N3O3 | 415.493 | 4 / 1 | 5.8 | No |
| 533932 | 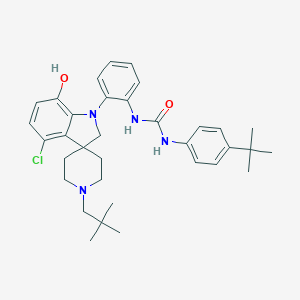 CHEMBL3946708 CHEMBL3946708 | C34H43ClN4O2 | 575.194 | 4 / 3 | 8.2 | No |
| 557476 | 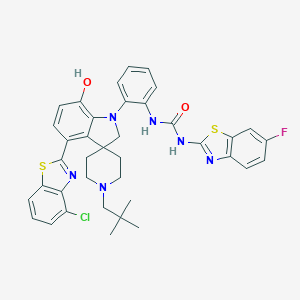 CHEMBL3892685 CHEMBL3892685 | C38H36ClFN6O2S2 | 727.314 | 9 / 3 | 9.5 | No |
| 6823 | 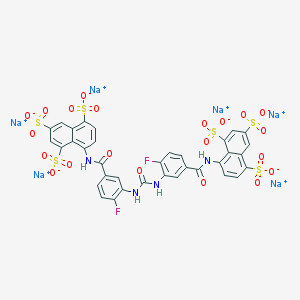 CHEMBL268784 CHEMBL268784 | C35H18F2N4Na6O21S6 | 1198.83 | 23 / 4 | N/A | No |
| 441978 | 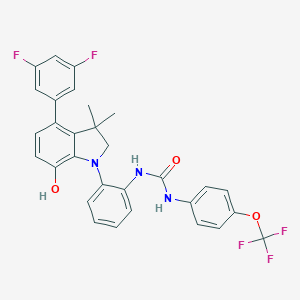 CHEMBL3314303 CHEMBL3314303 | C30H24F5N3O3 | 569.532 | 9 / 3 | 7.7 | No |
| 536158 | 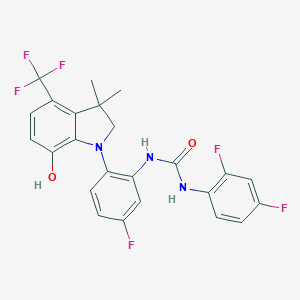 SCHEMBL15597032 SCHEMBL15597032 | C24H19F6N3O2 | 495.425 | 9 / 3 | 5.9 | No |
| 533937 | 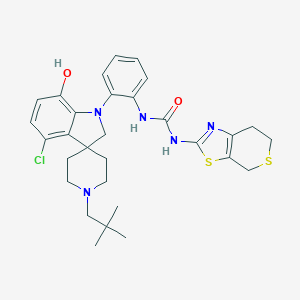 CHEMBL3954066 CHEMBL3954066 | C30H36ClN5O2S2 | 598.221 | 7 / 3 | 6.4 | No |
| 7447 | 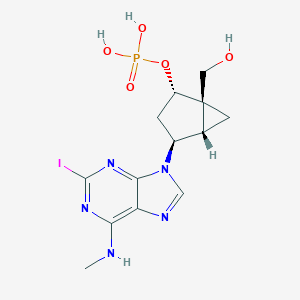 CHEMBL375022 CHEMBL375022 | C13H17IN5O5P | 481.187 | 9 / 4 | -0.7 | Yes |
| 7463 | 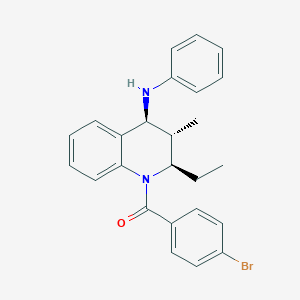 CHEMBL454910 CHEMBL454910 | C25H25BrN2O | 449.392 | 2 / 1 | 6.7 | No |
| 7531 | 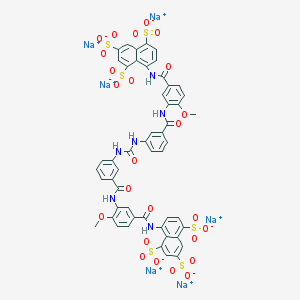 CHEMBL413910 CHEMBL413910 | C51H34N6Na6O25S6 | 1461.15 | 25 / 6 | N/A | No |
| 557511 | 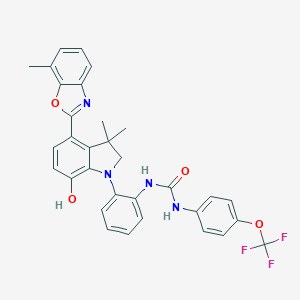 SCHEMBL15596905 SCHEMBL15596905 | C32H27F3N4O4 | 588.587 | 9 / 3 | 7.8 | No |
| 8302 | 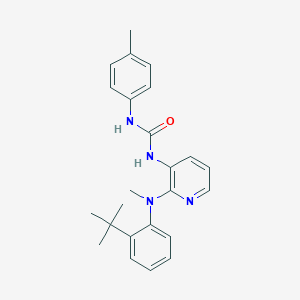 CHEMBL2333769 CHEMBL2333769 | C24H28N4O | 388.515 | 3 / 2 | 5.5 | No |
| 536189 | 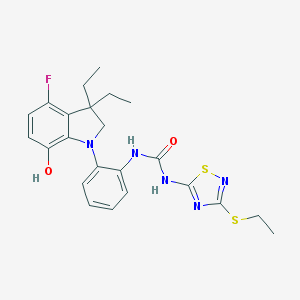 SCHEMBL17077212 SCHEMBL17077212 | C23H26FN5O2S2 | 487.612 | 8 / 3 | 5.9 | No |
| 8745 | 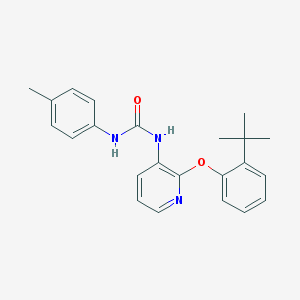 CHEMBL2333766 CHEMBL2333766 | C23H25N3O2 | 375.472 | 3 / 2 | 5.3 | No |
| 536236 | 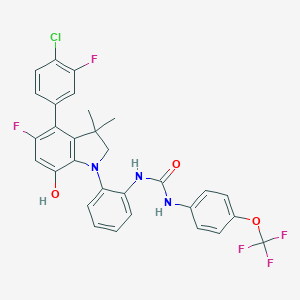 SCHEMBL15596991 SCHEMBL15596991 | C30H23ClF5N3O3 | 603.974 | 9 / 3 | 8.4 | No |
| 9745 | 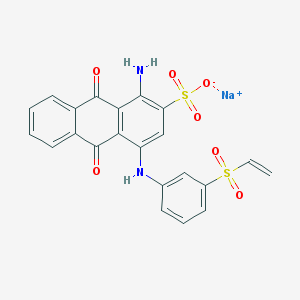 Uniblue A sodium salt Uniblue A sodium salt | C22H15N2NaO7S2 | 506.479 | 9 / 2 | N/A | No |
| 10007 | 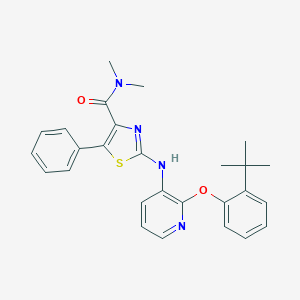 CHEMBL2401802 CHEMBL2401802 | C27H28N4O2S | 472.607 | 6 / 1 | 6.7 | No |
| 459313 | 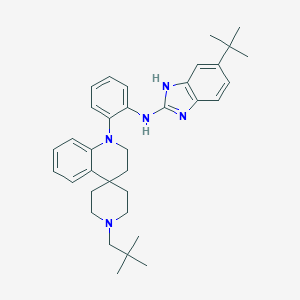 SCHEMBL1572821 SCHEMBL1572821 | C35H45N5 | 535.78 | 4 / 2 | 9.2 | No |
| 557592 | 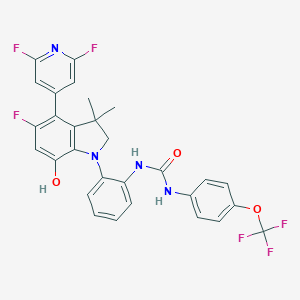 CHEMBL3314316 CHEMBL3314316 | C29H22F6N4O3 | 588.51 | 11 / 3 | 7.4 | No |
| 557594 | 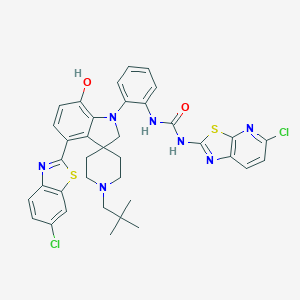 CHEMBL3906376 CHEMBL3906376 | C37H35Cl2N7O2S2 | 744.754 | 9 / 3 | 9.7 | No |
| 11228 | 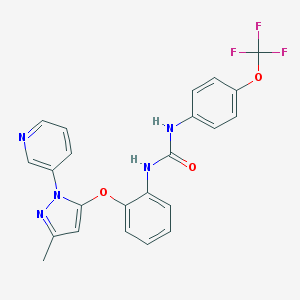 CHEMBL402793 CHEMBL402793 | C23H18F3N5O3 | 469.424 | 8 / 2 | 4.8 | Yes |
| 11250 | 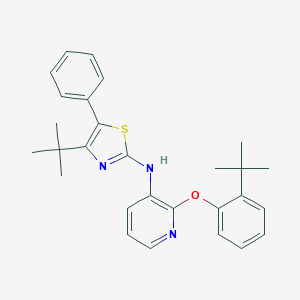 CHEMBL2401799 CHEMBL2401799 | C28H31N3OS | 457.636 | 5 / 1 | 8.7 | No |
| 11479 | 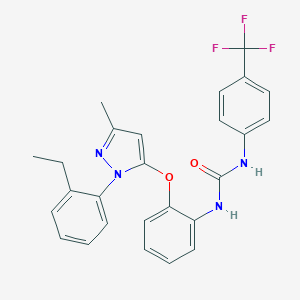 CHEMBL402837 CHEMBL402837 | C26H23F3N4O2 | 480.491 | 6 / 2 | 6.4 | No |
| 557624 | 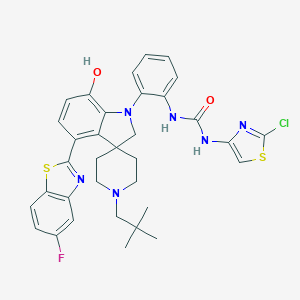 CHEMBL3924070 CHEMBL3924070 | C34H34ClFN6O2S2 | 677.254 | 9 / 3 | 8.4 | No |
| 557627 | 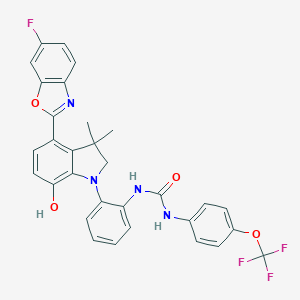 SCHEMBL15596961 SCHEMBL15596961 | C31H24F4N4O4 | 592.551 | 10 / 3 | 7.5 | No |
| 12090 | 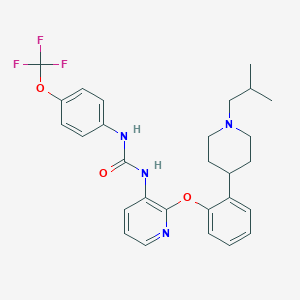 CHEMBL3092632 CHEMBL3092632 | C28H31F3N4O3 | 528.576 | 8 / 2 | 6.4 | No |
| 12537 | 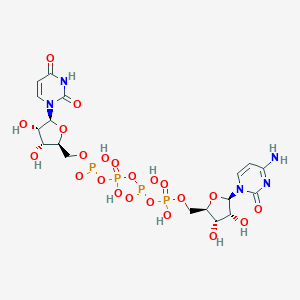 CHEMBL1162198 CHEMBL1162198 | C18H27N5O22P4 | 789.322 | 22 / 10 | -9.4 | No |
| 12587 | 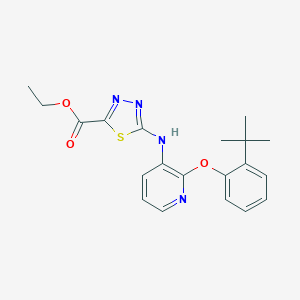 CHEMBL2393195 CHEMBL2393195 | C20H22N4O3S | 398.481 | 8 / 1 | 5.3 | No |
| 533967 | 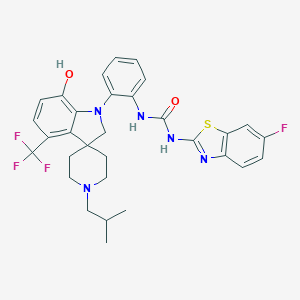 CHEMBL3925641 CHEMBL3925641 | C31H31F4N5O2S | 613.676 | 10 / 3 | 7.3 | No |
| 536326 | 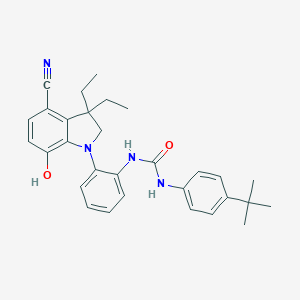 SCHEMBL15597029 SCHEMBL15597029 | C30H34N4O2 | 482.628 | 4 / 3 | 6.8 | No |
| 12993 | 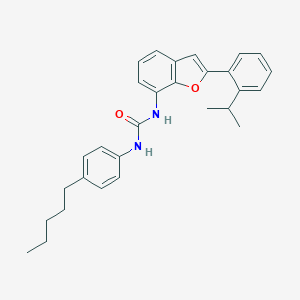 CHEMBL1169909 CHEMBL1169909 | C29H32N2O2 | 440.587 | 2 / 2 | 8.1 | No |
| 13196 | 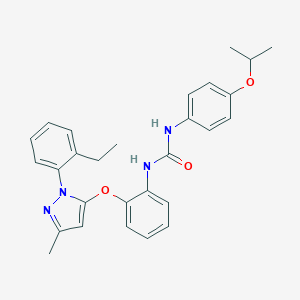 CHEMBL404869 CHEMBL404869 | C28H30N4O3 | 470.573 | 4 / 2 | 6.3 | No |
| 13285 | 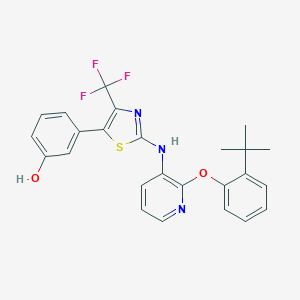 CHEMBL2401863 CHEMBL2401863 | C25H22F3N3O2S | 485.525 | 9 / 2 | 7.5 | No |
| 14457 | 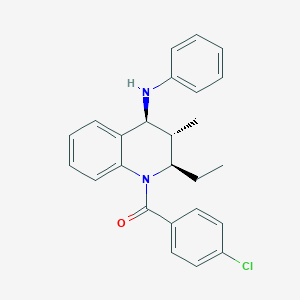 CHEMBL461532 CHEMBL461532 | C25H25ClN2O | 404.938 | 2 / 1 | 6.6 | No |
| 521883 | 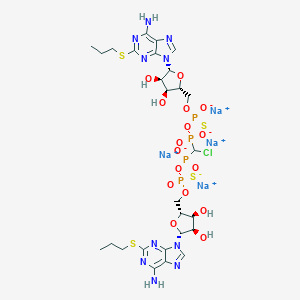 CHEMBL3745863 CHEMBL3745863 | C27H37ClN10Na4O16P4S4 | 1137.19 | 28 / 6 | N/A | No |
| 15351 | 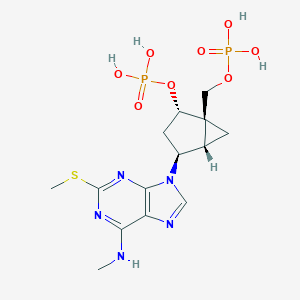 CHEMBL2112865 CHEMBL2112865 | C14H21N5O8P2S | 481.357 | 13 / 5 | -1.7 | No |
| 15621 | 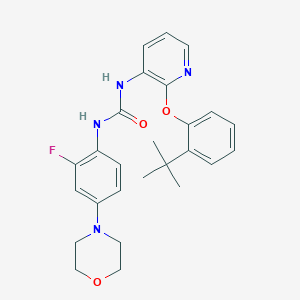 CHEMBL2381896 CHEMBL2381896 | C26H29FN4O3 | 464.541 | 6 / 2 | 4.8 | Yes |
| 16323 | 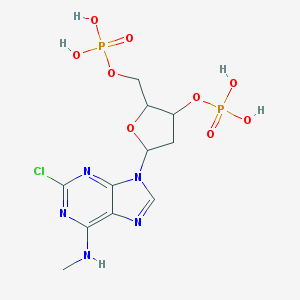 CHEMBL288798 CHEMBL288798 | C11H16ClN5O9P2 | 459.673 | 13 / 5 | -1.3 | No |
| 18267 | 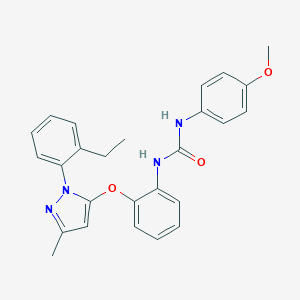 CHEMBL401657 CHEMBL401657 | C26H26N4O3 | 442.519 | 4 / 2 | 5.5 | No |
| 18309 | 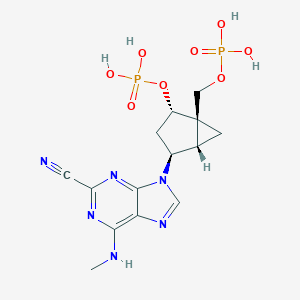 CHEMBL375682 CHEMBL375682 | C14H18N6O8P2 | 460.28 | 13 / 5 | -2.5 | No |
| 557837 | 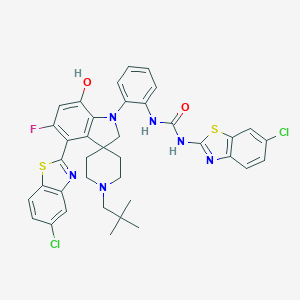 CHEMBL3961183 CHEMBL3961183 | C38H35Cl2FN6O2S2 | 761.756 | 9 / 3 | 10.2 | No |
| 522027 | 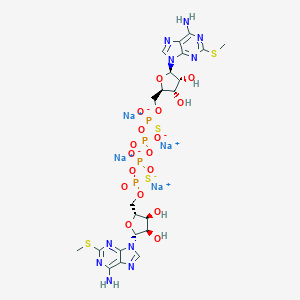 CHEMBL3746558 CHEMBL3746558 | C22H28N10Na4O17P4S4 | 1048.61 | 29 / 6 | N/A | No |
| 459391 | 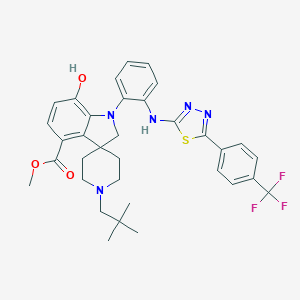 CHEMBL3926419 CHEMBL3926419 | C34H36F3N5O3S | 651.749 | 12 / 2 | 8.1 | No |
| 19867 | 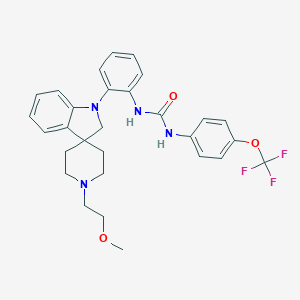 CHEMBL3104629 CHEMBL3104629 | C29H31F3N4O3 | 540.587 | 8 / 2 | 5.6 | No |
| 533987 | 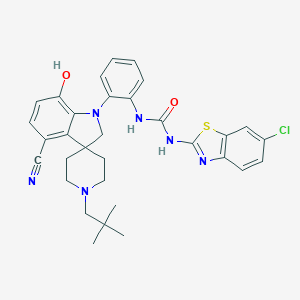 CHEMBL3964813 CHEMBL3964813 | C32H33ClN6O2S | 601.166 | 7 / 3 | 7.0 | No |
| 459395 | 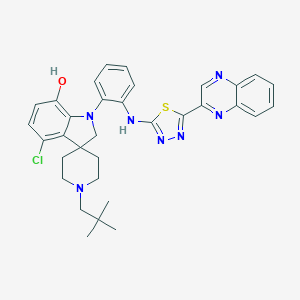 CHEMBL3972991 CHEMBL3972991 | C33H34ClN7OS | 612.193 | 9 / 2 | 7.3 | No |
| 557941 | 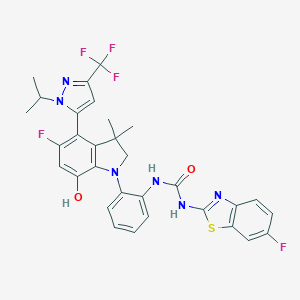 SCHEMBL15597008 SCHEMBL15597008 | C31H27F5N6O2S | 642.649 | 11 / 3 | 7.3 | No |
| 459404 | 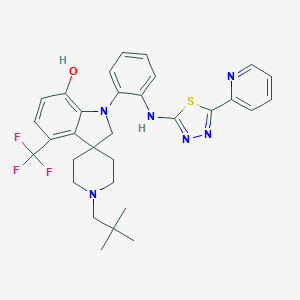 CHEMBL3900332 CHEMBL3900332 | C31H33F3N6OS | 594.701 | 11 / 2 | 7.2 | No |
| 442517 | 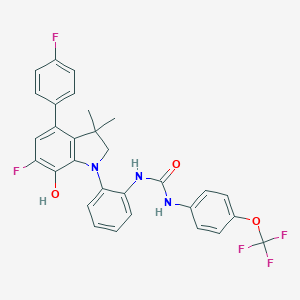 CHEMBL3314310 CHEMBL3314310 | C30H24F5N3O3 | 569.532 | 9 / 3 | 7.7 | No |
| 557957 | 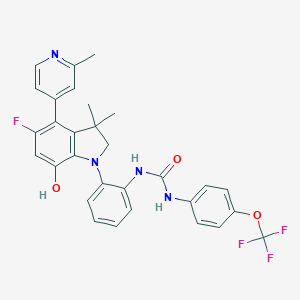 SCHEMBL17077215 SCHEMBL17077215 | C30H26F4N4O3 | 566.557 | 9 / 3 | 7.0 | No |
| 21947 | 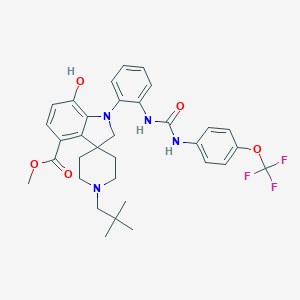 CHEMBL3287052 CHEMBL3287052 | C33H37F3N4O5 | 626.677 | 10 / 3 | 6.9 | No |
| 536557 | 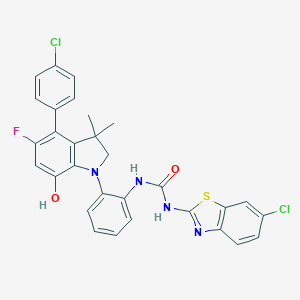 SCHEMBL15596945 SCHEMBL15596945 | C30H23Cl2FN4O2S | 593.498 | 6 / 3 | 8.5 | No |
| 536558 | 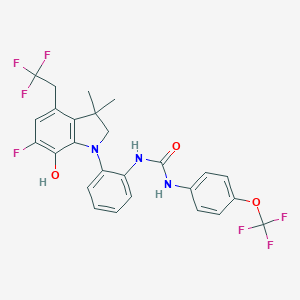 SCHEMBL15596944 SCHEMBL15596944 | C26H22F7N3O3 | 557.469 | 11 / 3 | 7.4 | No |
| 557974 | 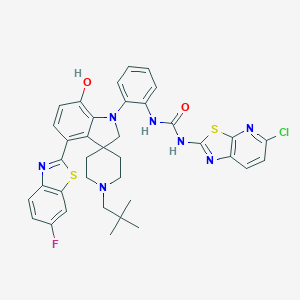 CHEMBL3951413 CHEMBL3951413 | C37H35ClFN7O2S2 | 728.302 | 10 / 3 | 9.1 | No |
| 533998 | 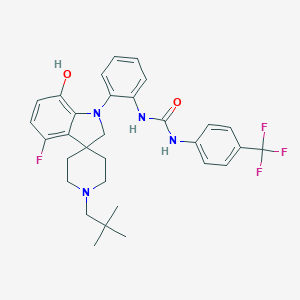 CHEMBL3930322 CHEMBL3930322 | C31H34F4N4O2 | 570.633 | 8 / 3 | 6.9 | No |
| 536569 | 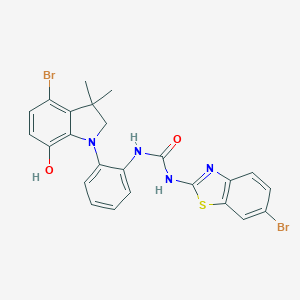 SCHEMBL15597099 SCHEMBL15597099 | C24H20Br2N4O2S | 588.318 | 5 / 3 | 6.9 | No |
| 23708 | 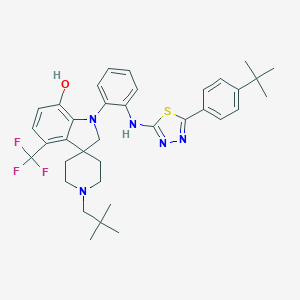 CHEMBL3263067 CHEMBL3263067 | C36H42F3N5OS | 649.821 | 10 / 2 | 9.9 | No |
| 23819 | 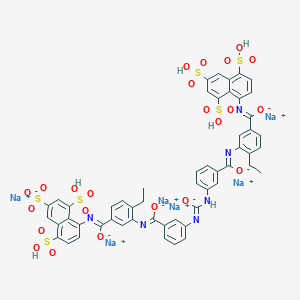 CID 136825090 CID 136825090 | C53H38N6Na6O23S6 | 1457.2 | 28 / 6 | N/A | No |
| 536595 | 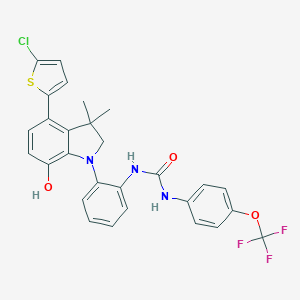 SCHEMBL15597151 SCHEMBL15597151 | C28H23ClF3N3O3S | 574.015 | 8 / 3 | 8.2 | No |
| 24785 | 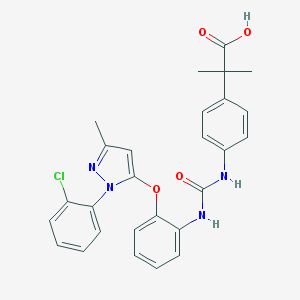 CHEMBL255454 CHEMBL255454 | C27H25ClN4O4 | 504.971 | 5 / 3 | 5.7 | No |
| 558048 | 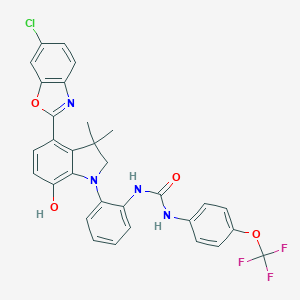 SCHEMBL15597039 SCHEMBL15597039 | C31H24ClF3N4O4 | 609.002 | 9 / 3 | 8.0 | No |
| 25281 | 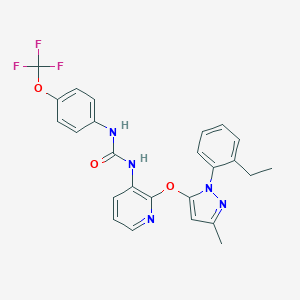 CHEMBL255307 CHEMBL255307 | C25H22F3N5O3 | 497.478 | 8 / 2 | 6.0 | No |
| 25645 | 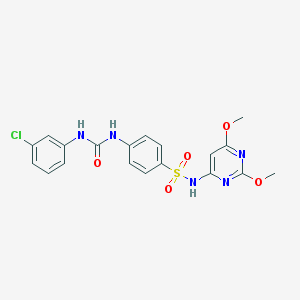 CHEMBL2071529 CHEMBL2071529 | C19H18ClN5O5S | 463.893 | 8 / 3 | 3.5 | Yes |
| 26655 | 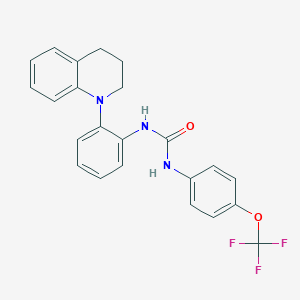 CHEMBL3103629 CHEMBL3103629 | C23H20F3N3O2 | 427.427 | 6 / 2 | 5.7 | No |
| 26863 | 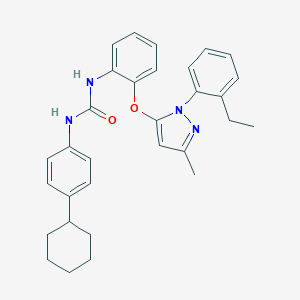 CHEMBL255724 CHEMBL255724 | C31H34N4O2 | 494.639 | 3 / 2 | 8.0 | No |
| 558101 | 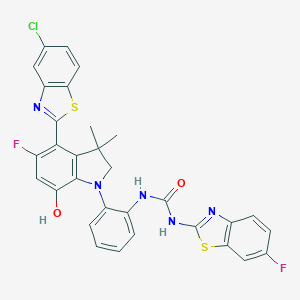 SCHEMBL17077171 SCHEMBL17077171 | C31H22ClF2N5O2S2 | 634.117 | 9 / 3 | 8.5 | No |
| 536697 | 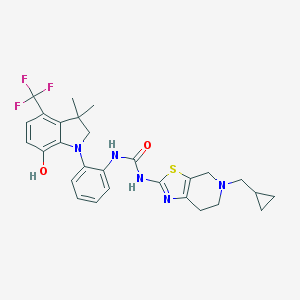 SCHEMBL17077132 SCHEMBL17077132 | C28H30F3N5O2S | 557.636 | 9 / 3 | 5.7 | No |
| 27899 | 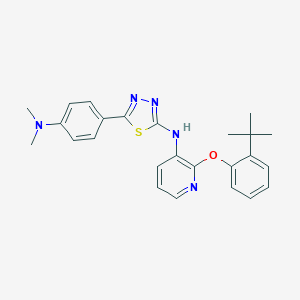 CHEMBL2393202 CHEMBL2393202 | C25H27N5OS | 445.585 | 7 / 1 | 6.5 | No |
| 558116 | 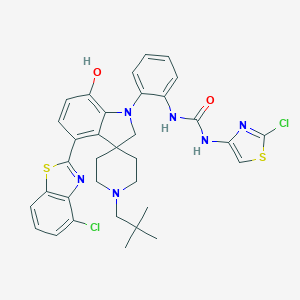 CHEMBL3981726 CHEMBL3981726 | C34H34Cl2N6O2S2 | 693.706 | 8 / 3 | 9.0 | No |
| 28814 | 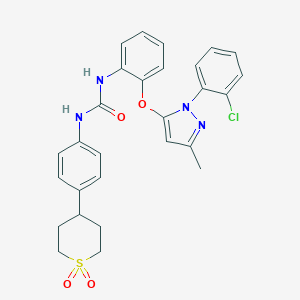 CHEMBL256944 CHEMBL256944 | C28H27ClN4O4S | 551.058 | 5 / 2 | 5.3 | No |
| 558153 | 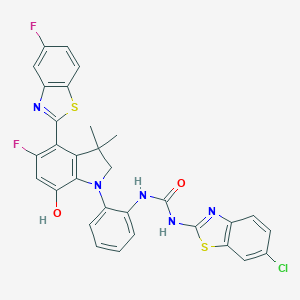 SCHEMBL17077140 SCHEMBL17077140 | C31H22ClF2N5O2S2 | 634.117 | 9 / 3 | 8.5 | No |
| 29164 | 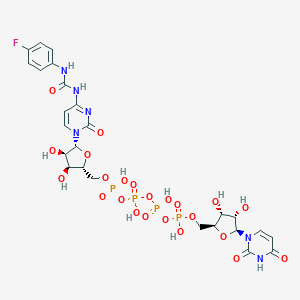 CHEMBL1162203 CHEMBL1162203 | C25H31FN6O23P4 | 926.435 | 24 / 11 | -7.8 | No |
| 534020 | 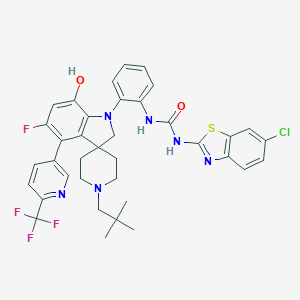 CHEMBL3894893 CHEMBL3894893 | C37H35ClF4N6O2S | 739.231 | 11 / 3 | 8.9 | No |
| 30401 | 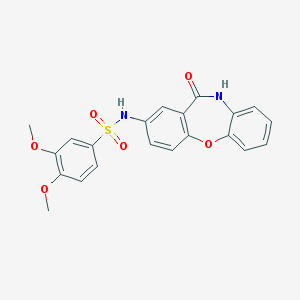 CHEMBL2071542 CHEMBL2071542 | C21H18N2O6S | 426.443 | 7 / 2 | 2.8 | Yes |
| 459478 | 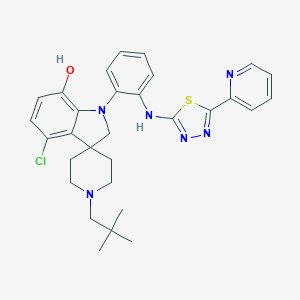 CHEMBL3932713 CHEMBL3932713 | C30H33ClN6OS | 561.145 | 8 / 2 | 6.9 | No |
| 30825 | 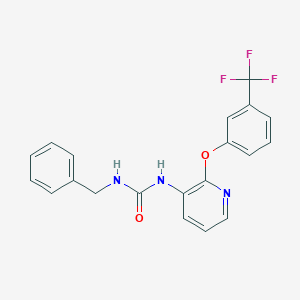 CHEMBL2333757 CHEMBL2333757 | C20H16F3N3O2 | 387.362 | 6 / 2 | 4.1 | Yes |
| 534027 | 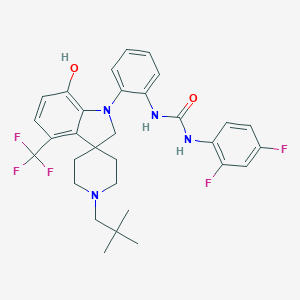 CHEMBL3944036 CHEMBL3944036 | C31H33F5N4O2 | 588.623 | 9 / 3 | 7.0 | No |
| 536775 | 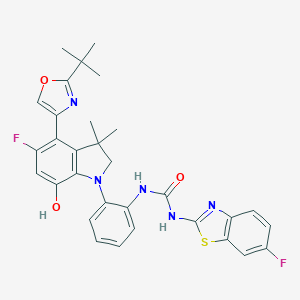 SCHEMBL15597046 SCHEMBL15597046 | C31H29F2N5O3S | 589.662 | 9 / 3 | 7.5 | No |
| 534029 | 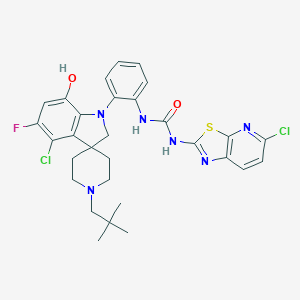 CHEMBL3955518 CHEMBL3955518 | C30H31Cl2FN6O2S | 629.576 | 8 / 3 | 7.6 | No |
| 558239 | 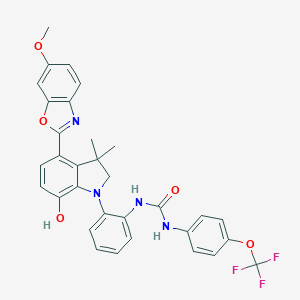 SCHEMBL15597131 SCHEMBL15597131 | C32H27F3N4O5 | 604.586 | 10 / 3 | 7.4 | No |
| 33949 | 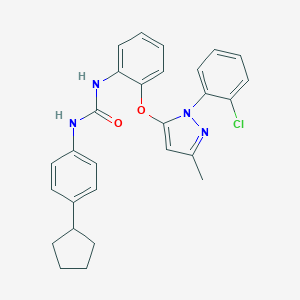 CHEMBL256088 CHEMBL256088 | C28H27ClN4O2 | 487.0 | 3 / 2 | 7.3 | No |
| 536868 | 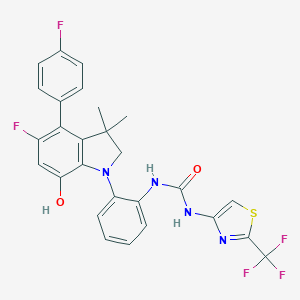 SCHEMBL15597162 SCHEMBL15597162 | C27H21F5N4O2S | 560.543 | 10 / 3 | 6.8 | No |
| 536894 | 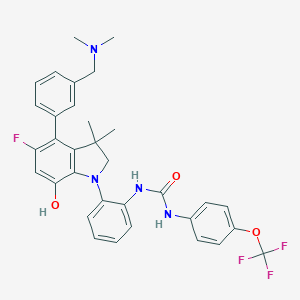 SCHEMBL15596998 SCHEMBL15596998 | C33H32F4N4O3 | 608.638 | 9 / 3 | 7.5 | No |
| 36208 | 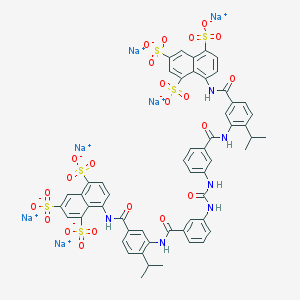 CHEMBL405500 CHEMBL405500 | C55H42N6Na6O23S6 | 1485.26 | 23 / 6 | N/A | No |
| 36505 | 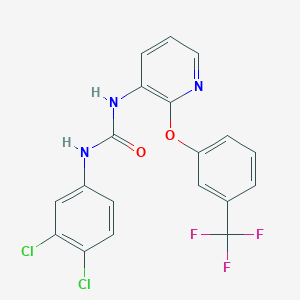 CHEMBL2333361 CHEMBL2333361 | C19H12Cl2F3N3O2 | 442.219 | 6 / 2 | 5.4 | No |
| 558407 | 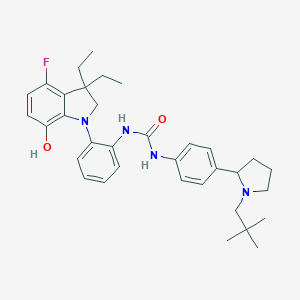 SCHEMBL15596915 SCHEMBL15596915 | C34H43FN4O2 | 558.742 | 5 / 3 | 7.6 | No |
| 37981 | 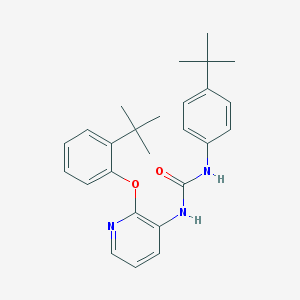 CHEMBL2333771 CHEMBL2333771 | C26H31N3O2 | 417.553 | 3 / 2 | 6.6 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218