

You can:
| Name | Corticotropin-releasing factor receptor 1 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | CRHR1 |
| Synonym | CRHR CRH-R1 CRH-R-1 CRFR1 CRFR-1 [ Show all ] |
| Disease | Major depressive disorder; Severe mood disorder Depression; Anxiety Depression Irritable bowel syndrome Anxiety disorder; Depression [ Show all ] |
| Length | 444 |
| Amino acid sequence | MGGHPQLRLVKALLLLGLNPVSASLQDQHCESLSLASNISGLQCNASVDLIGTCWPRSPAGQLVVRPCPAFFYGVRYNTTNNGYRECLANGSWAARVNYSECQEILNEEKKSKVHYHVAVIINYLGHCISLVALLVAFVLFLRLRPGCTHWGDQADGALEVGAPWSGAPFQVRRSIRCLRNIIHWNLISAFILRNATWFVVQLTMSPEVHQSNVGWCRLVTAAYNYFHVTNFFWMFGEGCYLHTAIVLTYSTDRLRKWMFICIGWGVPFPIIVAWAIGKLYYDNEKCWFGKRPGVYTDYIYQGPMILVLLINFIFLFNIVRILMTKLRASTTSETIQYRKAVKATLVLLPLLGITYMLFFVNPGEDEVSRVVFIYFNSFLESFQGFFVSVFYCFLNSEVRSAIRKRWHRWQDKHSIRARVARAMSIPTSPTRVSFHSIKQSTAV |
| UniProt | P34998 |
| Protein Data Bank | 4z9g, 4k5y |
| GPCR-HGmod model | P34998 |
| 3D structure model | This structure is from PDB ID 4z9g. |
| BioLiP | BL0350036,BL0350037,BL0350038, BL0251208 |
| Therapeutic Target Database | T45262 |
| ChEMBL | CHEMBL1800 |
| IUPHAR | 212 |
| DrugBank | BE0008658 |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 160 | 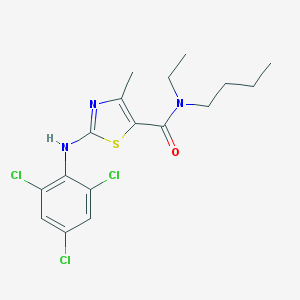 CHEMBL132055 CHEMBL132055 | C17H20Cl3N3OS | 420.777 | 4 / 1 | 6.5 | No |
| 583 | 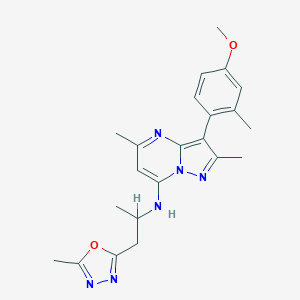 CHEMBL1287997 CHEMBL1287997 | C22H26N6O2 | 406.49 | 7 / 1 | 4.1 | Yes |
| 607 | 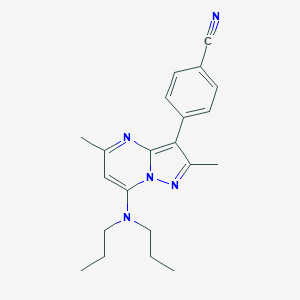 CHEMBL331539 CHEMBL331539 | C21H25N5 | 347.466 | 4 / 0 | 4.7 | Yes |
| 2293 | 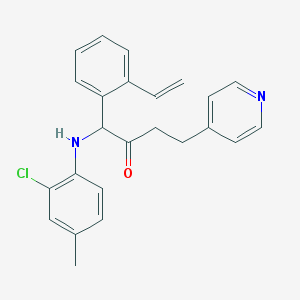 CHEMBL81376 CHEMBL81376 | C24H23ClN2O | 390.911 | 3 / 1 | 5.9 | No |
| 3033 | 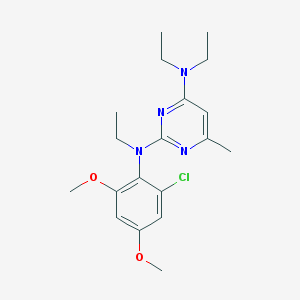 CHEMBL352378 CHEMBL352378 | C19H27ClN4O2 | 378.901 | 6 / 0 | 4.7 | Yes |
| 3088 | 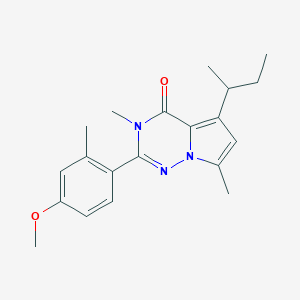 CHEMBL1939595 CHEMBL1939595 | C20H25N3O2 | 339.439 | 3 / 0 | 4.4 | Yes |
| 3542 | 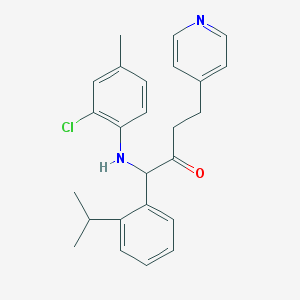 CHEMBL310343 CHEMBL310343 | C25H27ClN2O | 406.954 | 3 / 1 | 6.3 | No |
| 3704 | 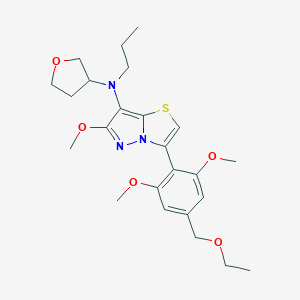 SCHEMBL1614189 SCHEMBL1614189 | C24H33N3O5S | 475.604 | 8 / 0 | 4.1 | Yes |
| 4074 | 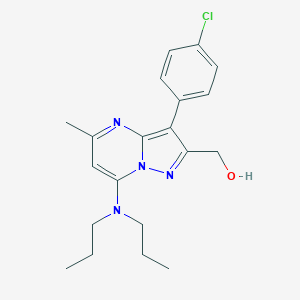 CHEMBL330954 CHEMBL330954 | C20H25ClN4O | 372.897 | 4 / 1 | 4.4 | Yes |
| 4171 | 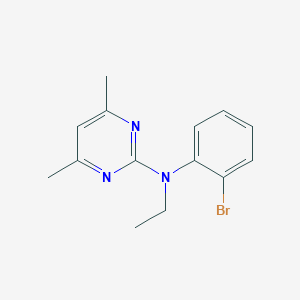 CHEMBL354871 CHEMBL354871 | C14H16BrN3 | 306.207 | 3 / 0 | 4.1 | Yes |
| 5127 | 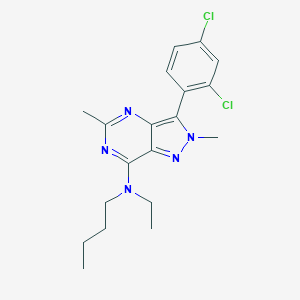 CHEMBL303380 CHEMBL303380 | C19H23Cl2N5 | 392.328 | 4 / 0 | 5.3 | No |
| 5492 | 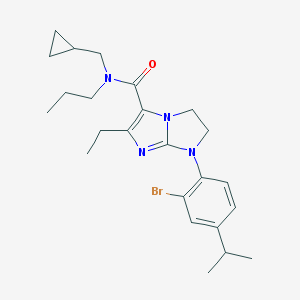 CHEMBL182624 CHEMBL182624 | C24H33BrN4O | 473.459 | 3 / 0 | 5.8 | No |
| 521630 | 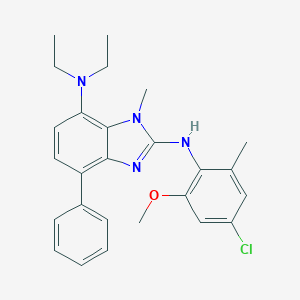 CHEMBL3793159 CHEMBL3793159 | C26H29ClN4O | 448.995 | 4 / 1 | 6.8 | No |
| 5834 | 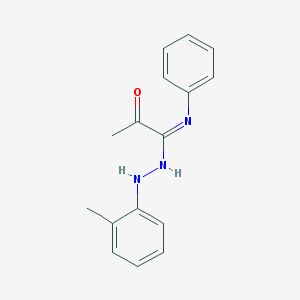 CHEMBL65708 CHEMBL65708 | C16H17N3O | 267.332 | 3 / 2 | 4.0 | Yes |
| 5898 | 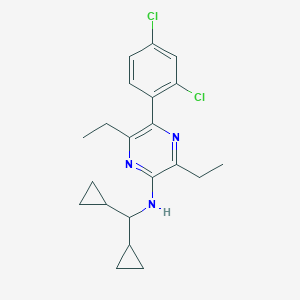 CHEMBL1807021 CHEMBL1807021 | C21H25Cl2N3 | 390.352 | 3 / 1 | 6.2 | No |
| 6306 | 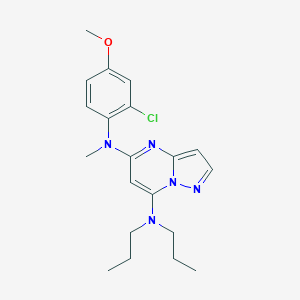 CHEMBL1828896 CHEMBL1828896 | C20H26ClN5O | 387.912 | 5 / 0 | 5.2 | No |
| 6571 | 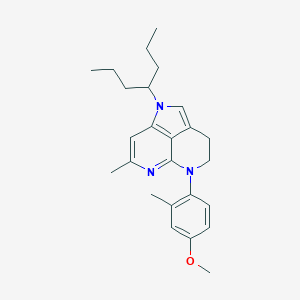 CHEMBL430396 CHEMBL430396 | C25H33N3O | 391.559 | 3 / 0 | 6.2 | No |
| 6575 | 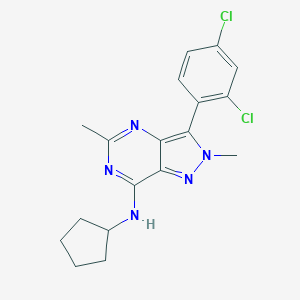 CHEMBL69968 CHEMBL69968 | C18H19Cl2N5 | 376.285 | 4 / 1 | 4.8 | Yes |
| 6910 | 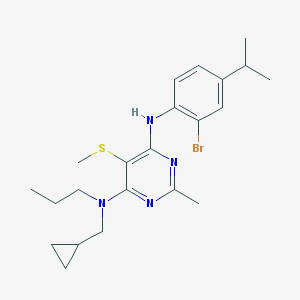 CHEMBL3235711 CHEMBL3235711 | C22H31BrN4S | 463.482 | 5 / 1 | 6.9 | No |
| 7248 | 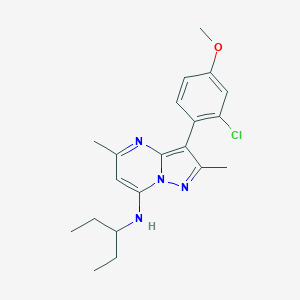 CHEMBL1819084 CHEMBL1819084 | C20H25ClN4O | 372.897 | 4 / 1 | 5.5 | No |
| 7398 | 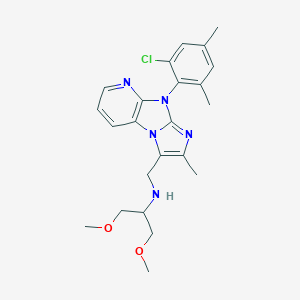 CHEMBL245506 CHEMBL245506 | C23H28ClN5O2 | 441.96 | 5 / 1 | 4.2 | Yes |
| 7503 | 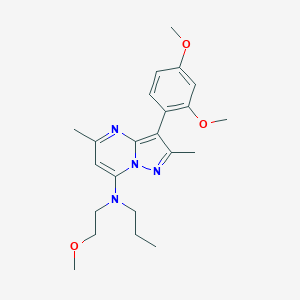 UNII-XBY7Y3DR1J UNII-XBY7Y3DR1J | C22H30N4O3 | 398.507 | 6 / 0 | 3.9 | Yes |
| 7625 | 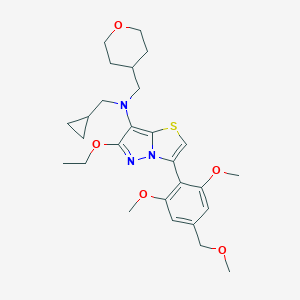 CHEMBL3664600 CHEMBL3664600 | C27H37N3O5S | 515.669 | 8 / 0 | 4.5 | No |
| 7635 | 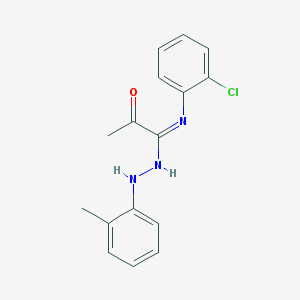 CHEMBL68119 CHEMBL68119 | C16H16ClN3O | 301.774 | 3 / 2 | 4.7 | Yes |
| 7663 | 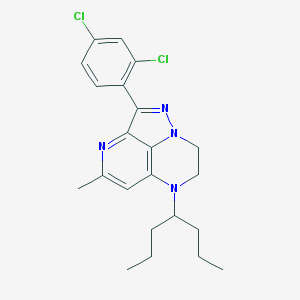 UNII-874OEY3JEJ UNII-874OEY3JEJ | C22H26Cl2N4 | 417.378 | 3 / 0 | 6.3 | No |
| 8173 | 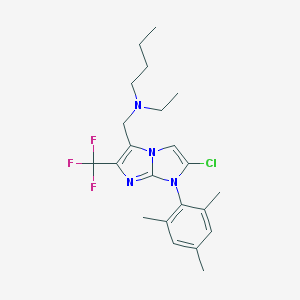 CHEMBL1086142 CHEMBL1086142 | C22H28ClF3N4 | 440.939 | 5 / 0 | 7.0 | No |
| 8930 | 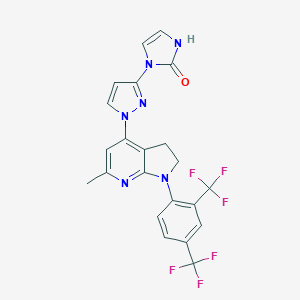 CHEMBL464185 CHEMBL464185 | C22H16F6N6O | 494.401 | 10 / 1 | 4.3 | Yes |
| 9131 | 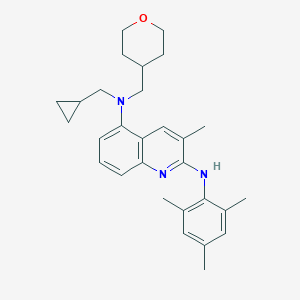 CHEMBL2049195 CHEMBL2049195 | C29H37N3O | 443.635 | 4 / 1 | 7.0 | No |
| 9183 | 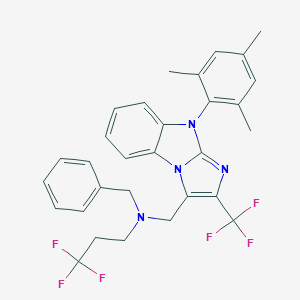 CHEMBL187975 CHEMBL187975 | C30H28F6N4 | 558.572 | 8 / 0 | 8.9 | No |
| 9383 | 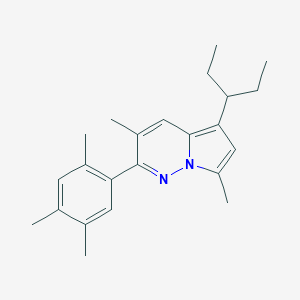 CHEMBL1939573 CHEMBL1939573 | C23H30N2 | 334.507 | 1 / 0 | 6.8 | No |
| 9446 | 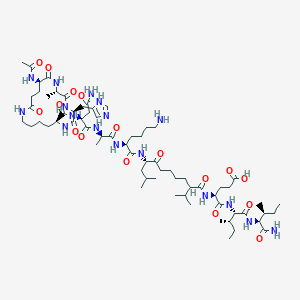 CHEMBL438724 CHEMBL438724 | C67H113N17O17 | 1428.74 | 19 / 17 | -2.5 | No |
| 9690 | 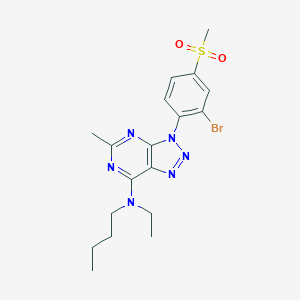 CHEMBL170844 CHEMBL170844 | C18H23BrN6O2S | 467.386 | 7 / 0 | 3.8 | Yes |
| 9713 | 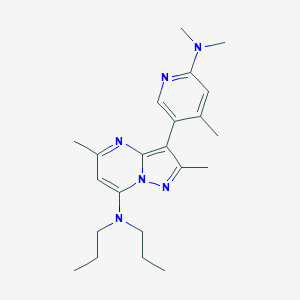 195055-03-9 195055-03-9 | C22H32N6 | 380.54 | 5 / 0 | 4.7 | Yes |
| 9724 | 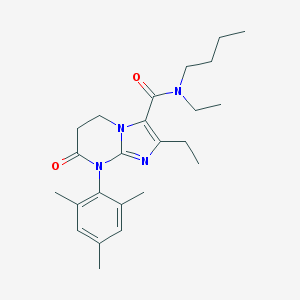 CHEMBL1077155 CHEMBL1077155 | C24H34N4O2 | 410.562 | 3 / 0 | 4.2 | Yes |
| 10039 | 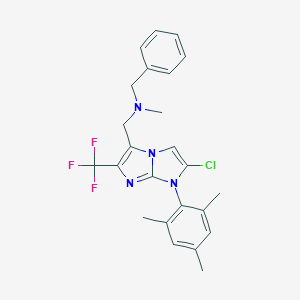 CHEMBL1082339 CHEMBL1082339 | C24H24ClF3N4 | 460.929 | 5 / 0 | 6.9 | No |
| 555518 |  OCRF OCRF | C205H339N59O63S | 4670.38 | 71 / 64 | -19.4 | No |
| 11111 | 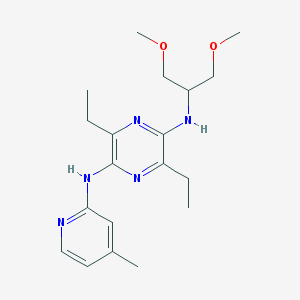 CHEMBL400861 CHEMBL400861 | C19H29N5O2 | 359.474 | 7 / 2 | 3.0 | Yes |
| 11472 |  CHEMBL2370927 CHEMBL2370927 | C183H307N47O56 | 4061.74 | 63 / 55 | -17.2 | No |
| 521774 | 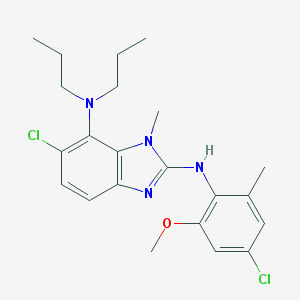 CHEMBL3793845 CHEMBL3793845 | C22H28Cl2N4O | 435.393 | 4 / 1 | 6.8 | No |
| 11557 | 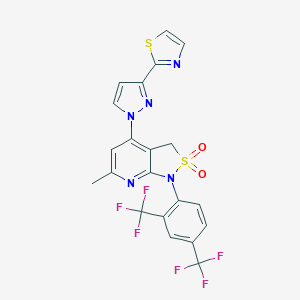 CHEMBL514717 CHEMBL514717 | C21H13F6N5O2S2 | 545.478 | 13 / 0 | 4.3 | No |
| 11567 | 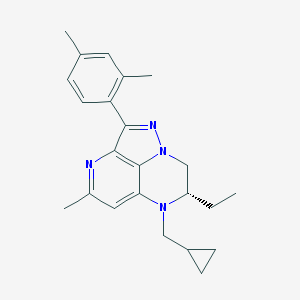 CHEMBL381850 CHEMBL381850 | C23H28N4 | 360.505 | 3 / 0 | 4.9 | Yes |
| 12166 | 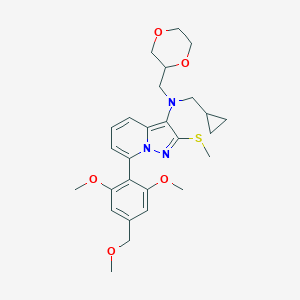 CHEMBL2087799 CHEMBL2087799 | C27H35N3O5S | 513.653 | 8 / 0 | 3.6 | No |
| 12378 | 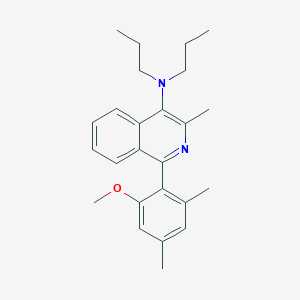 CHEMBL438888 CHEMBL438888 | C25H32N2O | 376.544 | 3 / 0 | 6.8 | No |
| 12556 | 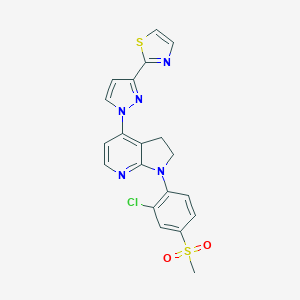 CHEMBL515070 CHEMBL515070 | C20H16ClN5O2S2 | 457.951 | 7 / 0 | 3.5 | Yes |
| 521826 | 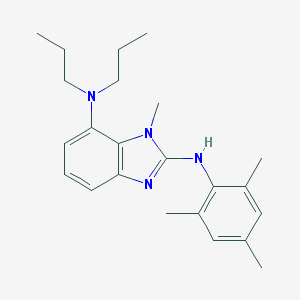 CHEMBL3794051 CHEMBL3794051 | C23H32N4 | 364.537 | 3 / 1 | 6.3 | No |
| 12883 | 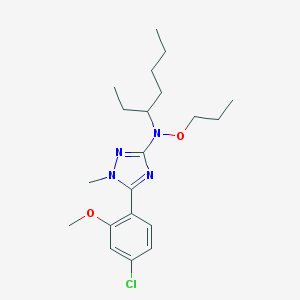 CHEMBL178667 CHEMBL178667 | C20H31ClN4O2 | 394.944 | 5 / 0 | 6.4 | No |
| 12894 | 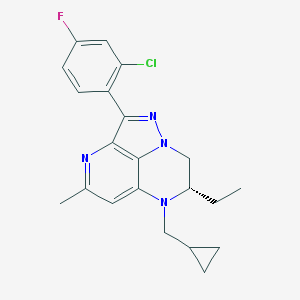 CHEMBL197187 CHEMBL197187 | C21H22ClFN4 | 384.883 | 4 / 0 | 4.9 | Yes |
| 13351 | 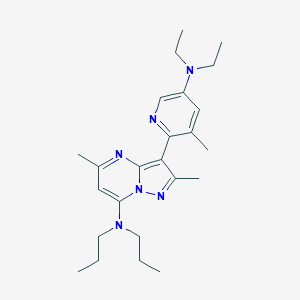 CHEMBL185055 CHEMBL185055 | C24H36N6 | 408.594 | 5 / 0 | 5.2 | No |
| 13409 | 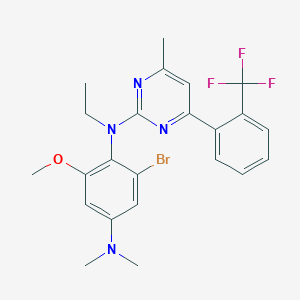 CHEMBL10015 CHEMBL10015 | C23H24BrF3N4O | 509.371 | 8 / 0 | 6.3 | No |
| 14099 | 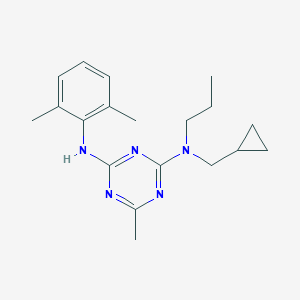 CHEMBL137123 CHEMBL137123 | C19H27N5 | 325.46 | 5 / 1 | 5.1 | No |
| 14427 | 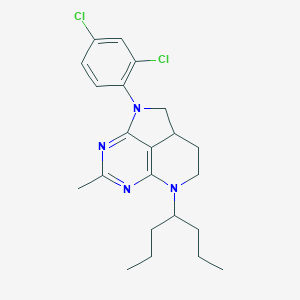 CHEMBL179608 CHEMBL179608 | C22H28Cl2N4 | 419.394 | 4 / 0 | 6.9 | No |
| 14456 | 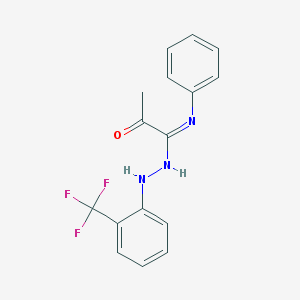 CHEMBL65721 CHEMBL65721 | C16H14F3N3O | 321.303 | 6 / 2 | 4.6 | Yes |
| 14540 | 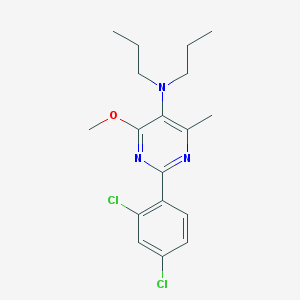 CHEMBL496270 CHEMBL496270 | C18H23Cl2N3O | 368.302 | 4 / 0 | 5.7 | No |
| 14615 | 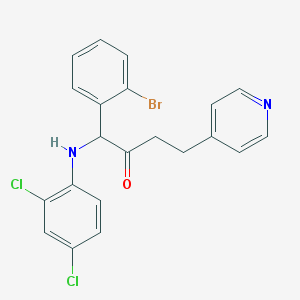 CHEMBL81726 CHEMBL81726 | C21H17BrCl2N2O | 464.184 | 3 / 1 | 6.1 | No |
| 14785 | 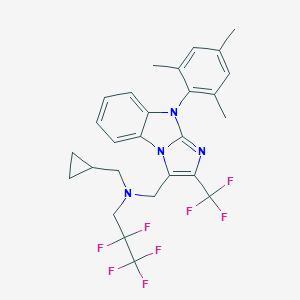 CHEMBL427527 CHEMBL427527 | C27H26F8N4 | 558.52 | 10 / 0 | 8.4 | No |
| 14838 | 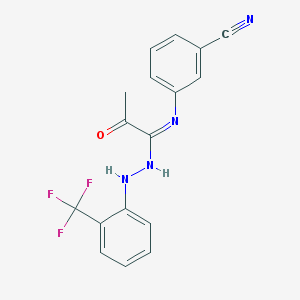 CHEMBL66572 CHEMBL66572 | C17H13F3N4O | 346.313 | 7 / 2 | 4.3 | Yes |
| 15310 | 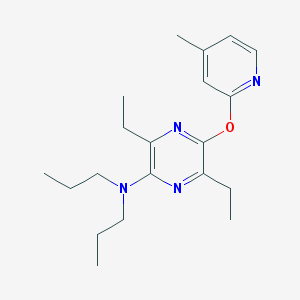 CHEMBL400154 CHEMBL400154 | C20H30N4O | 342.487 | 5 / 0 | 5.2 | No |
| 15335 | 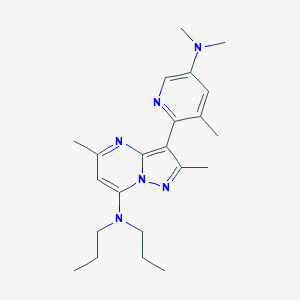 CHEMBL184850 CHEMBL184850 | C22H32N6 | 380.54 | 5 / 0 | 4.5 | Yes |
| 15577 | 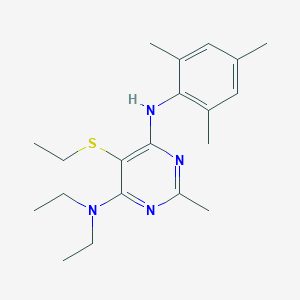 CHEMBL3235704 CHEMBL3235704 | C20H30N4S | 358.548 | 5 / 1 | 5.6 | No |
| 15820 | 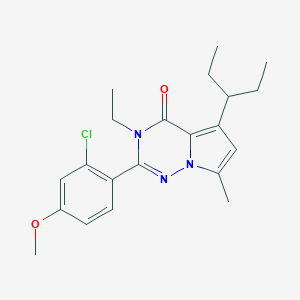 CHEMBL1939583 CHEMBL1939583 | C21H26ClN3O2 | 387.908 | 3 / 0 | 5.6 | No |
| 15913 | 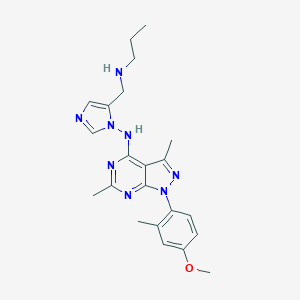 CHEMBL567382 CHEMBL567382 | C22H28N8O | 420.521 | 7 / 2 | 4.2 | Yes |
| 15981 | 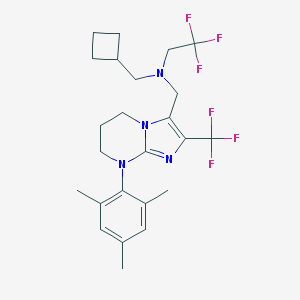 CHEMBL1088624 CHEMBL1088624 | C24H30F6N4 | 488.522 | 9 / 0 | 6.4 | No |
| 16227 | 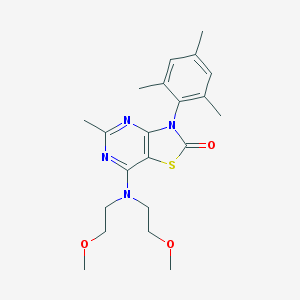 CHEMBL17656 CHEMBL17656 | C21H28N4O3S | 416.54 | 7 / 0 | 3.7 | Yes |
| 16371 | 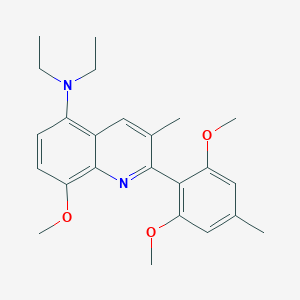 CHEMBL2160159 CHEMBL2160159 | C24H30N2O3 | 394.515 | 5 / 0 | 5.4 | No |
| 16430 | 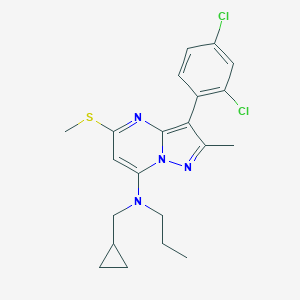 CHEMBL333735 CHEMBL333735 | C21H24Cl2N4S | 435.411 | 4 / 0 | 6.5 | No |
| 16583 | 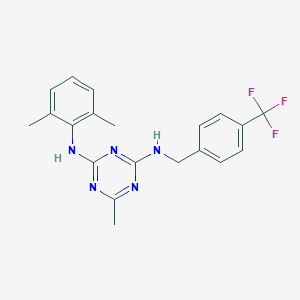 CHEMBL138597 CHEMBL138597 | C20H20F3N5 | 387.41 | 8 / 2 | 5.7 | No |
| 16934 | 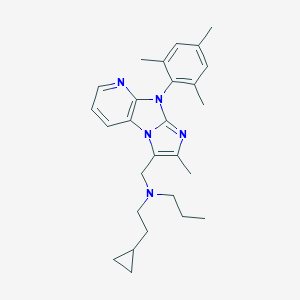 CHEMBL391170 CHEMBL391170 | C27H35N5 | 429.612 | 3 / 0 | 7.0 | No |
| 17005 | 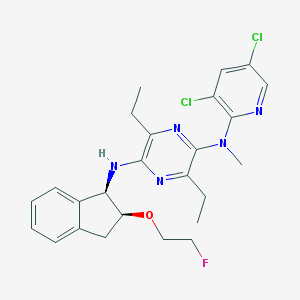 CHEMBL249212 CHEMBL249212 | C25H28Cl2FN5O | 504.431 | 7 / 1 | 6.1 | No |
| 17028 | 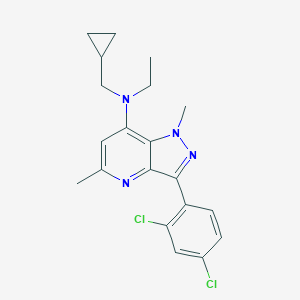 CHEMBL332721 CHEMBL332721 | C20H22Cl2N4 | 389.324 | 3 / 0 | 5.1 | No |
| 17157 | 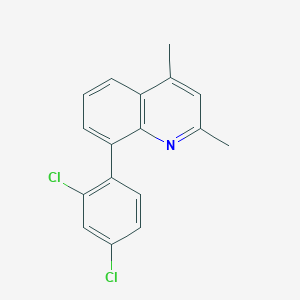 CHEMBL112389 CHEMBL112389 | C17H13Cl2N | 302.198 | 1 / 0 | 5.8 | No |
| 17193 | 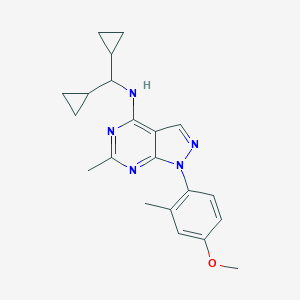 CHEMBL1836950 CHEMBL1836950 | C21H25N5O | 363.465 | 5 / 1 | 4.5 | Yes |
| 17414 | 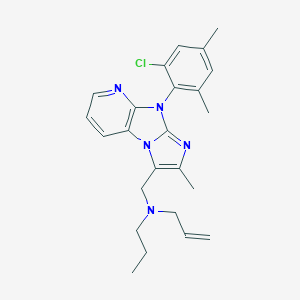 CHEMBL247151 CHEMBL247151 | C24H28ClN5 | 421.973 | 3 / 0 | 6.4 | No |
| 17490 | 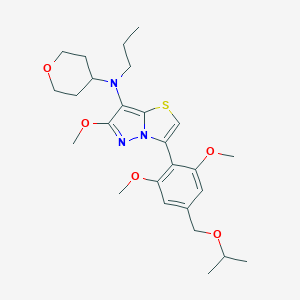 SCHEMBL1613428 SCHEMBL1613428 | C26H37N3O5S | 503.658 | 8 / 0 | 4.9 | No |
| 17669 | 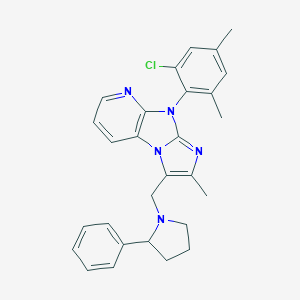 CHEMBL394112 CHEMBL394112 | C28H28ClN5 | 470.017 | 3 / 0 | 6.9 | No |
| 17732 | 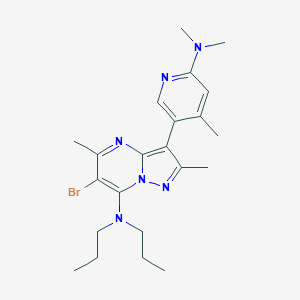 CHEMBL364168 CHEMBL364168 | C22H31BrN6 | 459.436 | 5 / 0 | 5.4 | No |
| 17800 | 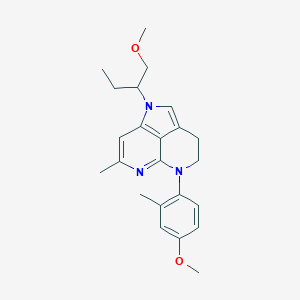 CHEMBL249333 CHEMBL249333 | C23H29N3O2 | 379.504 | 4 / 0 | 4.4 | Yes |
| 17839 | 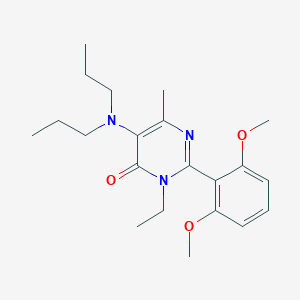 CHEMBL83789 CHEMBL83789 | C21H31N3O3 | 373.497 | 5 / 0 | 3.9 | Yes |
| 17951 | 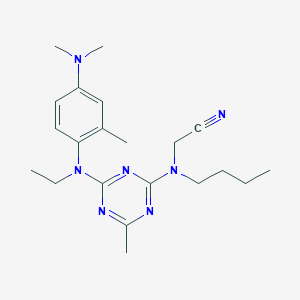 CHEMBL442773 CHEMBL442773 | C21H31N7 | 381.528 | 7 / 0 | 4.7 | Yes |
| 18275 | 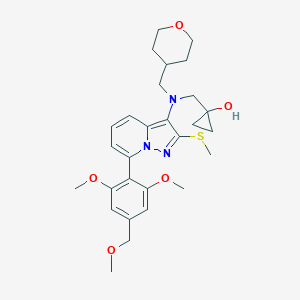 CHEMBL2087797 CHEMBL2087797 | C28H37N3O5S | 527.68 | 8 / 1 | 3.6 | No |
| 18373 | 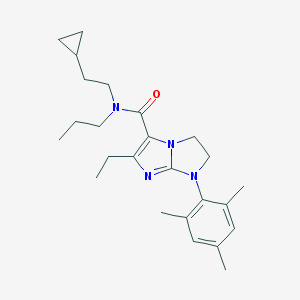 CHEMBL179499 CHEMBL179499 | C25H36N4O | 408.59 | 3 / 0 | 5.8 | No |
| 18447 | 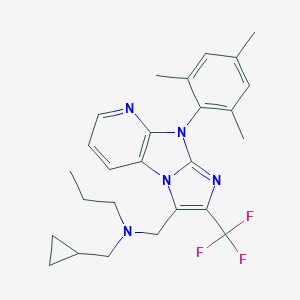 CHEMBL245671 CHEMBL245671 | C26H30F3N5 | 469.556 | 6 / 0 | 6.8 | No |
| 18685 | 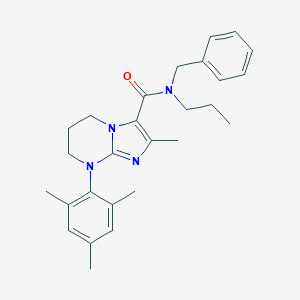 CHEMBL1083822 CHEMBL1083822 | C27H34N4O | 430.596 | 3 / 0 | 5.8 | No |
| 19154 | 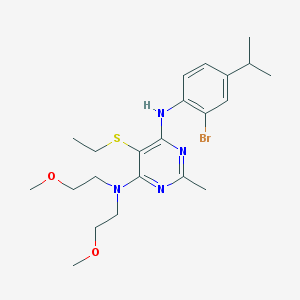 CHEMBL3235715 CHEMBL3235715 | C22H33BrN4O2S | 497.496 | 7 / 1 | 5.3 | No |
| 19261 | 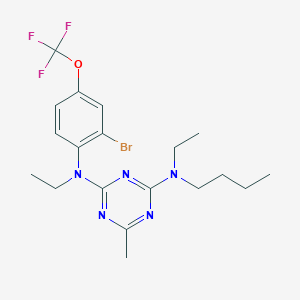 CHEMBL354367 CHEMBL354367 | C19H25BrF3N5O | 476.342 | 9 / 0 | 6.7 | No |
| 19279 | 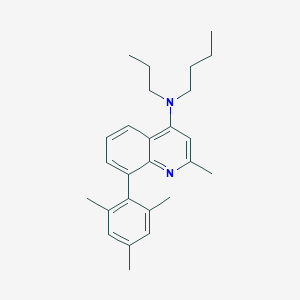 CHEMBL113653 CHEMBL113653 | C26H34N2 | 374.572 | 2 / 0 | 7.6 | No |
| 19289 | 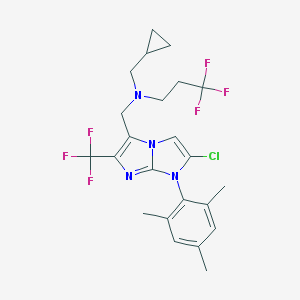 CHEMBL1083320 CHEMBL1083320 | C23H25ClF6N4 | 506.921 | 8 / 0 | 7.6 | No |
| 19337 | 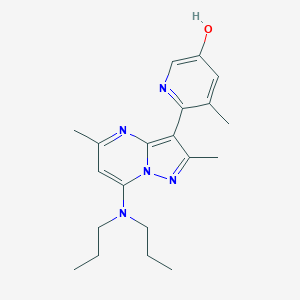 CHEMBL181860 CHEMBL181860 | C20H27N5O | 353.47 | 5 / 1 | 4.0 | Yes |
| 19490 | 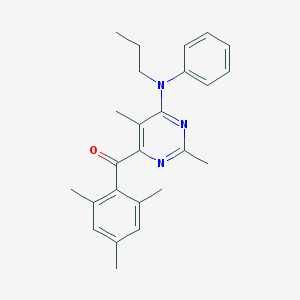 CHEMBL362224 CHEMBL362224 | C25H29N3O | 387.527 | 4 / 0 | 6.6 | No |
| 19630 |  CHEMBL2370916 CHEMBL2370916 | C211H362N58O66 | 4767.56 | 73 / 64 | -23.3 | No |
| 19646 | 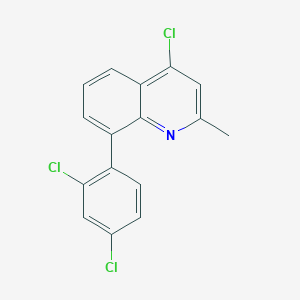 CHEMBL115119 CHEMBL115119 | C16H10Cl3N | 322.613 | 1 / 0 | 6.1 | No |
| 19865 | 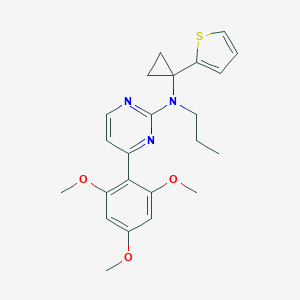 CHEMBL302396 CHEMBL302396 | C23H27N3O3S | 425.547 | 7 / 0 | 4.7 | Yes |
| 20013 | 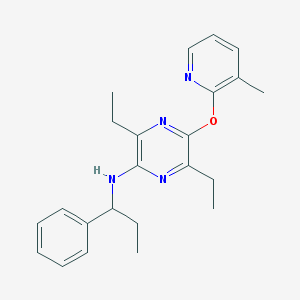 CHEMBL248816 CHEMBL248816 | C23H28N4O | 376.504 | 5 / 1 | 5.7 | No |
| 20452 | 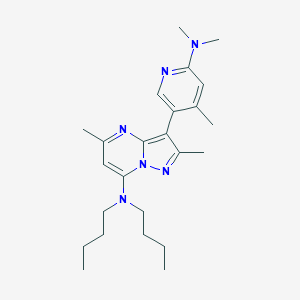 CHEMBL360851 CHEMBL360851 | C24H36N6 | 408.594 | 5 / 0 | 5.5 | No |
| 20997 | 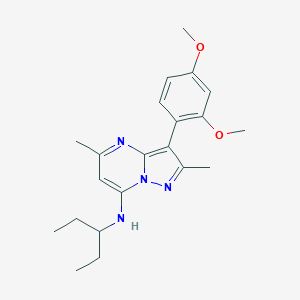 CHEMBL334237 CHEMBL334237 | C21H28N4O2 | 368.481 | 5 / 1 | 4.9 | Yes |
| 20999 | 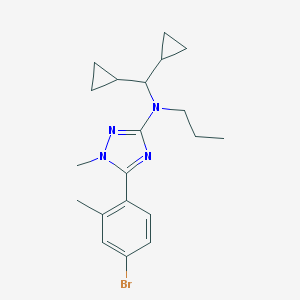 CHEMBL108673 CHEMBL108673 | C20H27BrN4 | 403.368 | 3 / 0 | 5.6 | No |
| 21467 | 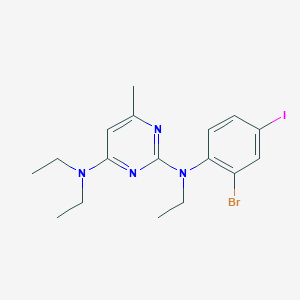 CHEMBL353973 CHEMBL353973 | C17H22BrIN4 | 489.199 | 4 / 0 | 5.5 | No |
| 21524 | 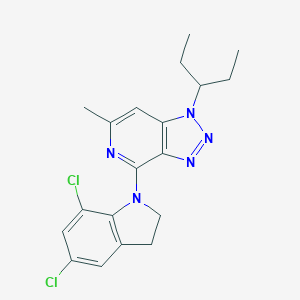 CHEMBL297868 CHEMBL297868 | C19H21Cl2N5 | 390.312 | 4 / 0 | 5.5 | No |
| 21644 | 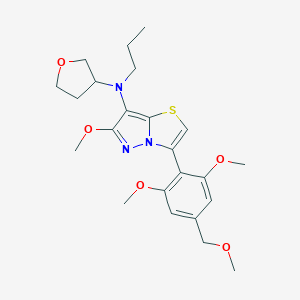 SCHEMBL1614465 SCHEMBL1614465 | C23H31N3O5S | 461.577 | 8 / 0 | 3.7 | Yes |
| 21663 | 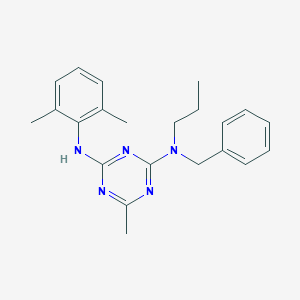 CHEMBL136596 CHEMBL136596 | C22H27N5 | 361.493 | 5 / 1 | 5.8 | No |
| 21749 | 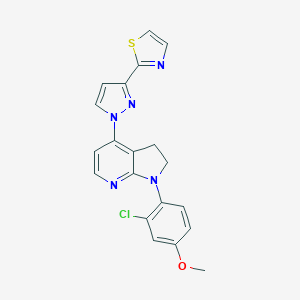 CHEMBL462789 CHEMBL462789 | C20H16ClN5OS | 409.892 | 6 / 0 | 4.2 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218