

You can:
| Name | Substance-P receptor |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | TACR1 |
| Synonym | NK-1 receptor Tachykinin receptor 1 TAC1R SPR Substance P receptor [ Show all ] |
| Disease | Cough Depression Depression; Anxiety Diabetes Eczema [ Show all ] |
| Length | 407 |
| Amino acid sequence | MDNVLPVDSDLSPNISTNTSEPNQFVQPAWQIVLWAAAYTVIVVTSVVGNVVVMWIILAHKRMRTVTNYFLVNLAFAEASMAAFNTVVNFTYAVHNEWYYGLFYCKFHNFFPIAAVFASIYSMTAVAFDRYMAIIHPLQPRLSATATKVVICVIWVLALLLAFPQGYYSTTETMPSRVVCMIEWPEHPNKIYEKVYHICVTVLIYFLPLLVIGYAYTVVGITLWASEIPGDSSDRYHEQVSAKRKVVKMMIVVVCTFAICWLPFHIFFLLPYINPDLYLKKFIQQVYLAIMWLAMSSTMYNPIIYCCLNDRFRLGFKHAFRCCPFISAGDYEGLEMKSTRYLQTQGSVYKVSRLETTISTVVGAHEEEPEDGPKATPSSLDLTSNCSSRSDSKTMTESFSFSSNVLS |
| UniProt | P25103 |
| Protein Data Bank | 2ksa, 6e59, 6hlo, 2ks9, 6hlp, 6hll, 2ksb |
| GPCR-HGmod model | P25103 |
| 3D structure model | This structure is from PDB ID 2ksa. |
| BioLiP | BL0101802, BL0437915, BL0437914, BL0437913, BL0434896, BL0101803, BL0101801 |
| Therapeutic Target Database | T47094 |
| ChEMBL | CHEMBL249 |
| IUPHAR | 360 |
| DrugBank | BE0000384 |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 312 | 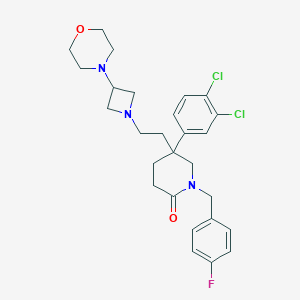 CHEMBL358600 CHEMBL358600 | C27H32Cl2FN3O2 | 520.47 | 5 / 0 | 4.3 | No |
| 347 | 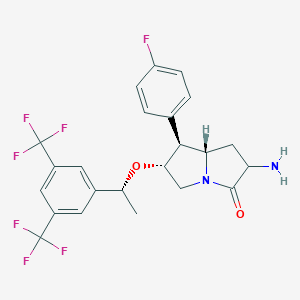 CHEMBL1097693 CHEMBL1097693 | C23H21F7N2O2 | 490.422 | 10 / 1 | 4.3 | Yes |
| 396 | 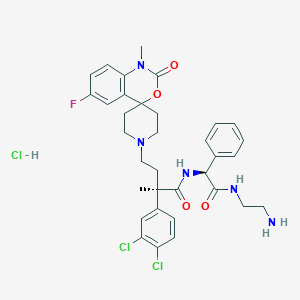 CHEMBL3288163 CHEMBL3288163 | C34H39Cl3FN5O4 | 707.065 | 7 / 4 | N/A | No |
| 777 | 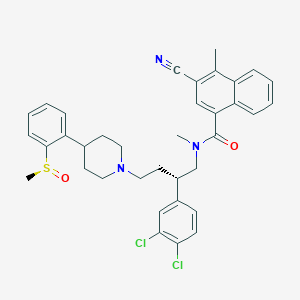 CHEMBL408242 CHEMBL408242 | C36H37Cl2N3O2S | 646.671 | 5 / 0 | 7.5 | No |
| 1038 | 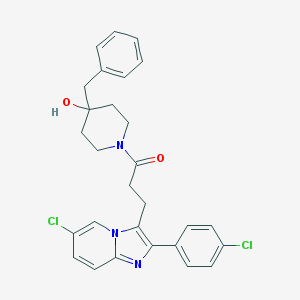 CHEMBL21607 CHEMBL21607 | C28H27Cl2N3O2 | 508.443 | 3 / 1 | 6.1 | No |
| 1104 | 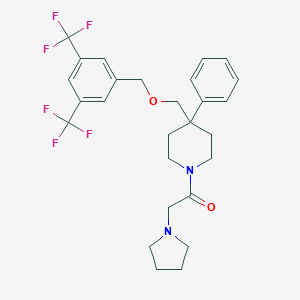 CHEMBL143156 CHEMBL143156 | C27H30F6N2O2 | 528.539 | 9 / 0 | 5.6 | No |
| 1179 | 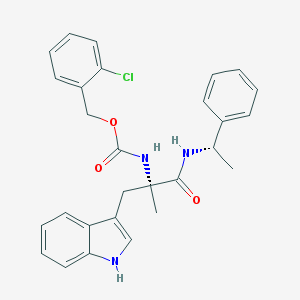 CHEMBL134737 CHEMBL134737 | C28H28ClN3O3 | 490.0 | 3 / 3 | 5.6 | No |
| 1274 | 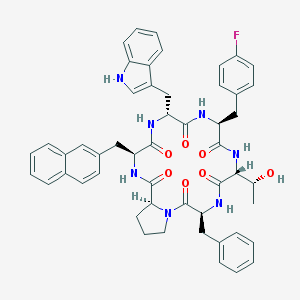 CHEMBL418757 CHEMBL418757 | C51H52FN7O7 | 894.017 | 8 / 7 | 6.6 | No |
| 1512 | 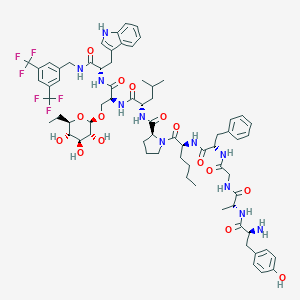 CHEMBL556537 CHEMBL556537 | C70H89F6N11O15 | 1438.53 | 22 / 14 | 5.8 | No |
| 1580 | 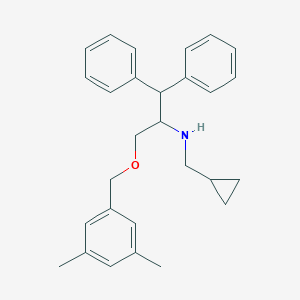 CHEMBL417766 CHEMBL417766 | C28H33NO | 399.578 | 2 / 1 | 6.1 | No |
| 1623 | 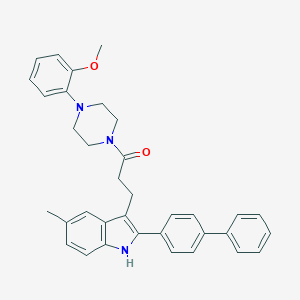 CHEMBL21939 CHEMBL21939 | C35H35N3O2 | 529.684 | 3 / 1 | 7.0 | No |
| 1720 | 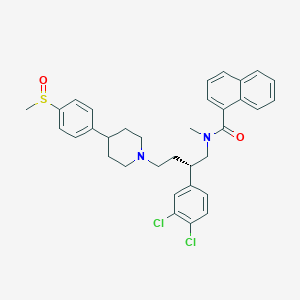 CHEMBL2373196 CHEMBL2373196 | C34H36Cl2N2O2S | 607.634 | 4 / 0 | 7.4 | No |
| 1836 | 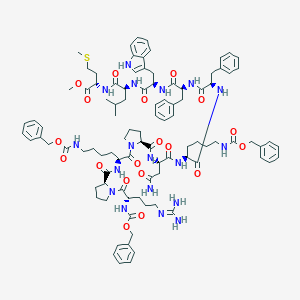 CHEMBL2304114 CHEMBL2304114 | C97H126N18O19S | 1880.24 | 21 / 15 | 7.8 | No |
| 1876 | 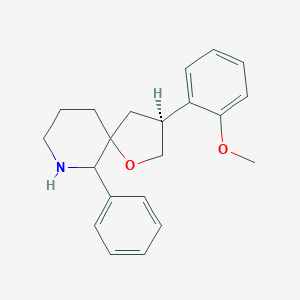 CHEMBL327928 CHEMBL327928 | C21H25NO2 | 323.436 | 3 / 1 | 3.4 | Yes |
| 1889 | 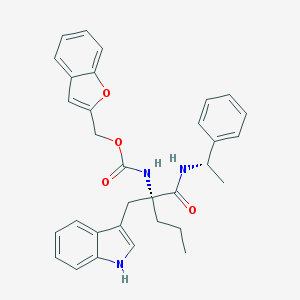 CHEMBL76455 CHEMBL76455 | C32H33N3O4 | 523.633 | 4 / 3 | 6.3 | No |
| 2342 | 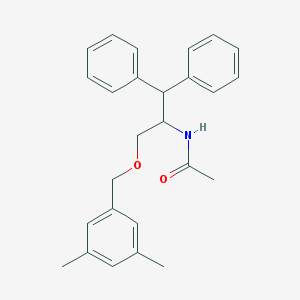 CHEMBL59310 CHEMBL59310 | C26H29NO2 | 387.523 | 2 / 1 | 5.1 | No |
| 2488 | 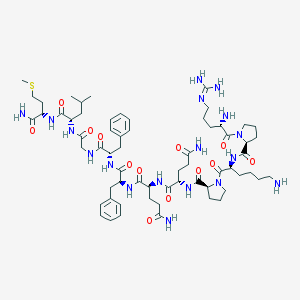 Substance P Substance P | C63H98N18O13S | 1347.65 | 17 / 15 | -2.3 | No |
| 2798 | 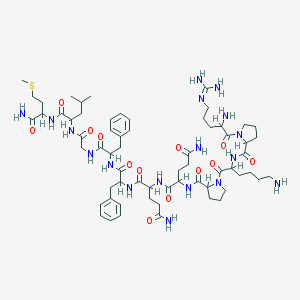 sub-stance p sub-stance p | C63H98N18O13S | 1347.65 | 17 / 15 | -2.3 | No |
| 463228 | 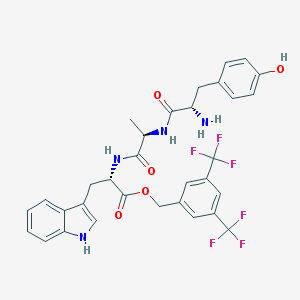 CHEMBL3609619 CHEMBL3609619 | C32H30F6N4O5 | 664.605 | 12 / 5 | 4.5 | No |
| 3121 | 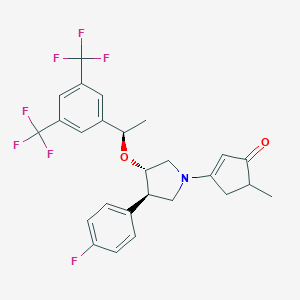 CHEMBL400148 CHEMBL400148 | C26H24F7NO2 | 515.472 | 10 / 0 | 6.2 | No |
| 3175 | 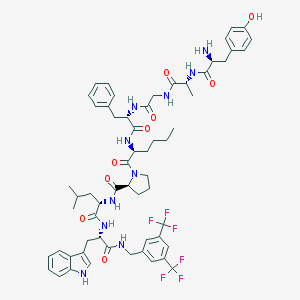 CHEMBL557188 CHEMBL557188 | C60H72F6N10O9 | 1191.29 | 16 / 10 | 7.4 | No |
| 3187 | 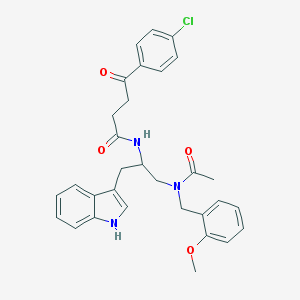 CHEMBL42653 CHEMBL42653 | C31H32ClN3O4 | 546.064 | 4 / 2 | 4.5 | No |
| 3200 | 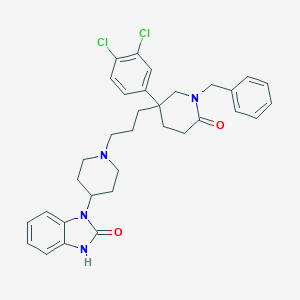 CHEMBL26790 CHEMBL26790 | C33H36Cl2N4O2 | 591.577 | 3 / 1 | 6.1 | No |
| 3429 | 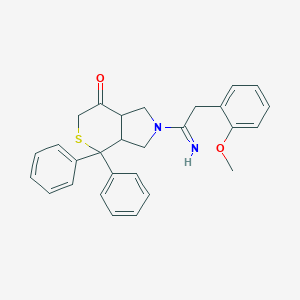 CHEMBL288371 CHEMBL288371 | C28H28N2O2S | 456.604 | 4 / 1 | 4.7 | Yes |
| 3447 | 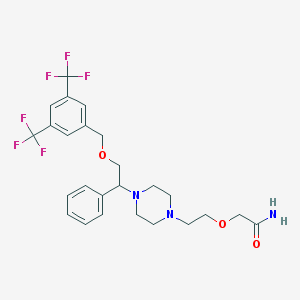 CHEMBL337789 CHEMBL337789 | C25H29F6N3O3 | 533.515 | 11 / 1 | 3.5 | No |
| 3448 | 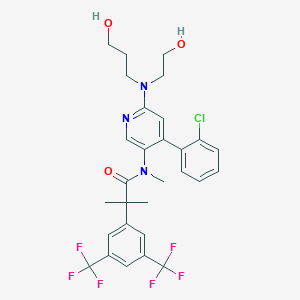 CHEMBL1084165 CHEMBL1084165 | C29H30ClF6N3O3 | 618.017 | 11 / 2 | 6.2 | No |
| 3515 | 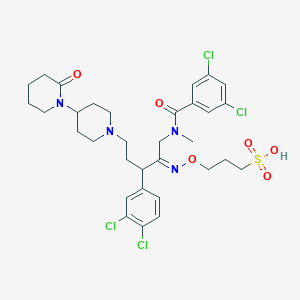 CHEMBL166827 CHEMBL166827 | C32H40Cl4N4O6S | 750.554 | 8 / 1 | 3.7 | No |
| 3849 | 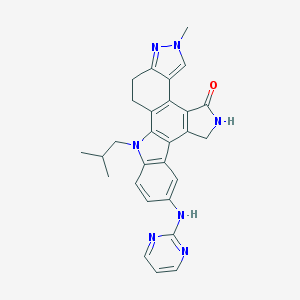 CEP-11981 CEP-11981 | C28H27N7O | 477.572 | 5 / 2 | 3.9 | Yes |
| 3973 | 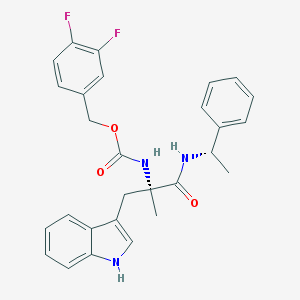 CHEMBL341681 CHEMBL341681 | C28H27F2N3O3 | 491.539 | 5 / 3 | 5.2 | No |
| 4672 | 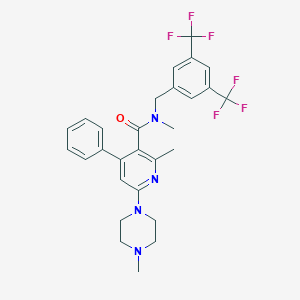 CHEMBL1765500 CHEMBL1765500 | C28H28F6N4O | 550.549 | 10 / 0 | 5.9 | No |
| 463457 | 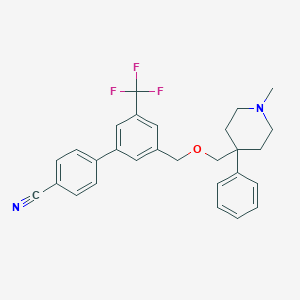 CHEMBL3596492 CHEMBL3596492 | C28H27F3N2O | 464.532 | 6 / 0 | 6.1 | No |
| 4838 | 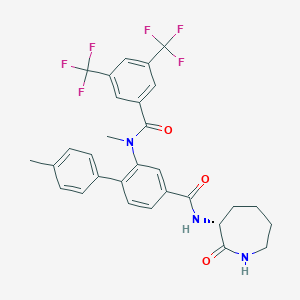 CHEMBL2112211 CHEMBL2112211 | C30H27F6N3O3 | 591.554 | 9 / 2 | 6.2 | No |
| 5216 | 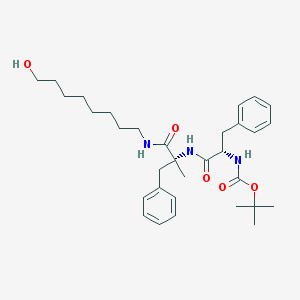 CHEMBL296014 CHEMBL296014 | C32H47N3O5 | 553.744 | 5 / 4 | 5.6 | No |
| 5397 | 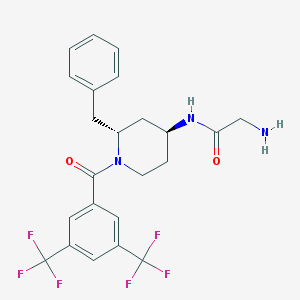 CHEMBL42800 CHEMBL42800 | C23H23F6N3O2 | 487.446 | 9 / 2 | 4.1 | Yes |
| 5405 | 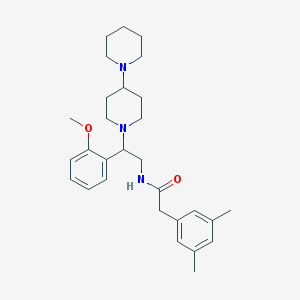 CHEMBL137972 CHEMBL137972 | C29H41N3O2 | 463.666 | 4 / 1 | 4.8 | Yes |
| 5705 | 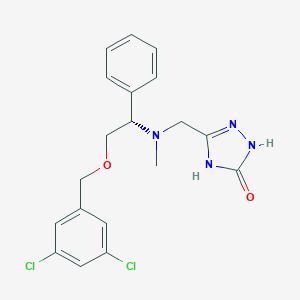 CHEMBL357303 CHEMBL357303 | C19H20Cl2N4O2 | 407.295 | 4 / 2 | 3.4 | Yes |
| 5755 | 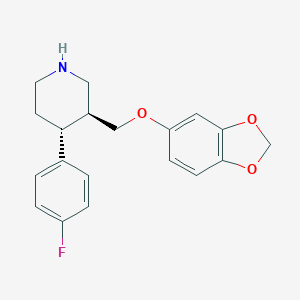 paroxetine paroxetine | C19H20FNO3 | 329.371 | 5 / 1 | 3.5 | Yes |
| 6022 | 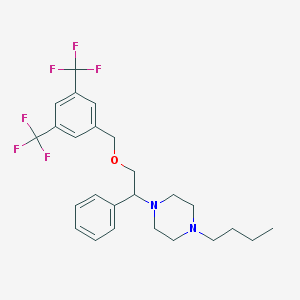 CHEMBL336251 CHEMBL336251 | C25H30F6N2O | 488.518 | 9 / 0 | 6.0 | No |
| 6026 | 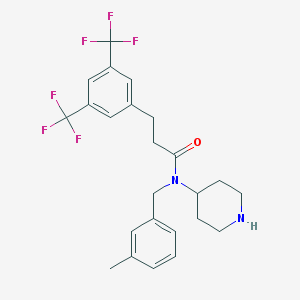 CHEMBL313070 CHEMBL313070 | C24H26F6N2O | 472.475 | 8 / 1 | 5.4 | No |
| 6045 | 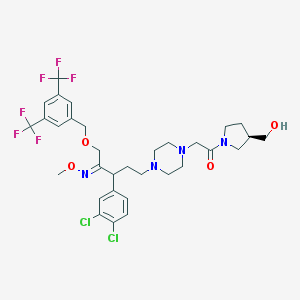 CHEMBL75367 CHEMBL75367 | C32H38Cl2F6N4O4 | 727.57 | 13 / 1 | 6.0 | No |
| 6047 | 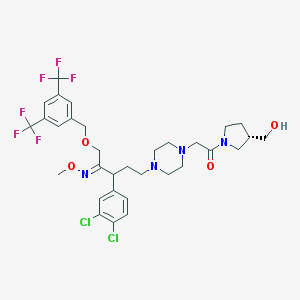 CHEMBL75436 CHEMBL75436 | C32H38Cl2F6N4O4 | 727.57 | 13 / 1 | 6.0 | No |
| 6179 | 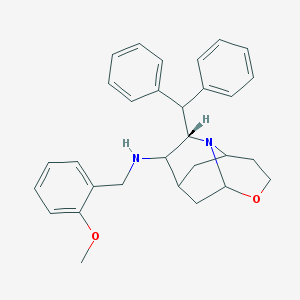 CHEMBL317297 CHEMBL317297 | C30H34N2O2 | 454.614 | 4 / 1 | 5.6 | No |
| 6195 | 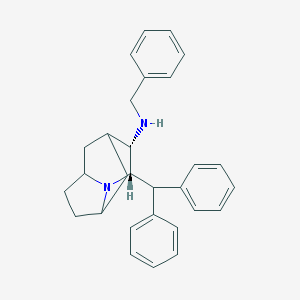 CHEMBL420876 CHEMBL420876 | C29H32N2 | 408.589 | 2 / 1 | 6.0 | No |
| 6196 | 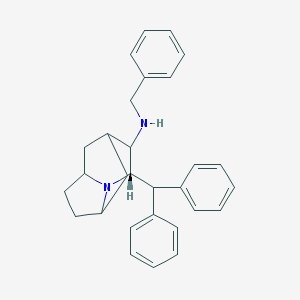 CHEMBL316842 CHEMBL316842 | C29H32N2 | 408.589 | 2 / 1 | 6.0 | No |
| 6452 | 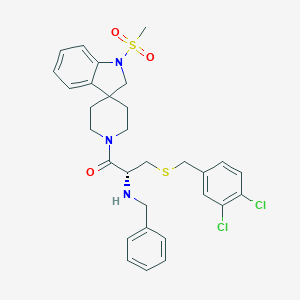 CHEMBL445357 CHEMBL445357 | C30H33Cl2N3O3S2 | 618.632 | 6 / 1 | 5.4 | No |
| 6649 | 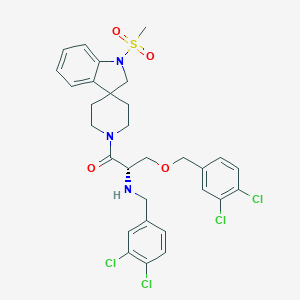 CHEMBL54540 CHEMBL54540 | C30H31Cl4N3O4S | 671.455 | 6 / 1 | 5.9 | No |
| 7266 | 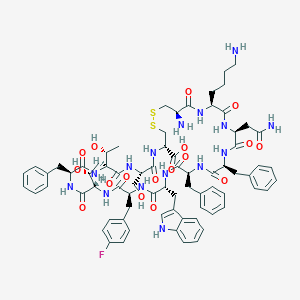 CHEMBL2112245 CHEMBL2112245 | C74H92FN15O17S2 | 1546.76 | 22 / 19 | -0.3 | No |
| 7547 | 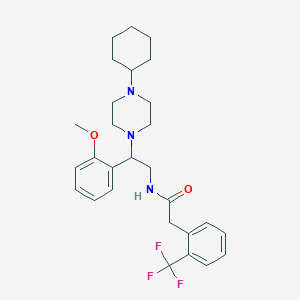 CHEMBL139256 CHEMBL139256 | C28H36F3N3O2 | 503.61 | 7 / 1 | 5.2 | No |
| 7953 | 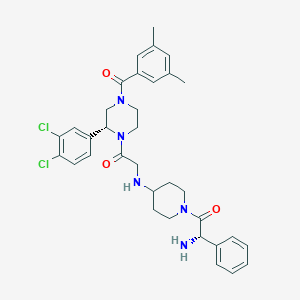 CHEMBL106300 CHEMBL106300 | C34H39Cl2N5O3 | 636.618 | 5 / 2 | 4.6 | No |
| 7954 | 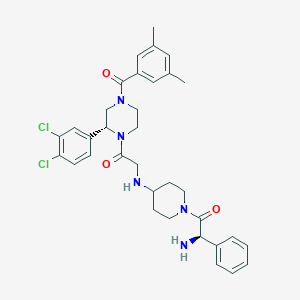 CHEMBL107306 CHEMBL107306 | C34H39Cl2N5O3 | 636.618 | 5 / 2 | 4.6 | No |
| 7957 | 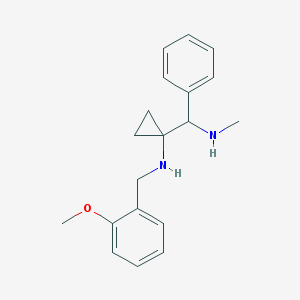 CHEMBL423597 CHEMBL423597 | C19H24N2O | 296.414 | 3 / 2 | 2.7 | Yes |
| 8035 | 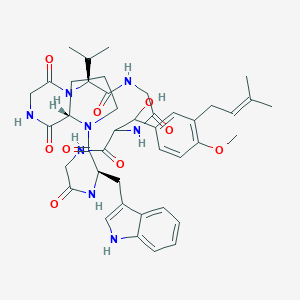 CHEMBL2370513 CHEMBL2370513 | C42H54N8O9 | 814.941 | 9 / 8 | 2.9 | No |
| 8221 | 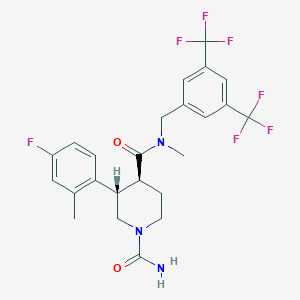 CHEMBL1951633 CHEMBL1951633 | C24H24F7N3O2 | 519.464 | 9 / 1 | 4.2 | No |
| 8304 | 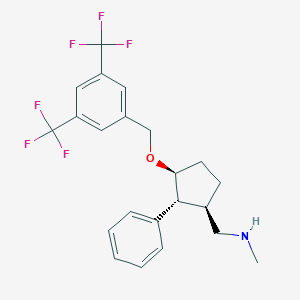 CHEMBL209531 CHEMBL209531 | C22H23F6NO | 431.422 | 8 / 1 | 5.5 | No |
| 8427 | 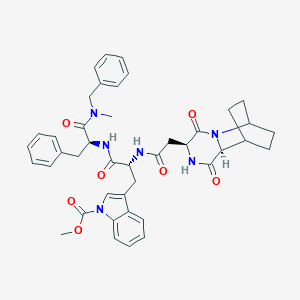 CHEMBL3144347 CHEMBL3144347 | C42H46N6O7 | 746.865 | 7 / 3 | 4.2 | No |
| 8432 | 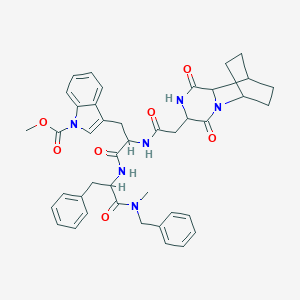 CHEMBL45085 CHEMBL45085 | C42H46N6O7 | 746.865 | 7 / 3 | 4.2 | No |
| 8982 | 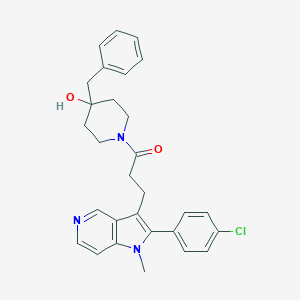 CHEMBL20826 CHEMBL20826 | C29H30ClN3O2 | 488.028 | 3 / 1 | 4.6 | Yes |
| 9058 | 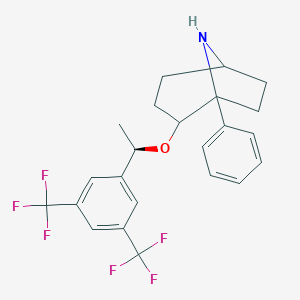 CHEMBL207698 CHEMBL207698 | C23H23F6NO | 443.433 | 8 / 1 | 5.8 | No |
| 9059 | 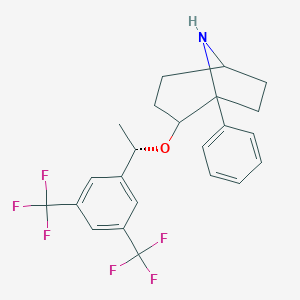 CHEMBL382988 CHEMBL382988 | C23H23F6NO | 443.433 | 8 / 1 | 5.8 | No |
| 9142 | 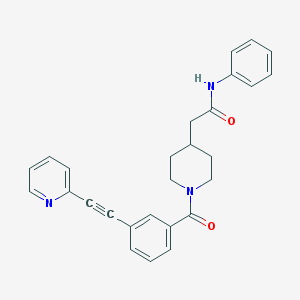 CHEMBL208900 CHEMBL208900 | C27H25N3O2 | 423.516 | 3 / 1 | 3.9 | Yes |
| 9226 | 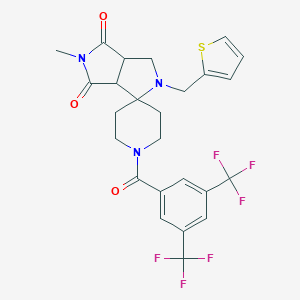 CHEMBL317209 CHEMBL317209 | C25H23F6N3O3S | 559.527 | 11 / 0 | 3.5 | No |
| 9301 | 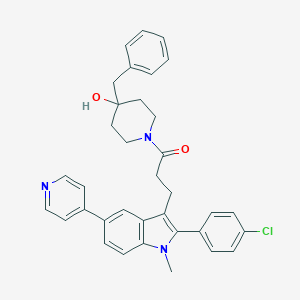 CHEMBL20736 CHEMBL20736 | C35H34ClN3O2 | 564.126 | 3 / 1 | 6.2 | No |
| 9359 | 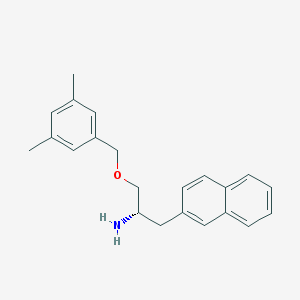 CHEMBL120589 CHEMBL120589 | C22H25NO | 319.448 | 2 / 1 | 4.6 | Yes |
| 9360 | 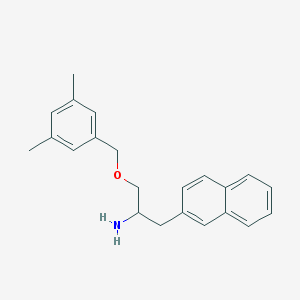 CHEMBL68670 CHEMBL68670 | C22H25NO | 319.448 | 2 / 1 | 4.6 | Yes |
| 9429 | 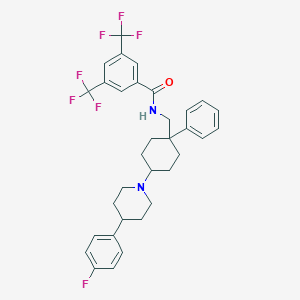 CHEMBL301618 CHEMBL301618 | C33H33F7N2O | 606.629 | 9 / 1 | 8.4 | No |
| 9451 | 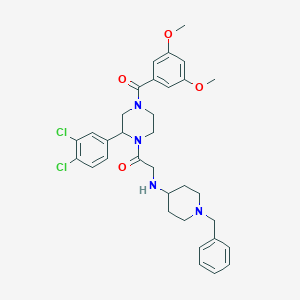 CHEMBL319474 CHEMBL319474 | C33H38Cl2N4O4 | 625.591 | 6 / 1 | 5.1 | No |
| 9776 | 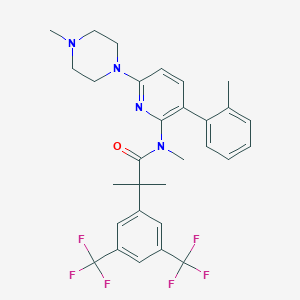 CHEMBL380510 CHEMBL380510 | C30H32F6N4O | 578.603 | 10 / 0 | 7.1 | No |
| 9820 | 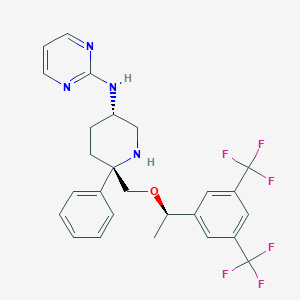 CHEMBL1270464 CHEMBL1270464 | C26H26F6N4O | 524.511 | 11 / 2 | 5.5 | No |
| 9869 | 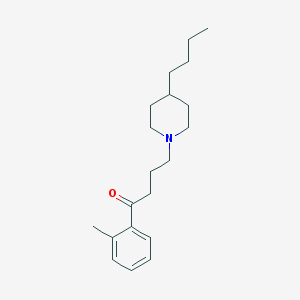 AC-42 AC-42 | C20H31NO | 301.474 | 2 / 0 | 5.2 | No |
| 9956 | 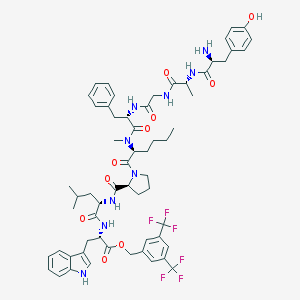 CHEMBL387670 CHEMBL387670 | C61H73F6N9O10 | 1206.3 | 17 / 8 | 8.1 | No |
| 10040 | 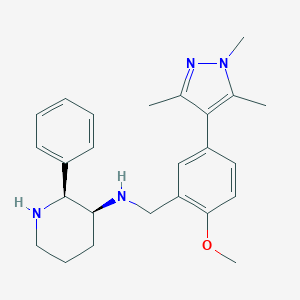 CHEMBL144793 CHEMBL144793 | C25H32N4O | 404.558 | 4 / 2 | 3.7 | Yes |
| 557571 | 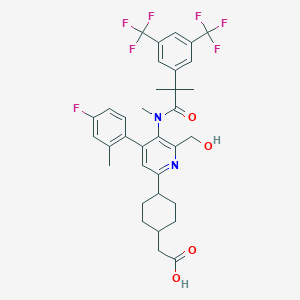 SCHEMBL17254991 SCHEMBL17254991 | C34H35F7N2O4 | 668.653 | 12 / 2 | 7.2 | No |
| 10200 | 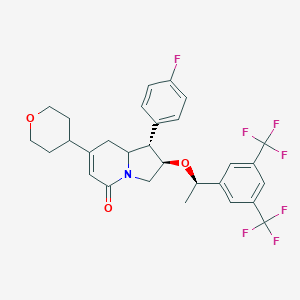 CHEMBL1090438 CHEMBL1090438 | C29H28F7NO3 | 571.536 | 10 / 0 | 5.5 | No |
| 10246 | 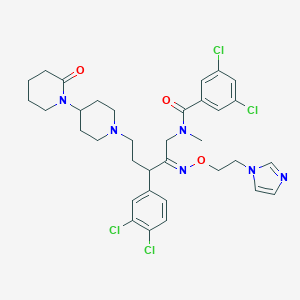 CHEMBL168144 CHEMBL168144 | C34H40Cl4N6O3 | 722.533 | 6 / 0 | 6.4 | No |
| 10603 | 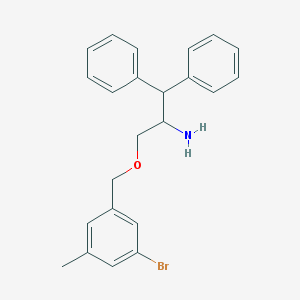 CHEMBL56372 CHEMBL56372 | C23H24BrNO | 410.355 | 2 / 1 | 5.2 | No |
| 521746 | 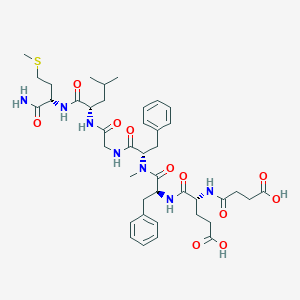 CHEMBL3735396 CHEMBL3735396 | C41H57N7O11S | 856.005 | 12 / 8 | 1.5 | No |
| 521749 | 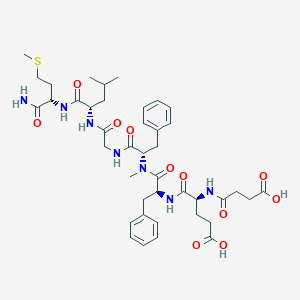 CHEMBL3735210 CHEMBL3735210 | C41H57N7O11S | 856.005 | 12 / 8 | 1.5 | No |
| 11050 | 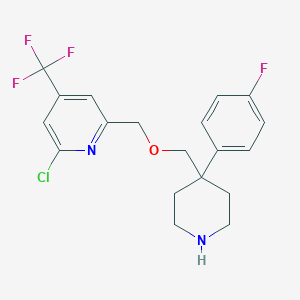 CHEMBL2333642 CHEMBL2333642 | C19H19ClF4N2O | 402.818 | 7 / 1 | 4.3 | Yes |
| 11074 | 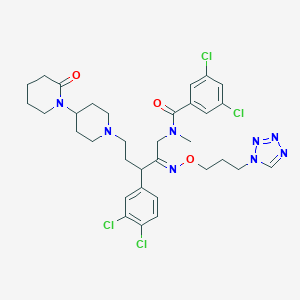 CHEMBL287369 CHEMBL287369 | C33H40Cl4N8O3 | 738.536 | 8 / 0 | 6.4 | No |
| 11141 | 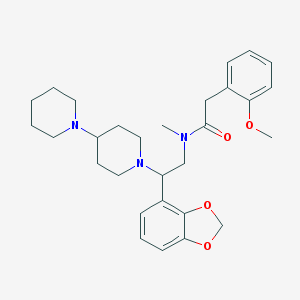 CHEMBL139057 CHEMBL139057 | C29H39N3O4 | 493.648 | 6 / 0 | 4.1 | Yes |
| 12343 | 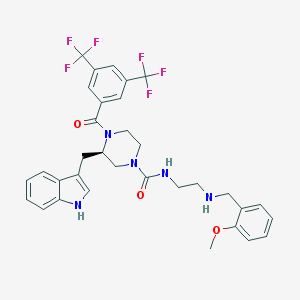 CHEMBL203113 CHEMBL203113 | C33H33F6N5O3 | 661.649 | 10 / 3 | 5.4 | No |
| 12445 | 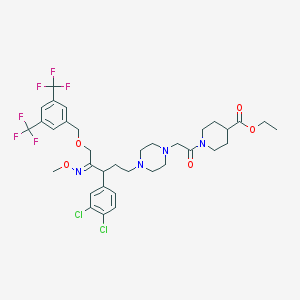 CHEMBL41068 CHEMBL41068 | C35H42Cl2F6N4O5 | 783.634 | 14 / 0 | 7.0 | No |
| 12539 | 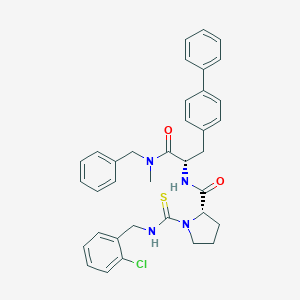 CHEMBL105847 CHEMBL105847 | C36H37ClN4O2S | 625.228 | 3 / 2 | 6.8 | No |
| 521828 | 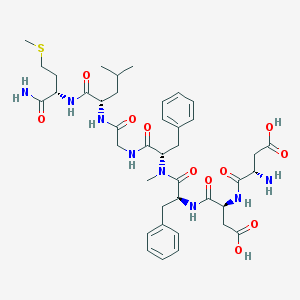 CHEMBL3736313 CHEMBL3736313 | C40H56N8O11S | 856.993 | 13 / 9 | -2.0 | No |
| 12653 | 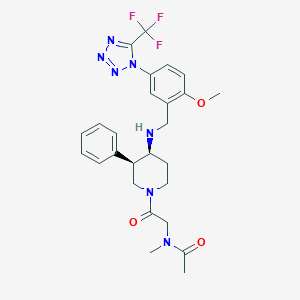 CHEMBL1917867 CHEMBL1917867 | C26H30F3N7O3 | 545.567 | 10 / 1 | 2.5 | No |
| 12963 | 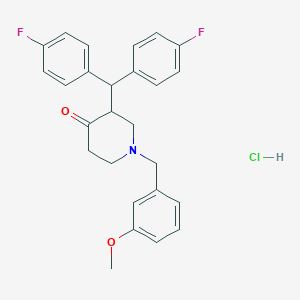 CHEMBL1817830 CHEMBL1817830 | C26H26ClF2NO2 | 457.946 | 5 / 1 | N/A | N/A |
| 13477 | 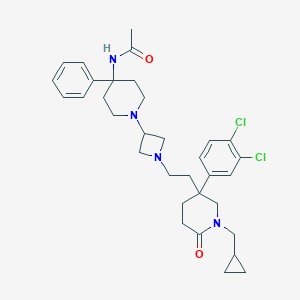 CHEMBL151356 CHEMBL151356 | C33H42Cl2N4O2 | 597.625 | 4 / 1 | 5.0 | No |
| 13533 | 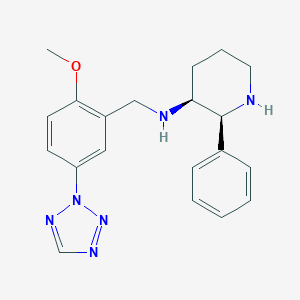 CHEMBL424301 CHEMBL424301 | C20H24N6O | 364.453 | 6 / 2 | 2.8 | Yes |
| 13712 | 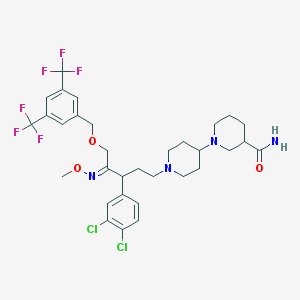 CHEMBL342341 CHEMBL342341 | C32H38Cl2F6N4O3 | 711.571 | 12 / 1 | 6.9 | No |
| 13771 | 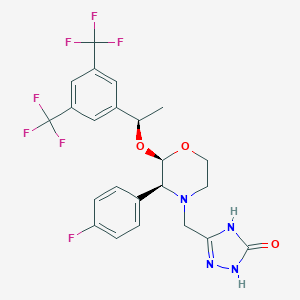 Aprepitant Aprepitant | C23H21F7N4O3 | 534.435 | 12 / 2 | 4.2 | No |
| 13773 | 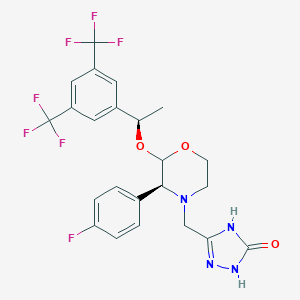 ent-Aprepitant ent-Aprepitant | C23H21F7N4O3 | 534.435 | 12 / 2 | 4.2 | No |
| 14024 | 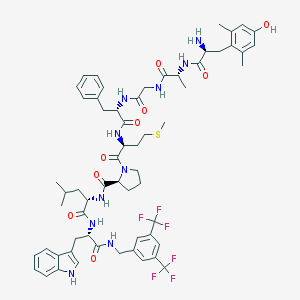 CHEMBL1766208 CHEMBL1766208 | C61H74F6N10O9S | 1237.37 | 17 / 10 | 7.3 | No |
| 14089 | 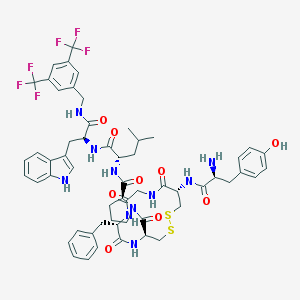 CHEMBL611925 CHEMBL611925 | C57H64F6N10O9S2 | 1211.31 | 18 / 10 | 6.0 | No |
| 14093 | 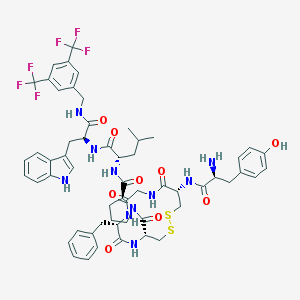 CHEMBL611924 CHEMBL611924 | C57H64F6N10O9S2 | 1211.31 | 18 / 10 | 6.0 | No |
| 14107 | 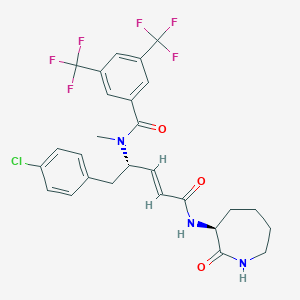 CHEMBL2114398 CHEMBL2114398 | C27H26ClF6N3O3 | 589.963 | 9 / 2 | 5.6 | No |
| 14110 | 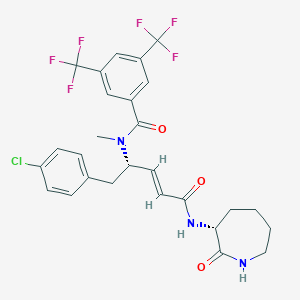 CHEMBL2112070 CHEMBL2112070 | C27H26ClF6N3O3 | 589.963 | 9 / 2 | 5.6 | No |
| 14111 | 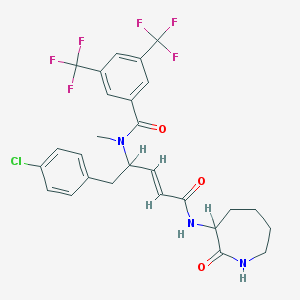 CHEMBL35857 CHEMBL35857 | C27H26ClF6N3O3 | 589.963 | 9 / 2 | 5.6 | No |
| 14113 | 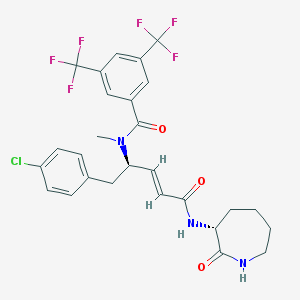 CHEMBL104954 CHEMBL104954 | C27H26ClF6N3O3 | 589.963 | 9 / 2 | 5.6 | No |
| 14115 | 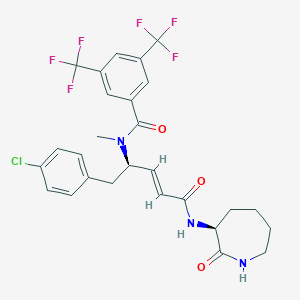 CHEMBL2115343 CHEMBL2115343 | C27H26ClF6N3O3 | 589.963 | 9 / 2 | 5.6 | No |
| 14137 | 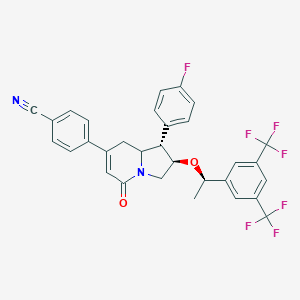 CHEMBL1090285 CHEMBL1090285 | C31H23F7N2O2 | 588.526 | 10 / 0 | 6.5 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218