

You can:
| Name | Platelet-activating factor receptor |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | PTAFR |
| Synonym | PAFr PAF-R PAF receptor AGEPC receptor |
| Disease | Nerve injury Ocular allergy Pain Unspecified Psoriasis [ Show all ] |
| Length | 342 |
| Amino acid sequence | MEPHDSSHMDSEFRYTLFPIVYSIIFVLGVIANGYVLWVFARLYPCKKFNEIKIFMVNLTMADMLFLITLPLWIVYYQNQGNWILPKFLCNVAGCLFFINTYCSVAFLGVITYNRFQAVTRPIKTAQANTRKRGISLSLVIWVAIVGAASYFLILDSTNTVPDSAGSGNVTRCFEHYEKGSVPVLIIHIFIVFSFFLVFLIILFCNLVIIRTLLMQPVQQQRNAEVKRRALWMVCTVLAVFIICFVPHHVVQLPWTLAELGFQDSKFHQAINDAHQVTLCLLSTNCVLDPVIYCFLTKKFRKHLTEKFYSMRSSRKCSRATTDTVTEVVVPFNQIPGNSLKN |
| UniProt | P25105 |
| Protein Data Bank | 5zkq, 5zkp |
| GPCR-HGmod model | P25105 |
| 3D structure model | This structure is from PDB ID 5zkq. |
| BioLiP | BL0417415, BL0417417,BL0417419, BL0417416,BL0417418, BL0417414 |
| Therapeutic Target Database | T87023 |
| ChEMBL | CHEMBL250 |
| IUPHAR | 334 |
| DrugBank | BE0005561 |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 767 | 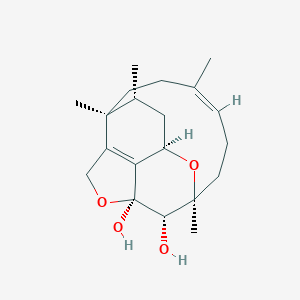 Phomactin A Phomactin A | C20H30O4 | 334.456 | 4 / 2 | 1.3 | Yes |
| 768 | 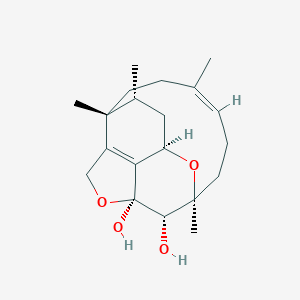 CHEMBL2373416 CHEMBL2373416 | C20H30O4 | 334.456 | 4 / 2 | 1.3 | Yes |
| 1606 | 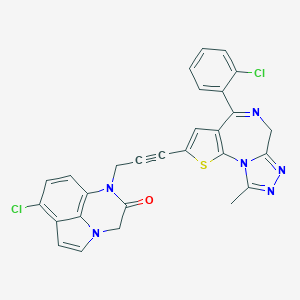 CHEMBL277678 CHEMBL277678 | C28H18Cl2N6OS | 557.453 | 5 / 0 | 5.1 | No |
| 1684 | 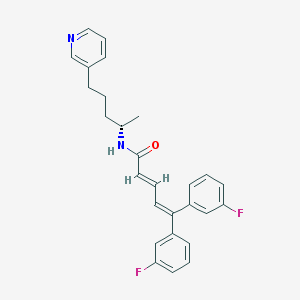 CHEMBL52933 CHEMBL52933 | C27H26F2N2O | 432.515 | 4 / 1 | 6.4 | No |
| 1686 | 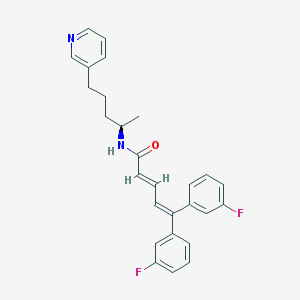 CHEMBL54935 CHEMBL54935 | C27H26F2N2O | 432.515 | 4 / 1 | 6.4 | No |
| 1736 | 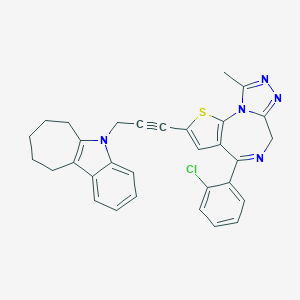 CHEMBL20644 CHEMBL20644 | C31H26ClN5S | 536.094 | 4 / 0 | 7.1 | No |
| 2248 | 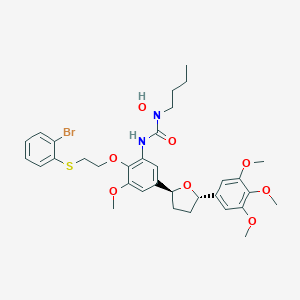 CHEMBL2113721 CHEMBL2113721 | C33H41BrN2O8S | 705.661 | 9 / 2 | 6.3 | No |
| 3309 | 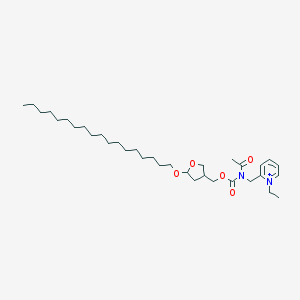 CHEMBL290840 CHEMBL290840 | C34H59N2O5+ | 575.855 | 5 / 0 | 9.9 | No |
| 4050 | 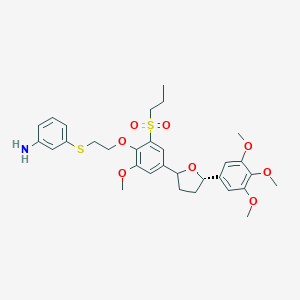 CHEMBL51798 CHEMBL51798 | C31H39NO8S2 | 617.772 | 10 / 1 | 5.1 | No |
| 4125 | 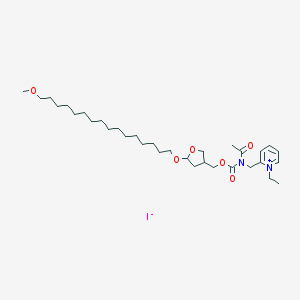 CHEMBL158974 CHEMBL158974 | C33H57IN2O6 | 704.731 | 7 / 0 | N/A | No |
| 4813 | 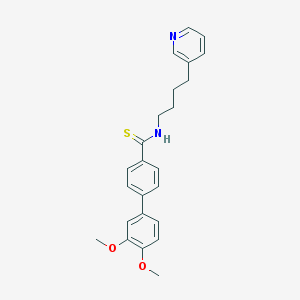 CHEMBL300644 CHEMBL300644 | C24H26N2O2S | 406.544 | 4 / 1 | 5.0 | Yes |
| 5785 | 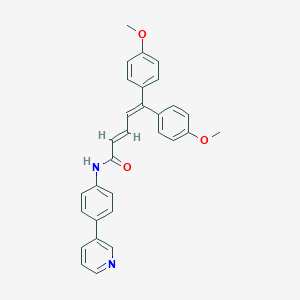 CHEMBL53014 CHEMBL53014 | C30H26N2O3 | 462.549 | 4 / 1 | 6.2 | No |
| 5863 | 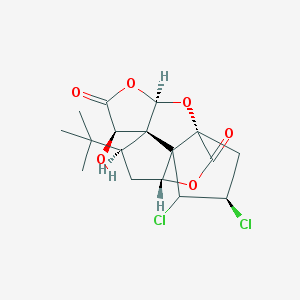 CHEMBL2304197 CHEMBL2304197 | C17H20Cl2O6 | 391.241 | 6 / 1 | 2.5 | Yes |
| 5970 | 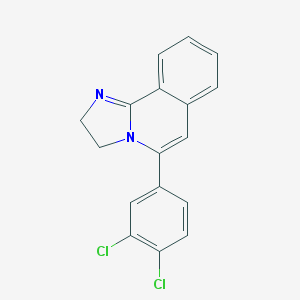 CHEMBL310866 CHEMBL310866 | C17H12Cl2N2 | 315.197 | 1 / 0 | 4.1 | Yes |
| 553270 | 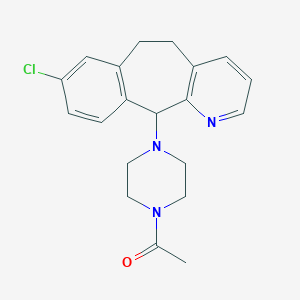 CHEMBL326821 CHEMBL326821 | C20H22ClN3O | 355.866 | 3 / 0 | 2.6 | Yes |
| 6724 | 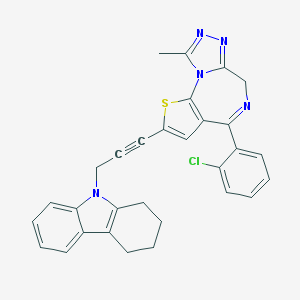 CHEMBL278072 CHEMBL278072 | C30H24ClN5S | 522.067 | 4 / 0 | 6.6 | No |
| 8286 | 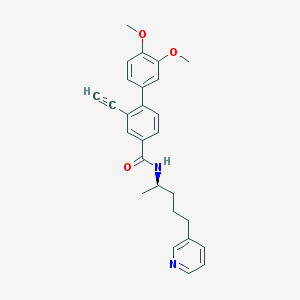 CHEMBL430969 CHEMBL430969 | C27H28N2O3 | 428.532 | 4 / 1 | 5.0 | Yes |
| 8345 | 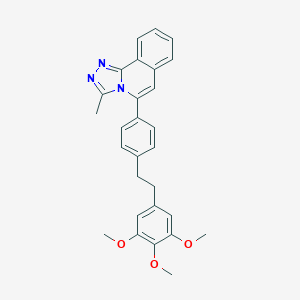 CHEMBL106358 CHEMBL106358 | C28H27N3O3 | 453.542 | 5 / 0 | 6.6 | No |
| 9088 | 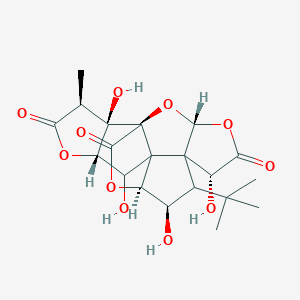 CHEBI:5357 CHEBI:5357 | C20H24O11 | 440.401 | 11 / 4 | -1.4 | No |
| 9610 | 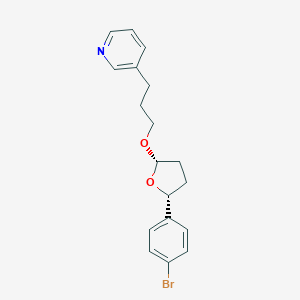 CHEMBL37547 CHEMBL37547 | C18H20BrNO2 | 362.267 | 3 / 0 | 4.0 | Yes |
| 9611 | 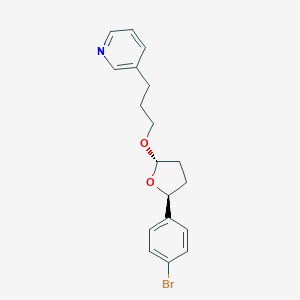 CHEMBL291082 CHEMBL291082 | C18H20BrNO2 | 362.267 | 3 / 0 | 4.0 | Yes |
| 10078 | 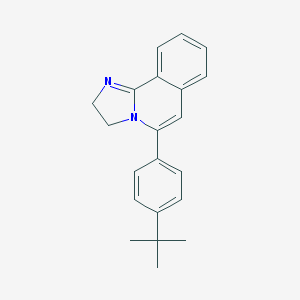 CHEMBL79089 CHEMBL79089 | C21H22N2 | 302.421 | 1 / 0 | 4.5 | Yes |
| 10573 | 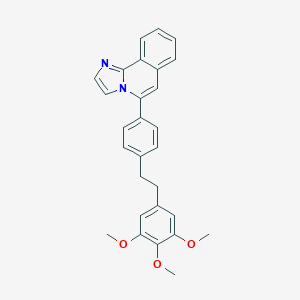 CHEMBL320577 CHEMBL320577 | C28H26N2O3 | 438.527 | 4 / 0 | 6.8 | No |
| 10586 | 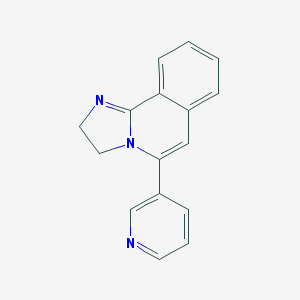 CHEMBL80505 CHEMBL80505 | C16H13N3 | 247.301 | 2 / 0 | 1.8 | Yes |
| 11225 | 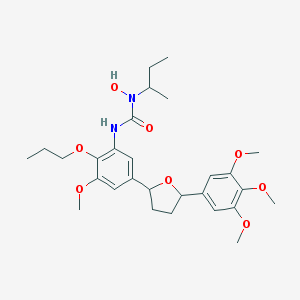 CHEMBL160290 CHEMBL160290 | C28H40N2O8 | 532.634 | 8 / 2 | 4.4 | No |
| 11373 | 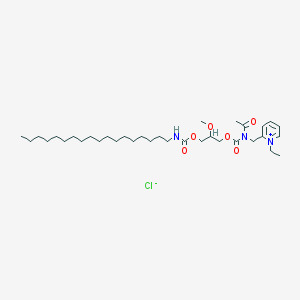 CV-6209 CV-6209 | C34H60ClN3O6 | 642.319 | 7 / 1 | N/A | No |
| 12047 | 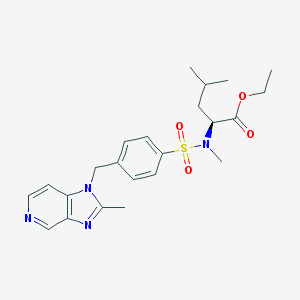 LEXIPAFANT LEXIPAFANT | C23H30N4O4S | 458.577 | 7 / 0 | 3.5 | Yes |
| 13751 | 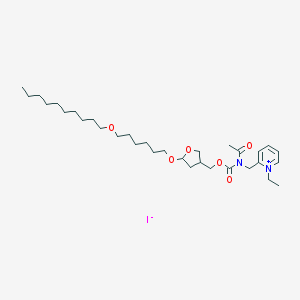 CHEMBL421843 CHEMBL421843 | C32H55IN2O6 | 690.704 | 7 / 0 | N/A | No |
| 13787 | 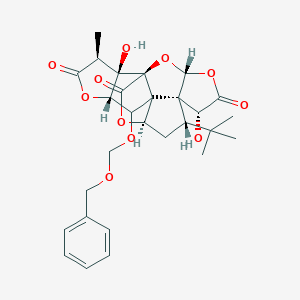 CHEMBL2304166 CHEMBL2304166 | C28H32O11 | 544.553 | 11 / 2 | 1.6 | No |
| 14081 | 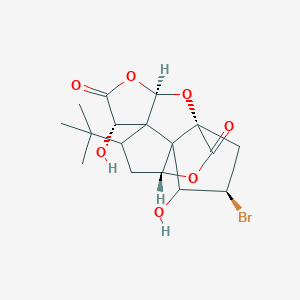 CHEMBL23779 CHEMBL23779 | C17H21BrO7 | 417.252 | 7 / 2 | 1.4 | Yes |
| 17141 | 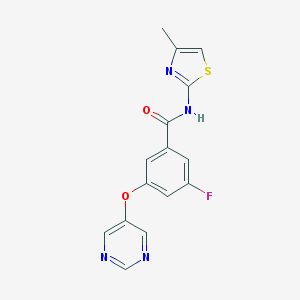 CHEMBL2440659 CHEMBL2440659 | C15H11FN4O2S | 330.337 | 7 / 1 | 2.4 | Yes |
| 17821 | 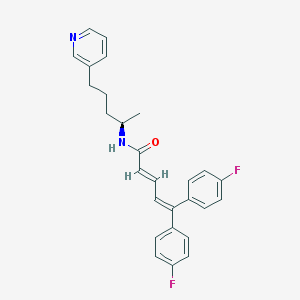 CHEMBL300742 CHEMBL300742 | C27H26F2N2O | 432.515 | 4 / 1 | 6.4 | No |
| 17823 | 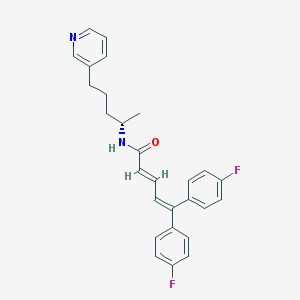 CHEMBL299780 CHEMBL299780 | C27H26F2N2O | 432.515 | 4 / 1 | 6.4 | No |
| 19376 | 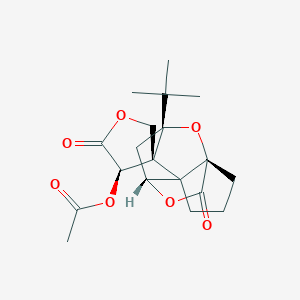 CHEMBL2304169 CHEMBL2304169 | C19H24O7 | 364.394 | 7 / 0 | 1.9 | Yes |
| 20427 | 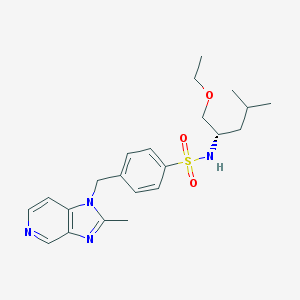 BB-823 BB-823 | C22H30N4O3S | 430.567 | 6 / 1 | 3.3 | Yes |
| 20828 | 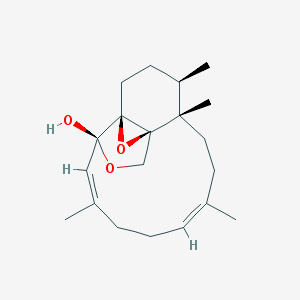 CID 21577214 CID 21577214 | C20H30O3 | 318.457 | 3 / 1 | 2.7 | Yes |
| 21180 | 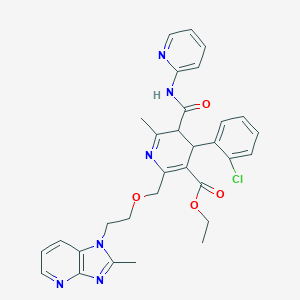 BDBM50286211 BDBM50286211 | C31H31ClN6O4 | 587.077 | 8 / 1 | 3.4 | No |
| 22057 | 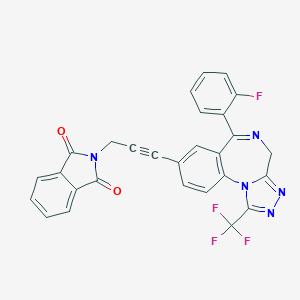 CHEMBL20194 CHEMBL20194 | C28H15F4N5O2 | 529.455 | 9 / 0 | 4.1 | No |
| 23472 | 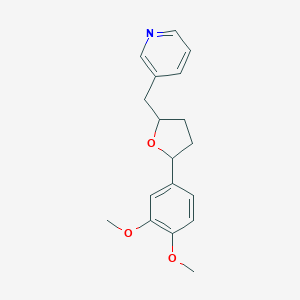 CHEMBL37643 CHEMBL37643 | C18H21NO3 | 299.37 | 4 / 0 | 3.0 | Yes |
| 24592 | 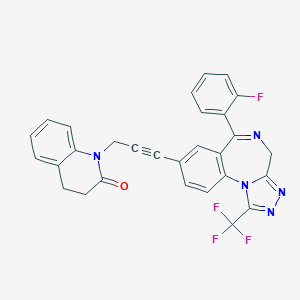 CHEMBL279084 CHEMBL279084 | C29H19F4N5O | 529.499 | 8 / 0 | 4.4 | No |
| 25647 | 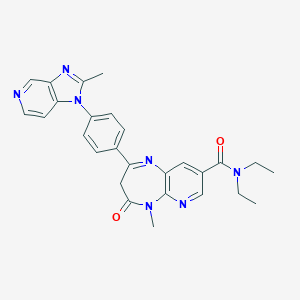 CHEMBL118672 CHEMBL118672 | C27H27N7O2 | 481.56 | 6 / 0 | 2.6 | Yes |
| 26361 | 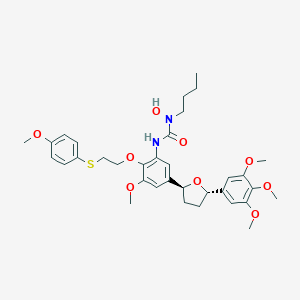 CHEMBL2113730 CHEMBL2113730 | C34H44N2O9S | 656.791 | 10 / 2 | 5.6 | No |
| 558118 | 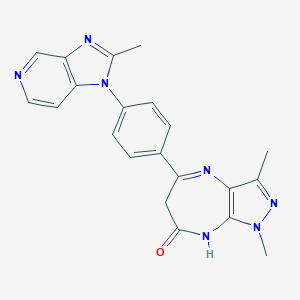 CHEMBL325928 CHEMBL325928 | C21H19N7O | 385.431 | 5 / 1 | 2.0 | Yes |
| 30787 | 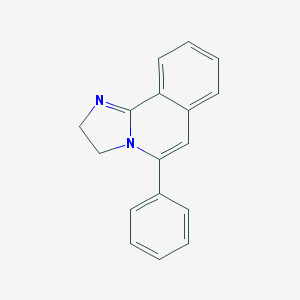 CHEMBL80130 CHEMBL80130 | C17H14N2 | 246.313 | 1 / 0 | 2.8 | Yes |
| 33427 | 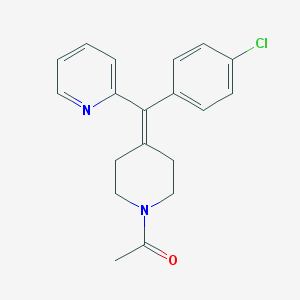 CHEMBL140919 CHEMBL140919 | C19H19ClN2O | 326.824 | 2 / 0 | 3.5 | Yes |
| 34732 | 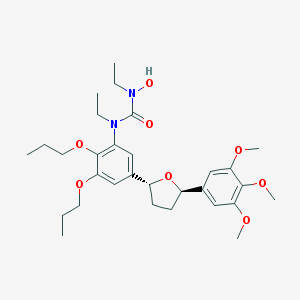 CHEMBL348609 CHEMBL348609 | C30H44N2O8 | 560.688 | 8 / 1 | 4.9 | No |
| 35124 | 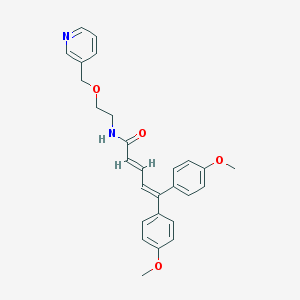 CHEMBL54068 CHEMBL54068 | C27H28N2O4 | 444.531 | 5 / 1 | 4.4 | Yes |
| 35292 | 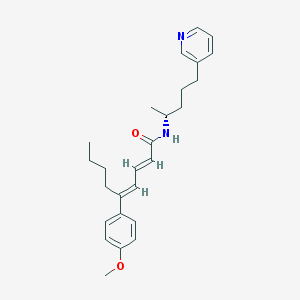 CHEMBL296600 CHEMBL296600 | C26H34N2O2 | 406.57 | 3 / 1 | 6.4 | No |
| 35540 | 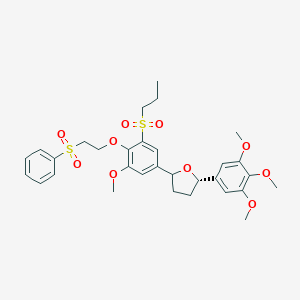 CHEMBL293773 CHEMBL293773 | C31H38O10S2 | 634.755 | 10 / 0 | 4.5 | No |
| 36825 | 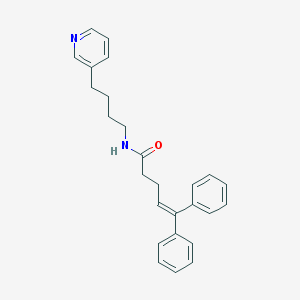 CHEMBL52821 CHEMBL52821 | C26H28N2O | 384.523 | 2 / 1 | 5.6 | No |
| 38835 | 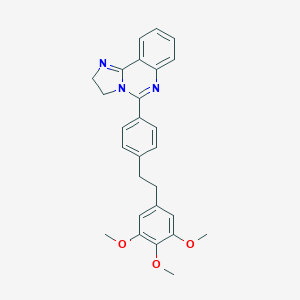 CHEMBL320871 CHEMBL320871 | C27H27N3O3 | 441.531 | 5 / 0 | 4.3 | Yes |
| 41279 | 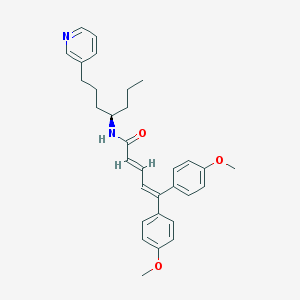 CHEMBL299773 CHEMBL299773 | C31H36N2O3 | 484.64 | 4 / 1 | 7.0 | No |
| 41782 | 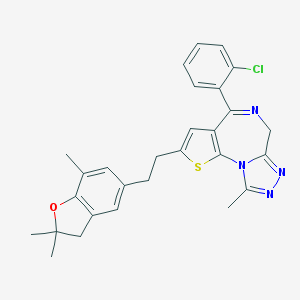 CHEMBL286337 CHEMBL286337 | C28H27ClN4OS | 503.061 | 5 / 0 | 5.0 | No |
| 42151 | 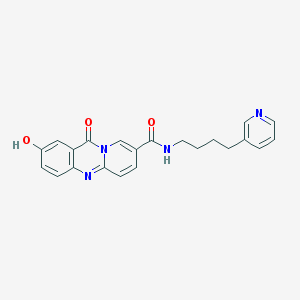 CHEMBL57480 CHEMBL57480 | C22H20N4O3 | 388.427 | 5 / 2 | 1.8 | Yes |
| 553454 | 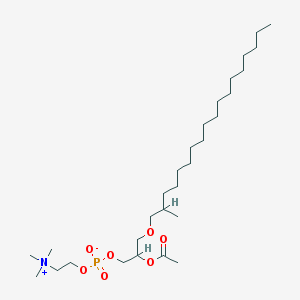 2-Methyloctadecyl paf 2-Methyloctadecyl paf | C29H60NO7P | 565.773 | 7 / 0 | 8.0 | No |
| 45376 | 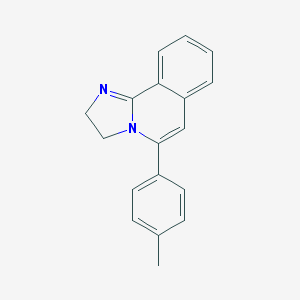 CHEMBL312650 CHEMBL312650 | C18H16N2 | 260.34 | 1 / 0 | 3.2 | Yes |
| 45482 | 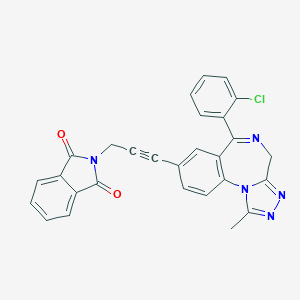 CHEMBL20033 CHEMBL20033 | C28H18ClN5O2 | 491.935 | 5 / 0 | 2.7 | Yes |
| 46213 | 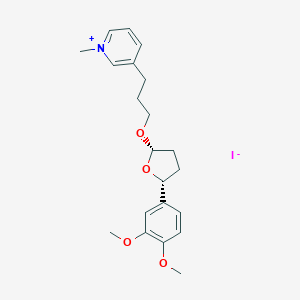 CHEMBL287821 CHEMBL287821 | C21H28INO4 | 485.362 | 5 / 0 | N/A | N/A |
| 46214 | 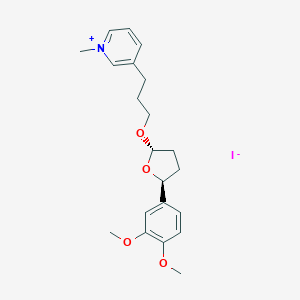 CHEMBL34063 CHEMBL34063 | C21H28INO4 | 485.362 | 5 / 0 | N/A | N/A |
| 522875 | 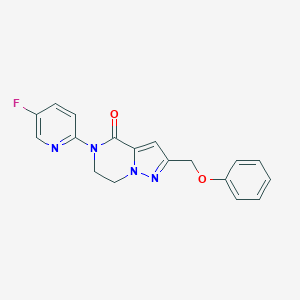 CHEMBL3746457 CHEMBL3746457 | C18H15FN4O2 | 338.342 | 5 / 0 | 2.2 | Yes |
| 47316 | 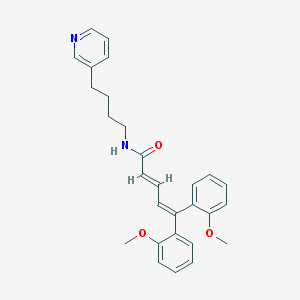 CHEMBL53804 CHEMBL53804 | C28H30N2O3 | 442.559 | 4 / 1 | 5.7 | No |
| 48624 | 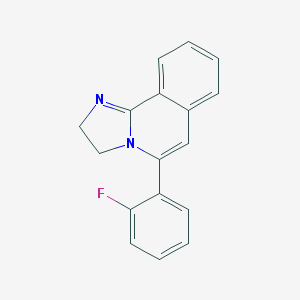 CHEMBL310238 CHEMBL310238 | C17H13FN2 | 264.303 | 2 / 0 | 2.9 | Yes |
| 49415 | 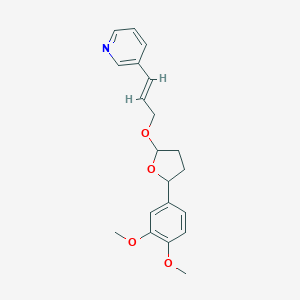 CHEMBL37952 CHEMBL37952 | C20H23NO4 | 341.407 | 5 / 0 | 3.1 | Yes |
| 50374 | 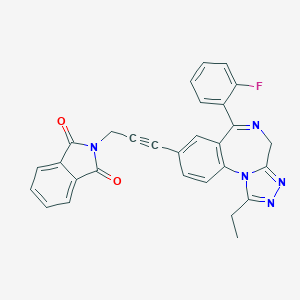 CHEMBL283152 CHEMBL283152 | C29H20FN5O2 | 489.51 | 6 / 0 | 4.0 | Yes |
| 50466 | 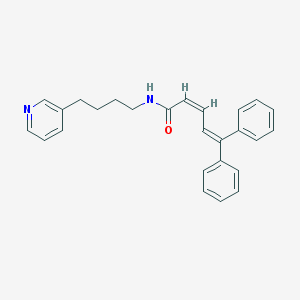 CHEMBL51696 CHEMBL51696 | C26H26N2O | 382.507 | 2 / 1 | 5.8 | No |
| 50763 | 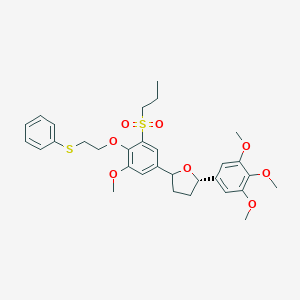 CHEMBL52369 CHEMBL52369 | C31H38O8S2 | 602.757 | 9 / 0 | 5.8 | No |
| 51196 | 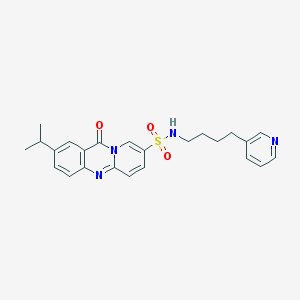 CHEMBL143037 CHEMBL143037 | C24H26N4O3S | 450.557 | 6 / 1 | 3.0 | Yes |
| 51896 | 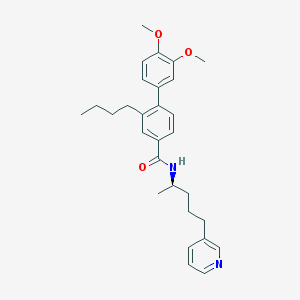 CHEMBL51692 CHEMBL51692 | C29H36N2O3 | 460.618 | 4 / 1 | 6.7 | No |
| 53960 | 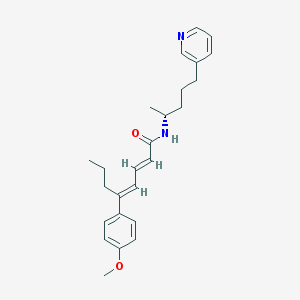 CHEMBL51855 CHEMBL51855 | C25H32N2O2 | 392.543 | 3 / 1 | 5.9 | No |
| 54574 | 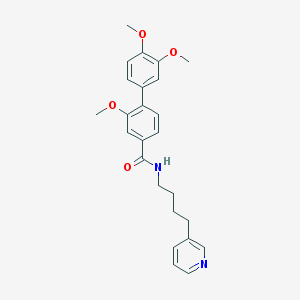 CHEMBL52031 CHEMBL52031 | C25H28N2O4 | 420.509 | 5 / 1 | 4.3 | Yes |
| 55559 | 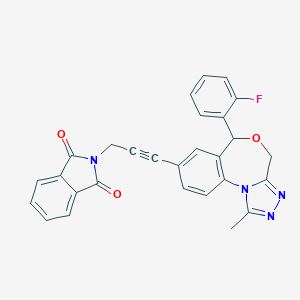 CHEMBL282666 CHEMBL282666 | C28H19FN4O3 | 478.483 | 6 / 0 | 3.4 | Yes |
| 55715 | 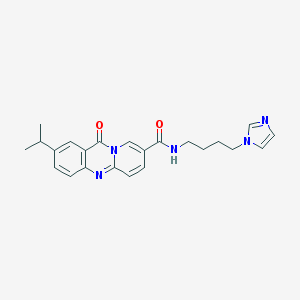 CHEMBL56434 CHEMBL56434 | C23H25N5O2 | 403.486 | 4 / 1 | 2.1 | Yes |
| 56526 | 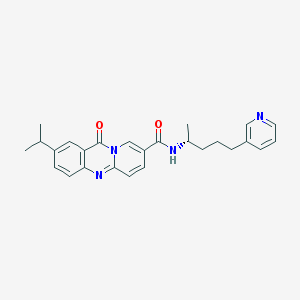 CHEMBL142433 CHEMBL142433 | C26H28N4O2 | 428.536 | 4 / 1 | 3.7 | Yes |
| 56527 | 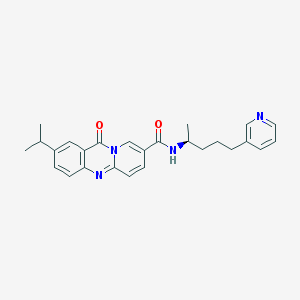 CHEMBL142903 CHEMBL142903 | C26H28N4O2 | 428.536 | 4 / 1 | 3.7 | Yes |
| 56528 | 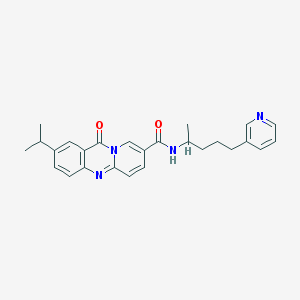 CHEMBL57825 CHEMBL57825 | C26H28N4O2 | 428.536 | 4 / 1 | 3.7 | Yes |
| 58309 | 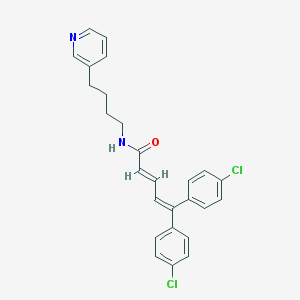 CHEMBL52615 CHEMBL52615 | C26H24Cl2N2O | 451.391 | 2 / 1 | 7.0 | No |
| 58658 | 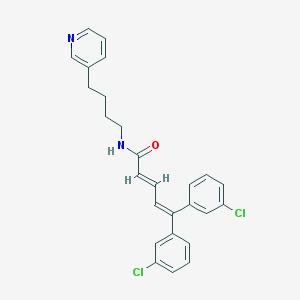 CHEMBL53149 CHEMBL53149 | C26H24Cl2N2O | 451.391 | 2 / 1 | 7.0 | No |
| 59740 | 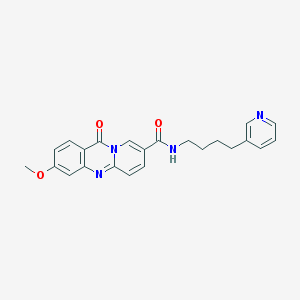 CHEMBL56418 CHEMBL56418 | C23H22N4O3 | 402.454 | 5 / 1 | 2.1 | Yes |
| 553543 | 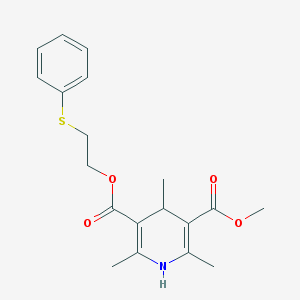 Pca 4248 Pca 4248 | C19H23NO4S | 361.456 | 6 / 1 | 3.6 | Yes |
| 59917 | 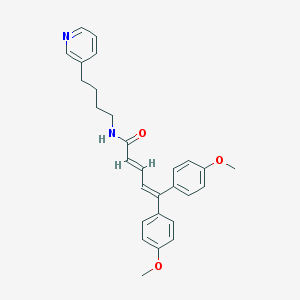 CHEMBL54029 CHEMBL54029 | C28H30N2O3 | 442.559 | 4 / 1 | 5.7 | No |
| 61829 | 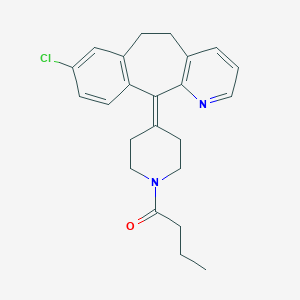 CHEMBL331603 CHEMBL331603 | C23H25ClN2O | 380.916 | 2 / 0 | 5.3 | No |
| 62639 | 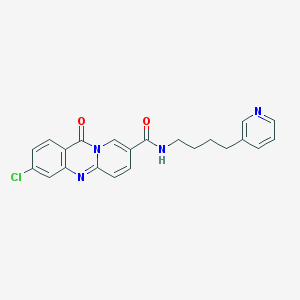 CHEMBL57489 CHEMBL57489 | C22H19ClN4O2 | 406.87 | 4 / 1 | 2.8 | Yes |
| 62658 | 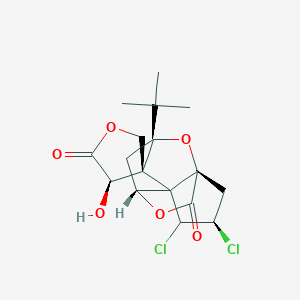 CHEMBL2304165 CHEMBL2304165 | C17H20Cl2O6 | 391.241 | 6 / 1 | 1.9 | Yes |
| 62659 | 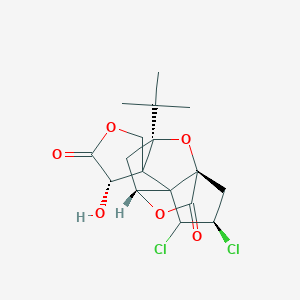 CHEMBL2368435 CHEMBL2368435 | C17H20Cl2O6 | 391.241 | 6 / 1 | 1.9 | Yes |
| 63648 | 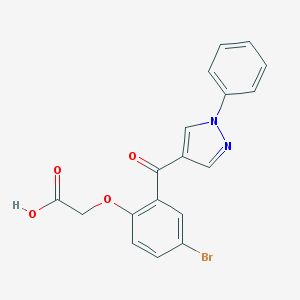 CHEMBL217053 CHEMBL217053 | C18H13BrN2O4 | 401.216 | 5 / 1 | 3.5 | Yes |
| 65724 | 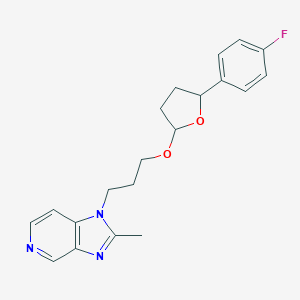 CHEMBL37358 CHEMBL37358 | C20H22FN3O2 | 355.413 | 5 / 0 | 3.0 | Yes |
| 67200 | 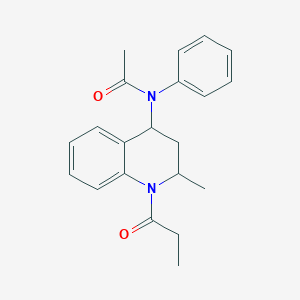 Probes1_000120 Probes1_000120 | C21H24N2O2 | 336.435 | 2 / 0 | 3.2 | Yes |
| 68849 | 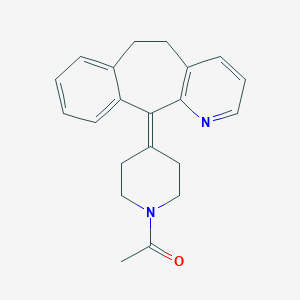 Azatadine M (nor), acetylated Azatadine M (nor), acetylated | C21H22N2O | 318.42 | 2 / 0 | 3.0 | Yes |
| 69932 | 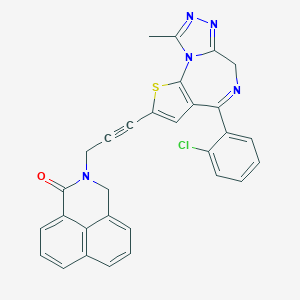 CHEMBL280210 CHEMBL280210 | C30H20ClN5OS | 534.034 | 5 / 0 | 5.5 | No |
| 72738 | 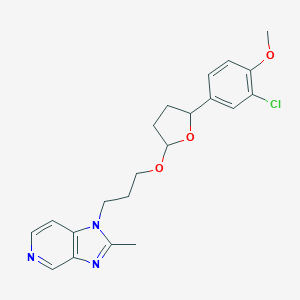 CHEMBL34548 CHEMBL34548 | C21H24ClN3O3 | 401.891 | 5 / 0 | 3.5 | Yes |
| 73130 | 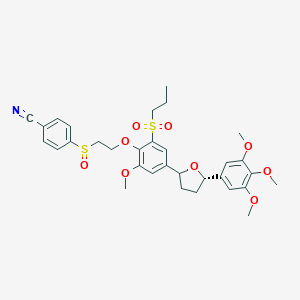 CHEMBL300910 CHEMBL300910 | C32H37NO9S2 | 643.766 | 11 / 0 | 4.1 | No |
| 74004 | 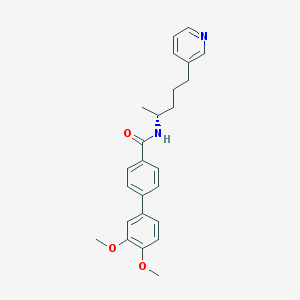 CHEMBL53585 CHEMBL53585 | C25H28N2O3 | 404.51 | 4 / 1 | 4.8 | Yes |
| 74840 | 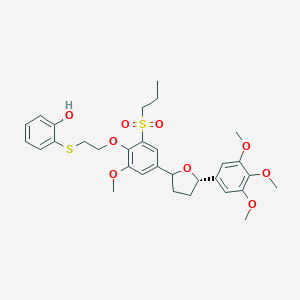 CHEMBL55071 CHEMBL55071 | C31H38O9S2 | 618.756 | 10 / 1 | 5.5 | No |
| 75141 | 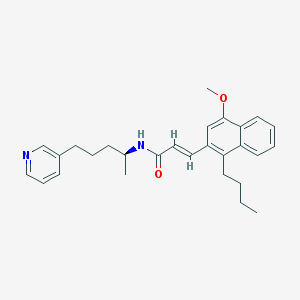 CHEMBL99294 CHEMBL99294 | C28H34N2O2 | 430.592 | 3 / 1 | 6.8 | No |
| 77699 | 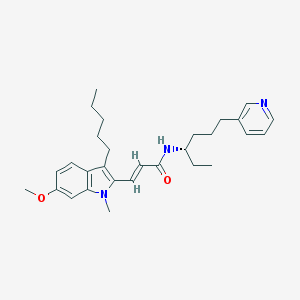 CHEMBL319384 CHEMBL319384 | C29H39N3O2 | 461.65 | 3 / 1 | 6.7 | No |
| 78440 | 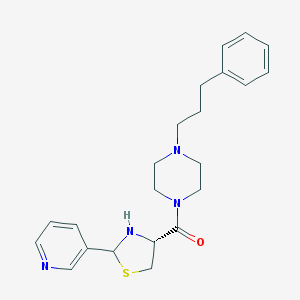 CHEMBL156461 CHEMBL156461 | C22H28N4OS | 396.553 | 5 / 1 | 2.5 | Yes |
| 79029 | 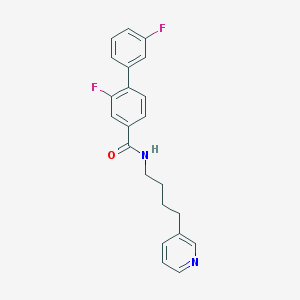 CHEMBL301388 CHEMBL301388 | C22H20F2N2O | 366.412 | 4 / 1 | 4.6 | Yes |
| 80758 | 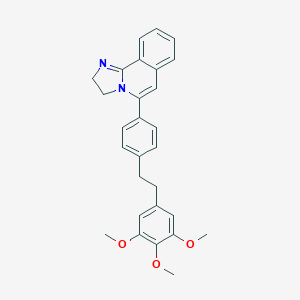 SDZ-64-412 SDZ-64-412 | C28H28N2O3 | 440.543 | 4 / 0 | 5.0 | Yes |
| 81067 | 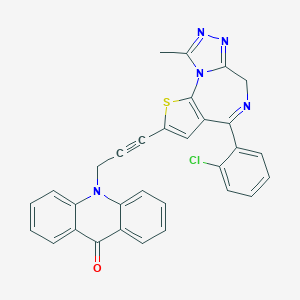 CHEMBL278635 CHEMBL278635 | C31H20ClN5OS | 546.045 | 6 / 0 | 4.9 | No |
| 81563 | 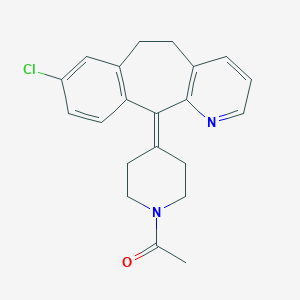 Sch-37370 Sch-37370 | C21H21ClN2O | 352.862 | 2 / 0 | 4.4 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218