

You can:
| Name | Melanocyte-stimulating hormone receptor |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | MC1R |
| Synonym | MSH-R Melanocortin receptor 1 melanocortin 1 receptor (alpha melanocyte stimulating hormone receptor) MC1-R MC1 receptor |
| Disease | Atopic dermatitis |
| Length | 317 |
| Amino acid sequence | MAVQGSQRRLLGSLNSTPTAIPQLGLAANQTGARCLEVSISDGLFLSLGLVSLVENALVVATIAKNRNLHSPMYCFICCLALSDLLVSGSNVLETAVILLLEAGALVARAAVLQQLDNVIDVITCSSMLSSLCFLGAIAVDRYISIFYALRYHSIVTLPRARRAVAAIWVASVVFSTLFIAYYDHVAVLLCLVVFFLAMLVLMAVLYVHMLARACQHAQGIARLHKRQRPVHQGFGLKGAVTLTILLGIFFLCWGPFFLHLTLIVLCPEHPTCGCIFKNFNLFLALIICNAIIDPLIYAFHSQELRRTLKEVLTCSW |
| UniProt | Q01726 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | Q01726 |
| 3D structure model | This predicted structure model is from GPCR-EXP Q01726. |
| BioLiP | N/A |
| Therapeutic Target Database | T35842 |
| ChEMBL | CHEMBL3795 |
| IUPHAR | 282 |
| DrugBank | BE0002447 |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 194 | 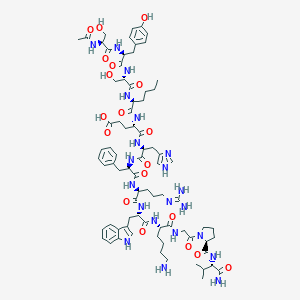 CHEMBL2372838 CHEMBL2372838 | C78H111N21O19 | 1646.87 | 22 / 22 | -3.9 | No |
| 1366 | 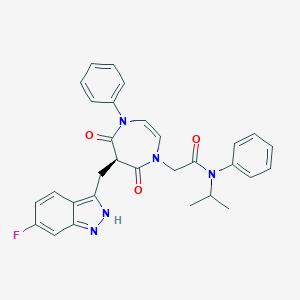 CHEMBL596804 CHEMBL596804 | C30H28FN5O3 | 525.584 | 5 / 1 | 4.3 | No |
| 1369 | 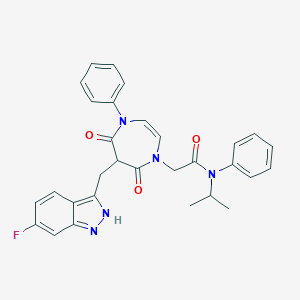 CHEMBL598021 CHEMBL598021 | C30H28FN5O3 | 525.584 | 5 / 1 | 4.3 | No |
| 1370 | 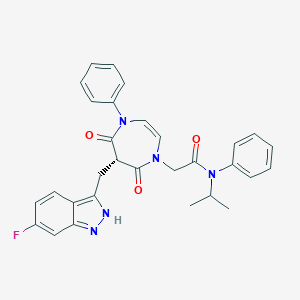 CHEMBL596602 CHEMBL596602 | C30H28FN5O3 | 525.584 | 5 / 1 | 4.3 | No |
| 1900 | 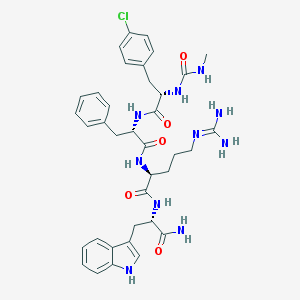 CHEMBL181161 CHEMBL181161 | C37H45ClN10O5 | 745.282 | 6 / 9 | 2.0 | No |
| 3305 | 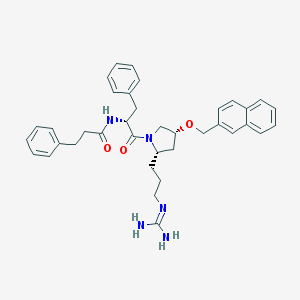 CHEMBL212976 CHEMBL212976 | C37H43N5O3 | 605.783 | 4 / 3 | 5.0 | No |
| 3993 | 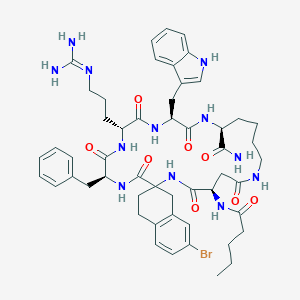 CHEMBL412174 CHEMBL412174 | C52H67BrN12O8 | 1068.09 | 9 / 11 | 3.0 | No |
| 441864 | 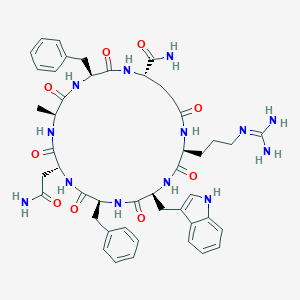 CHEMBL1775057 CHEMBL1775057 | C47H59N13O9 | 950.071 | 10 / 12 | -0.8 | No |
| 441870 | 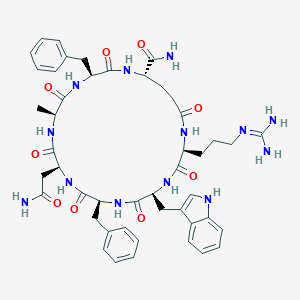 CHEMBL1775063 CHEMBL1775063 | C47H59N13O9 | 950.071 | 10 / 12 | -0.8 | No |
| 441871 | 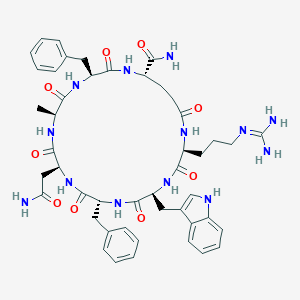 CHEMBL1775064 CHEMBL1775064 | C47H59N13O9 | 950.071 | 10 / 12 | -0.8 | No |
| 4344 | 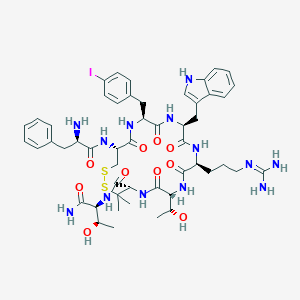 CHEMBL384813 CHEMBL384813 | C51H68IN13O10S2 | 1214.21 | 14 / 14 | 0.9 | No |
| 4709 | 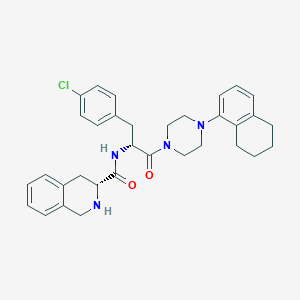 CHEMBL2113144 CHEMBL2113144 | C33H37ClN4O2 | 557.135 | 4 / 2 | 5.8 | No |
| 5648 | 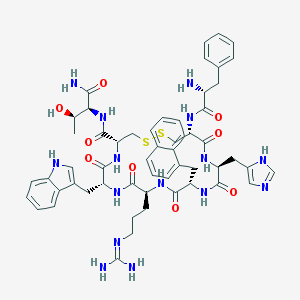 CHEMBL405408 CHEMBL405408 | C55H67N15O9S2 | 1146.36 | 14 / 14 | 0.7 | No |
| 5981 | 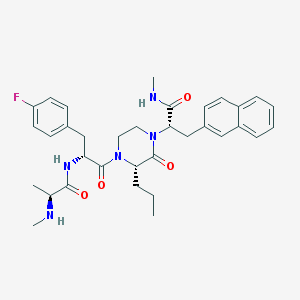 CHEMBL504349 CHEMBL504349 | C34H42FN5O4 | 603.739 | 6 / 3 | 4.3 | No |
| 6072 | 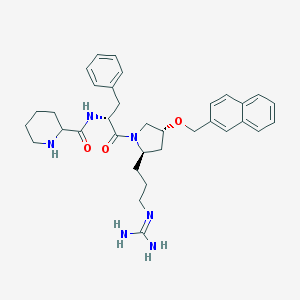 CHEMBL213752 CHEMBL213752 | C34H44N6O3 | 584.765 | 5 / 4 | 3.6 | No |
| 6074 | 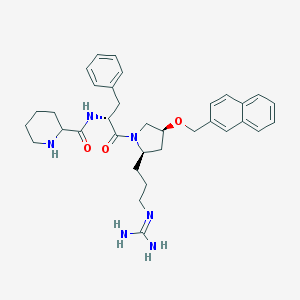 CHEMBL210009 CHEMBL210009 | C34H44N6O3 | 584.765 | 5 / 4 | 3.6 | No |
| 6077 | 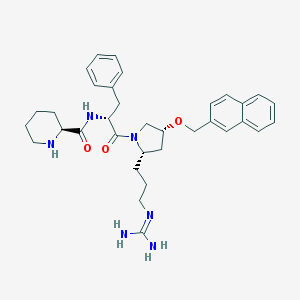 CHEMBL213956 CHEMBL213956 | C34H44N6O3 | 584.765 | 5 / 4 | 3.6 | No |
| 6079 | 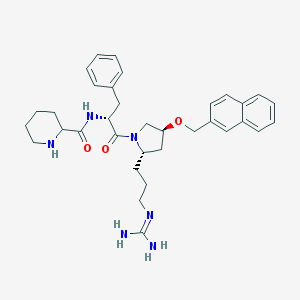 CHEMBL210011 CHEMBL210011 | C34H44N6O3 | 584.765 | 5 / 4 | 3.6 | No |
| 6469 | 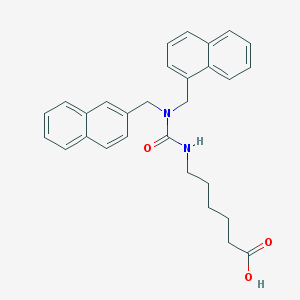 CHEMBL8825 CHEMBL8825 | C29H30N2O3 | 454.57 | 3 / 2 | 5.6 | No |
| 6520 | 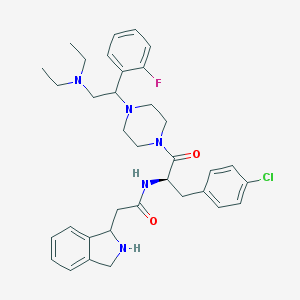 CHEMBL380855 CHEMBL380855 | C35H43ClFN5O2 | 620.21 | 6 / 2 | 4.5 | No |
| 6555 | 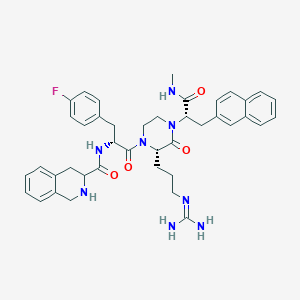 CHEMBL215895 CHEMBL215895 | C41H47FN8O4 | 734.877 | 7 / 5 | 3.4 | No |
| 6674 | 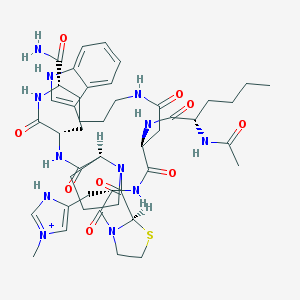 CHEMBL267360 CHEMBL267360 | C45H63N12O9S+ | 948.134 | 10 / 9 | 0.4 | No |
| 7141 | 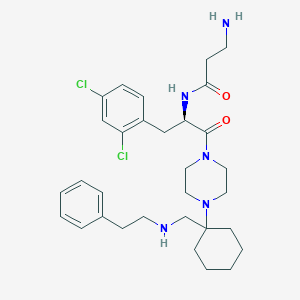 CHEMBL365237 CHEMBL365237 | C31H43Cl2N5O2 | 588.618 | 5 / 3 | 4.3 | No |
| 7587 | 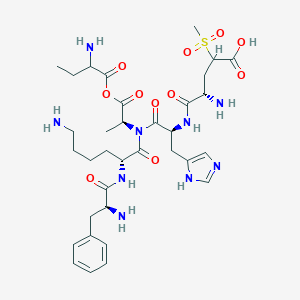 BDBM82410 BDBM82410 | C34H51N9O11S | 793.894 | 16 / 8 | -4.1 | No |
| 7709 | 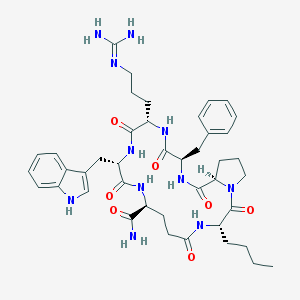 CHEMBL381739 CHEMBL381739 | C42H57N11O7 | 827.988 | 8 / 9 | 1.2 | No |
| 9537 | 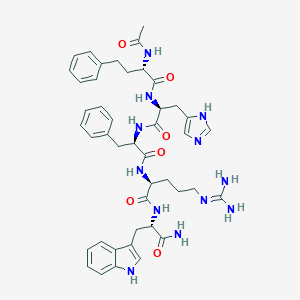 CHEMBL1923663 CHEMBL1923663 | C44H54N12O6 | 846.994 | 8 / 10 | 1.7 | No |
| 517370 | 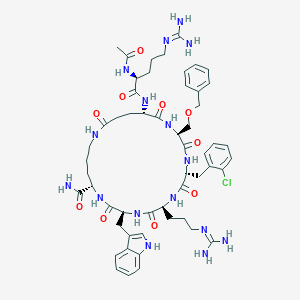 SCHEMBL8369982 SCHEMBL8369982 | C54H73ClN16O10 | 1141.73 | 12 / 14 | -0.8 | No |
| 10114 | 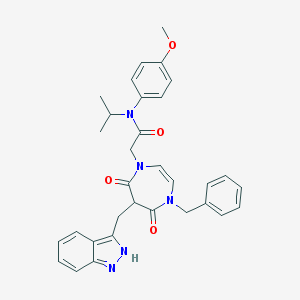 CHEMBL591026 CHEMBL591026 | C32H33N5O4 | 551.647 | 5 / 1 | 4.1 | No |
| 10141 | 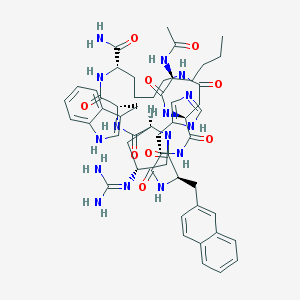 CHEMBL508501 CHEMBL508501 | C54H69N15O9 | 1072.24 | 11 / 12 | 0.9 | No |
| 10146 | 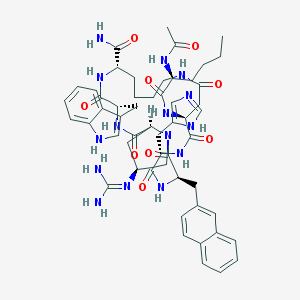 CHEMBL509582 CHEMBL509582 | C54H69N15O9 | 1072.24 | 11 / 12 | 0.9 | No |
| 10374 | 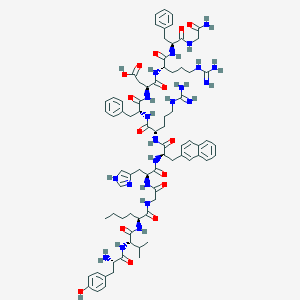 CID 16151595 CID 16151595 | C77H103N21O15 | 1562.8 | 19 / 22 | -0.3 | No |
| 10676 | 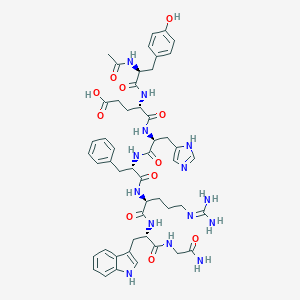 CHEMBL2028953 CHEMBL2028953 | C50H62N14O11 | 1035.13 | 13 / 14 | -0.4 | No |
| 12402 | 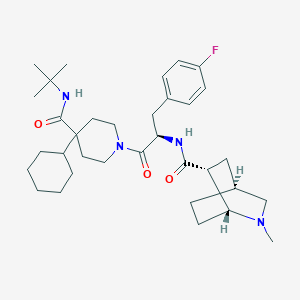 RY-764 RY-764 | C34H51FN4O3 | 582.805 | 5 / 2 | 5.4 | No |
| 13405 | 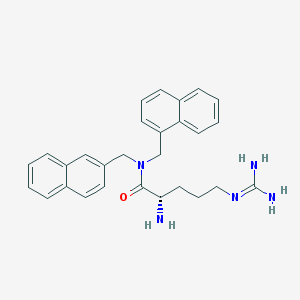 CHEMBL267020 CHEMBL267020 | C28H31N5O | 453.59 | 3 / 3 | 3.5 | Yes |
| 13527 | 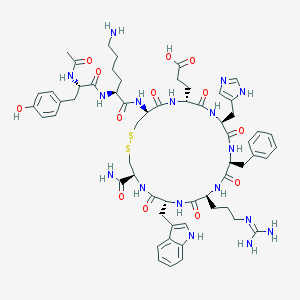 Ac-YK[CEHdFRWC]-NH2 Ac-YK[CEHdFRWC]-NH2 | C60H79N17O13S2 | 1310.52 | 18 / 17 | -3.3 | No |
| 14734 | 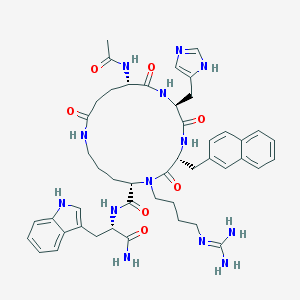 CHEMBL385561 CHEMBL385561 | C47H59N13O7 | 918.073 | 9 / 10 | 0.9 | No |
| 14869 | 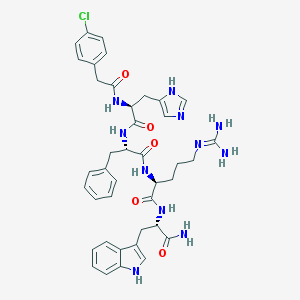 CHEMBL92481 CHEMBL92481 | C40H46ClN11O5 | 796.33 | 7 / 9 | 2.2 | No |
| 15529 | 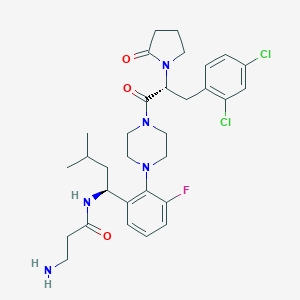 CHEMBL397580 CHEMBL397580 | C31H40Cl2FN5O3 | 620.591 | 6 / 2 | 4.1 | No |
| 15896 | 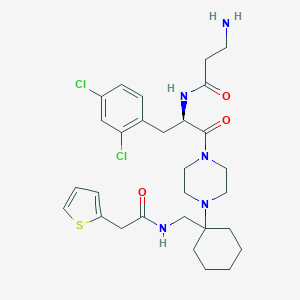 CHEMBL188651 CHEMBL188651 | C29H39Cl2N5O3S | 608.623 | 6 / 3 | 3.4 | No |
| 548060 | 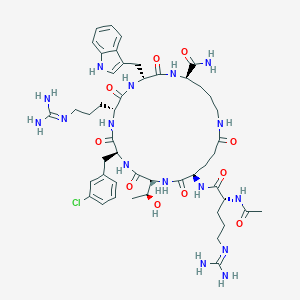 CHEMBL3971054 CHEMBL3971054 | C48H69ClN16O10 | 1065.63 | 12 / 15 | -1.8 | No |
| 557728 | 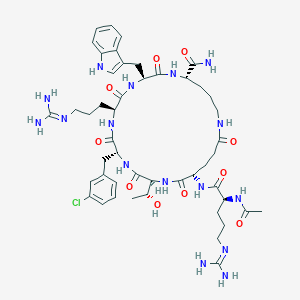 SCHEMBL8371033 SCHEMBL8371033 | C48H69ClN16O10 | 1065.63 | 12 / 15 | -1.8 | No |
| 464871 | 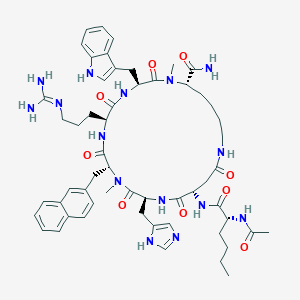 CHEMBL3600838 CHEMBL3600838 | C56H75N15O9 | 1102.31 | 11 / 11 | 1.6 | No |
| 517406 | 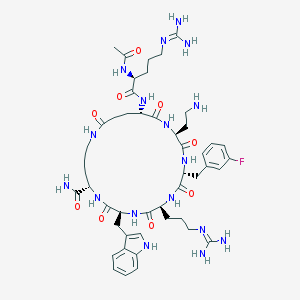 SCHEMBL8371464 SCHEMBL8371464 | C48H70FN17O9 | 1048.2 | 13 / 15 | -3.2 | No |
| 536408 | 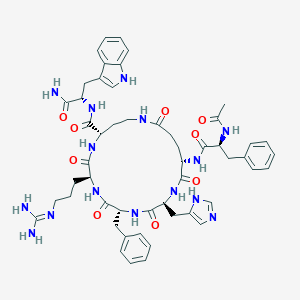 CHEMBL3898758 CHEMBL3898758 | C52H65N15O9 | 1044.19 | 11 / 13 | -0.2 | No |
| 536411 | 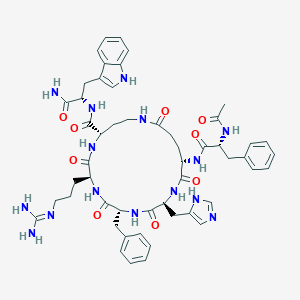 CHEMBL3936714 CHEMBL3936714 | C52H65N15O9 | 1044.19 | 11 / 13 | -0.2 | No |
| 536426 | 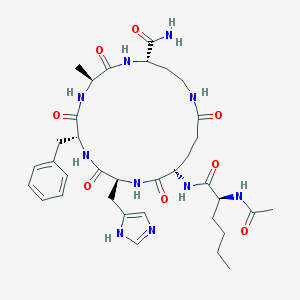 CHEMBL3965807 CHEMBL3965807 | C35H50N10O8 | 738.847 | 9 / 9 | -0.6 | No |
| 17525 | 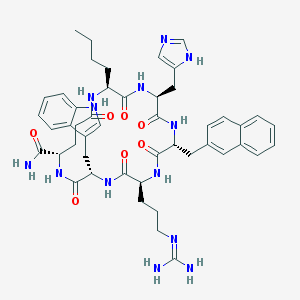 CHEMBL425591 CHEMBL425591 | C47H59N13O7 | 918.073 | 9 / 11 | 1.9 | No |
| 18722 | 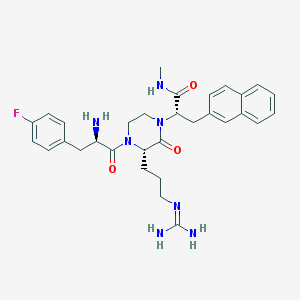 CHEMBL378837 CHEMBL378837 | C31H38FN7O3 | 575.689 | 6 / 4 | 2.0 | No |
| 465269 | 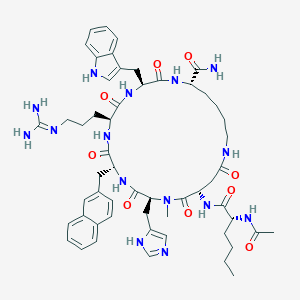 CHEMBL3600835 CHEMBL3600835 | C55H73N15O9 | 1088.29 | 11 / 12 | 1.5 | No |
| 19967 | 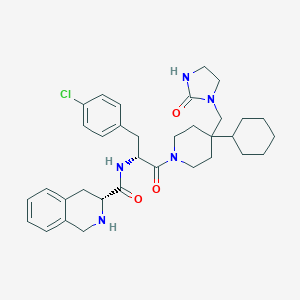 CHEMBL381503 CHEMBL381503 | C34H44ClN5O3 | 606.208 | 4 / 3 | 5.3 | No |
| 536496 | 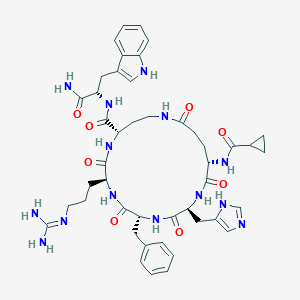 CHEMBL3911582 CHEMBL3911582 | C45H58N14O8 | 923.049 | 10 / 12 | -1.0 | No |
| 21528 | 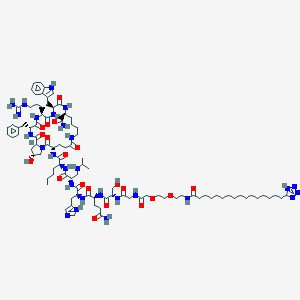 CID 70693082 CID 70693082 | C93H145N27O20 | 1961.35 | 26 / 24 | 1.5 | No |
| 517439 | 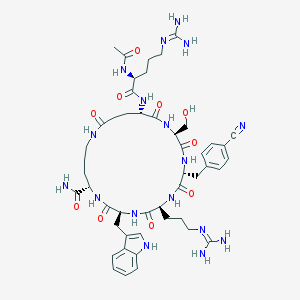 CHEMBL3663361 CHEMBL3663361 | C48H67N17O10 | 1042.17 | 13 / 15 | -3.1 | No |
| 517447 | 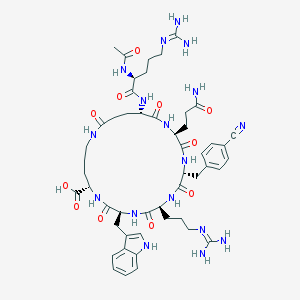 CHEMBL3667932 CHEMBL3667932 | C50H69N17O11 | 1084.21 | 14 / 15 | -3.2 | No |
| 23003 | 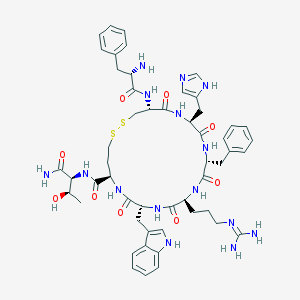 CHEMBL406891 CHEMBL406891 | C52H67N15O9S2 | 1110.32 | 14 / 14 | -0.2 | No |
| 23191 | 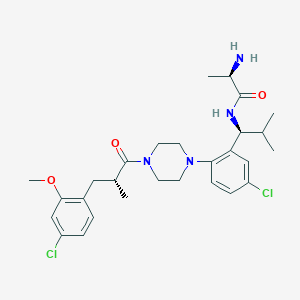 CHEMBL467587 CHEMBL467587 | C28H38Cl2N4O3 | 549.537 | 5 / 2 | 5.0 | No |
| 23308 | 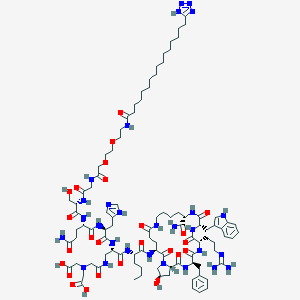 BDBM50389782 BDBM50389782 | C96H146N28O25 | 2092.4 | 31 / 25 | -3.0 | No |
| 517451 | 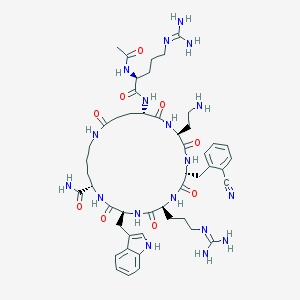 SCHEMBL8369995 SCHEMBL8369995 | C49H70N18O9 | 1055.22 | 13 / 15 | -3.6 | No |
| 23785 | 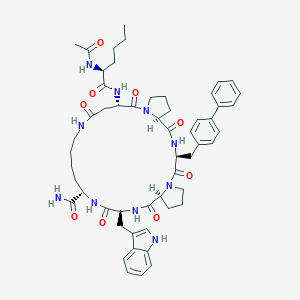 CHEMBL2371963 CHEMBL2371963 | C54H68N10O9 | 1001.2 | 9 / 8 | 3.7 | No |
| 24581 | 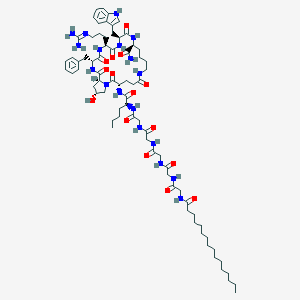 CHEMBL2070248 CHEMBL2070248 | C74H114N18O15 | 1495.84 | 16 / 17 | 4.0 | No |
| 24966 | 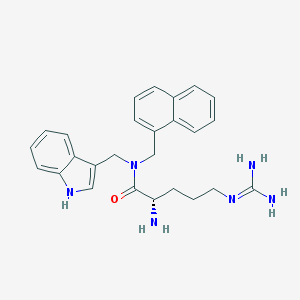 CHEMBL2113280 CHEMBL2113280 | C26H30N6O | 442.567 | 3 / 4 | 2.4 | Yes |
| 25682 | 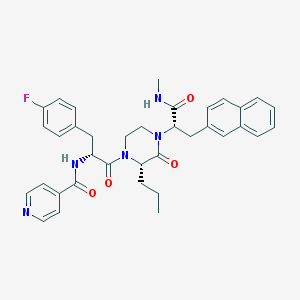 CHEMBL449131 CHEMBL449131 | C36H38FN5O4 | 623.729 | 6 / 2 | 4.9 | No |
| 466225 | 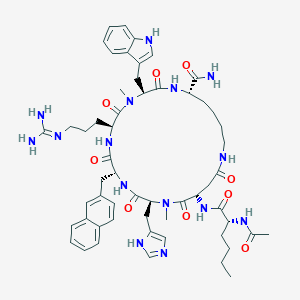 CHEMBL3600842 CHEMBL3600842 | C56H75N15O9 | 1102.31 | 11 / 11 | 1.6 | No |
| 27258 | 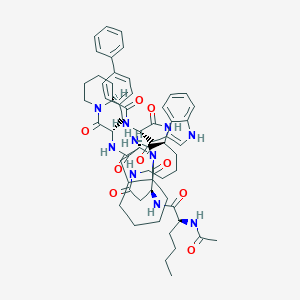 CHEMBL2371968 CHEMBL2371968 | C59H76N10O9 | 1069.32 | 9 / 8 | 5.4 | No |
| 28030 | 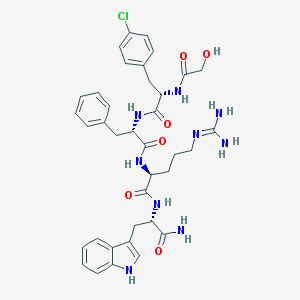 CHEMBL366040 CHEMBL366040 | C37H44ClN9O6 | 746.266 | 7 / 9 | 1.5 | No |
| 517461 | 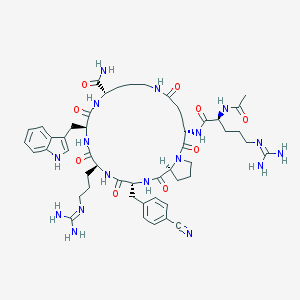 SCHEMBL8368801 SCHEMBL8368801 | C50H69N17O9 | 1052.21 | 12 / 13 | -2.3 | No |
| 517465 | 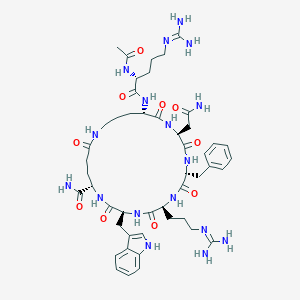 CHEMBL3646886 CHEMBL3646886 | C48H69N17O10 | 1044.19 | 12 / 15 | -3.9 | No |
| 517466 | 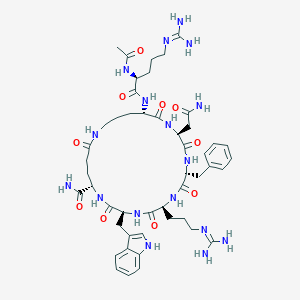 US8846601, 202 US8846601, 202 | C48H69N17O10 | 1044.19 | 12 / 15 | -3.9 | No |
| 29093 | 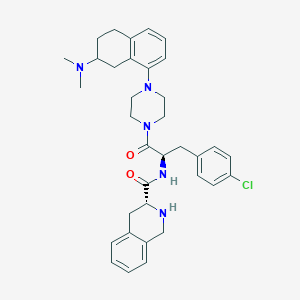 CHEMBL2113149 CHEMBL2113149 | C35H42ClN5O2 | 600.204 | 5 / 2 | 5.2 | No |
| 548212 | 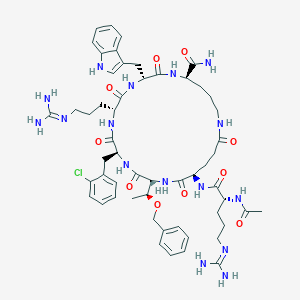 CHEMBL3929099 CHEMBL3929099 | C55H75ClN16O10 | 1155.76 | 12 / 14 | -0.3 | No |
| 558181 | 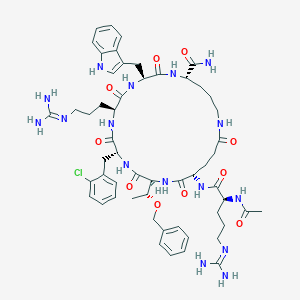 SCHEMBL8369075 SCHEMBL8369075 | C55H75ClN16O10 | 1155.76 | 12 / 14 | -0.3 | No |
| 30250 | 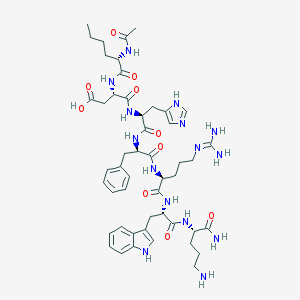 CHEMBL2372830 CHEMBL2372830 | C49H69N15O10 | 1028.19 | 13 / 14 | -3.1 | No |
| 33350 | 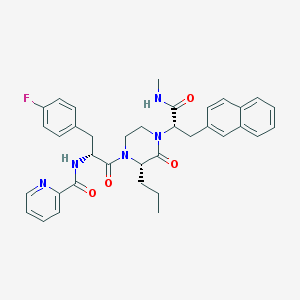 CHEMBL448337 CHEMBL448337 | C36H38FN5O4 | 623.729 | 6 / 2 | 5.3 | No |
| 33384 | 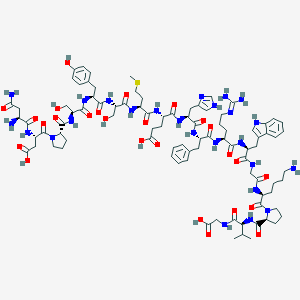 D0I8SJ D0I8SJ | C90H127N25O26S | 2007.21 | 31 / 27 | -10.1 | No |
| 442973 | 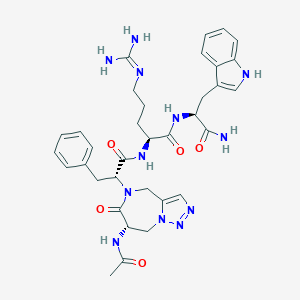 CHEMBL3421680 CHEMBL3421680 | C34H42N12O5 | 698.789 | 8 / 7 | -0.9 | No |
| 33997 | 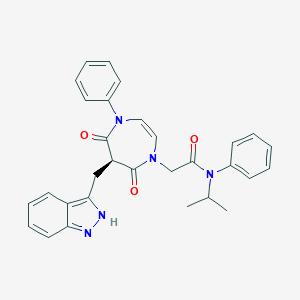 CHEMBL598643 CHEMBL598643 | C30H29N5O3 | 507.594 | 4 / 1 | 4.2 | No |
| 33998 | 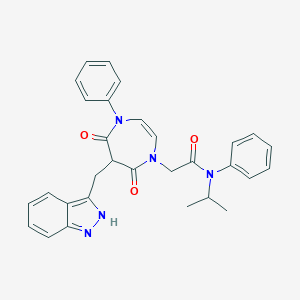 CHEMBL603468 CHEMBL603468 | C30H29N5O3 | 507.594 | 4 / 1 | 4.2 | No |
| 34001 | 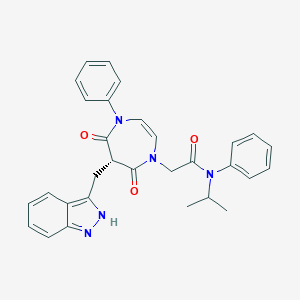 CHEMBL598442 CHEMBL598442 | C30H29N5O3 | 507.594 | 4 / 1 | 4.2 | No |
| 34057 | 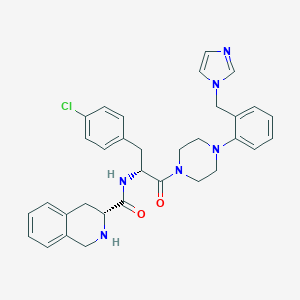 CHEMBL423101 CHEMBL423101 | C33H35ClN6O2 | 583.133 | 5 / 2 | 4.1 | No |
| 35070 | 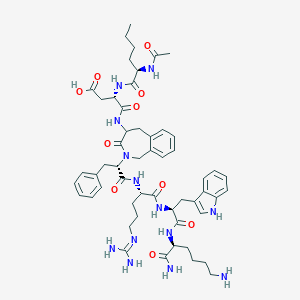 CHEMBL234456 CHEMBL234456 | C54H73N13O10 | 1064.26 | 12 / 12 | -1.1 | No |
| 36195 | 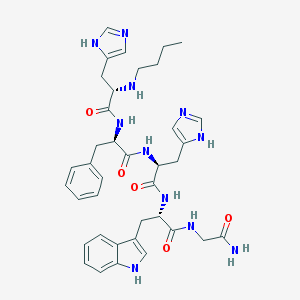 CHEMBL309213 CHEMBL309213 | C38H47N11O5 | 737.866 | 8 / 9 | 1.5 | No |
| 36354 | 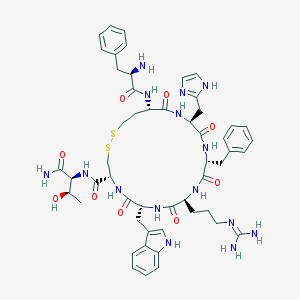 CHEMBL407098 CHEMBL407098 | C52H67N15O9S2 | 1110.32 | 14 / 14 | -0.2 | No |
| 36374 | 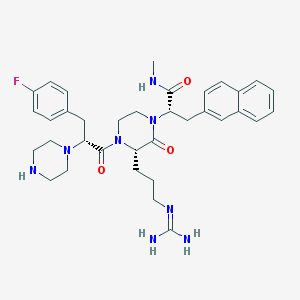 CHEMBL214410 CHEMBL214410 | C35H45FN8O3 | 644.796 | 7 / 4 | 2.3 | No |
| 37422 | 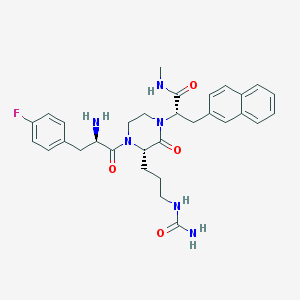 CHEMBL213284 CHEMBL213284 | C31H37FN6O4 | 576.673 | 6 / 4 | 2.3 | No |
| 536940 | 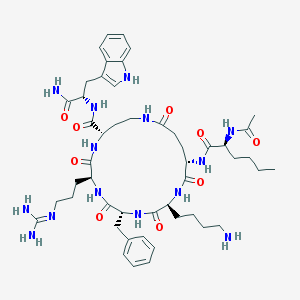 CHEMBL3955172 CHEMBL3955172 | C49H72N14O9 | 1001.2 | 11 / 13 | -0.3 | No |
| 536945 | 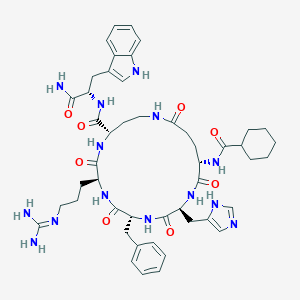 CHEMBL3981581 CHEMBL3981581 | C48H64N14O8 | 965.13 | 10 / 12 | 0.6 | No |
| 38379 | 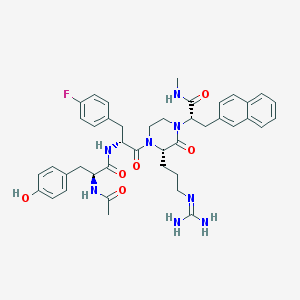 CHEMBL385000 CHEMBL385000 | C42H49FN8O6 | 780.902 | 8 / 6 | 3.2 | No |
| 39091 | 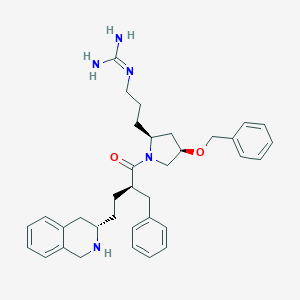 CHEMBL380437 CHEMBL380437 | C35H45N5O2 | 567.778 | 4 / 3 | 4.3 | No |
| 39761 | 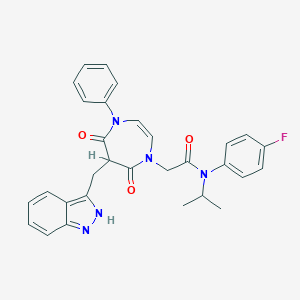 CHEMBL598616 CHEMBL598616 | C30H28FN5O3 | 525.584 | 5 / 1 | 4.3 | No |
| 39886 | 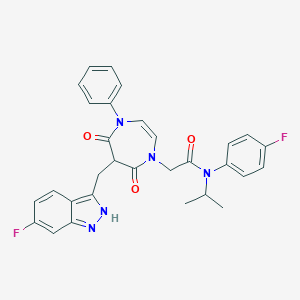 CHEMBL598617 CHEMBL598617 | C30H27F2N5O3 | 543.575 | 6 / 1 | 4.4 | No |
| 40255 | 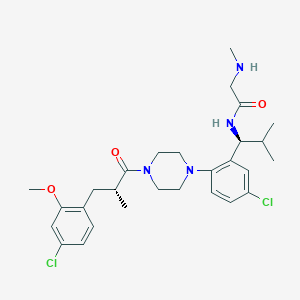 CHEMBL467586 CHEMBL467586 | C28H38Cl2N4O3 | 549.537 | 5 / 2 | 5.1 | No |
| 41140 | 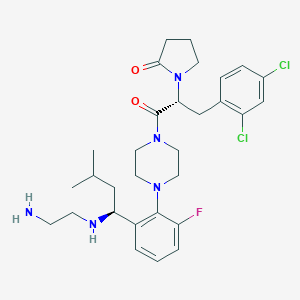 CHEMBL393576 CHEMBL393576 | C30H40Cl2FN5O2 | 592.581 | 6 / 2 | 4.4 | No |
| 41825 | 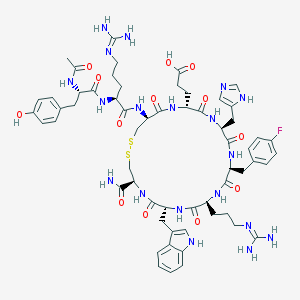 CHEMBL407809 CHEMBL407809 | C60H78FN19O13S2 | 1356.52 | 19 / 18 | -1.9 | No |
| 42057 | 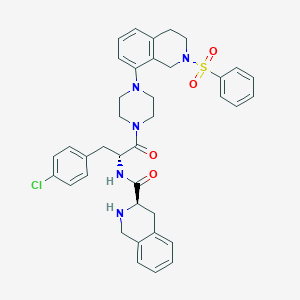 CHEMBL2113139 CHEMBL2113139 | C38H40ClN5O4S | 698.279 | 7 / 2 | 5.1 | No |
| 42154 | 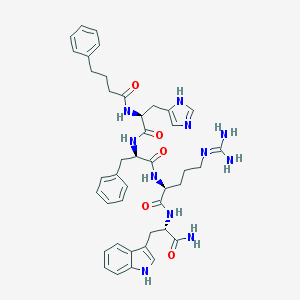 CHEMBL1923662 CHEMBL1923662 | C42H51N11O5 | 789.942 | 7 / 9 | 2.2 | No |
| 42158 | 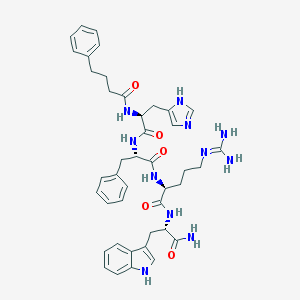 CHEMBL431801 CHEMBL431801 | C42H51N11O5 | 789.942 | 7 / 9 | 2.2 | No |
| 468092 | 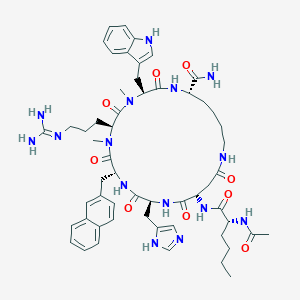 CHEMBL3600840 CHEMBL3600840 | C56H75N15O9 | 1102.31 | 11 / 11 | 1.6 | No |
| 42938 | 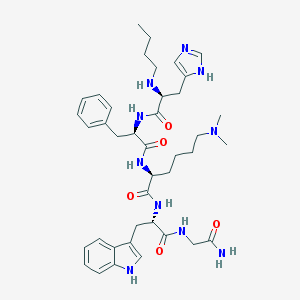 CHEMBL420581 CHEMBL420581 | C40H56N10O5 | 756.953 | 8 / 8 | 2.8 | No |
| 43875 | 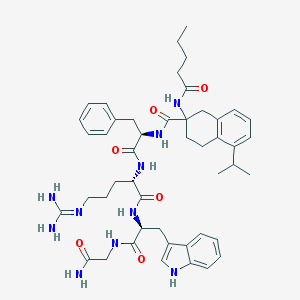 CHEMBL434966 CHEMBL434966 | C47H62N10O6 | 863.077 | 7 / 9 | 3.9 | No |
| 44324 | 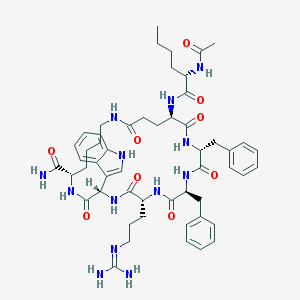 CHEMBL269353 CHEMBL269353 | C53H71N13O9 | 1034.23 | 10 / 12 | 1.9 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218