

You can:
| Name | P2Y purinoceptor 4 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | P2RY4 |
| Synonym | UNR pyrimidinoceptor pyrimidinergic receptor P2Y P2Y4R P2Y4 receptor [ Show all ] |
| Disease | N/A |
| Length | 365 |
| Amino acid sequence | MASTESSLLRSLGLSPGPGSSEVELDCWFDEDFKFILLPVSYAVVFVLGLGLNAPTLWLFIFRLRPWDATATYMFHLALSDTLYVLSLPTLIYYYAAHNHWPFGTEICKFVRFLFYWNLYCSVLFLTCISVHRYLGICHPLRALRWGRPRLAGLLCLAVWLVVAGCLVPNLFFVTTSNKGTTVLCHDTTRPEEFDHYVHFSSAVMGLLFGVPCLVTLVCYGLMARRLYQPLPGSAQSSSRLRSLRTIAVVLTVFAVCFVPFHITRTIYYLARLLEADCRVLNIVNVVYKVTRPLASANSCLDPVLYLLTGDKYRRQLRQLCGGGKPQPRTAASSLALVSLPEDSSCRWAATPQDSSCSTPRADRL |
| UniProt | P51582 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | P51582 |
| 3D structure model | This predicted structure model is from GPCR-EXP P51582. |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL2123 |
| IUPHAR | 325 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 3493 | 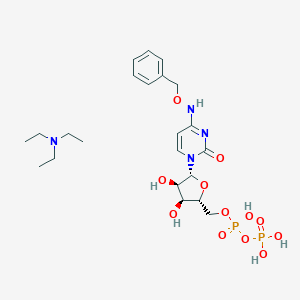 CHEMBL1084612 CHEMBL1084612 | C22H36N4O12P2 | 610.494 | 13 / 6 | N/A | No |
| 3879 | 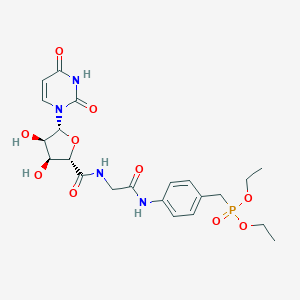 PSB-6426 PSB-6426 | C22H29N4O10P | 540.466 | 10 / 5 | -1.6 | No |
| 4532 | 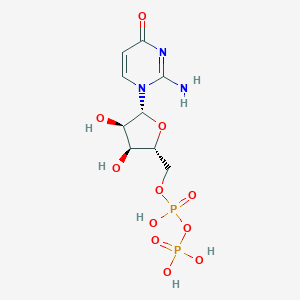 CHEMBL1767415 CHEMBL1767415 | C9H15N3O11P2 | 403.177 | 11 / 6 | -4.5 | No |
| 4767 | 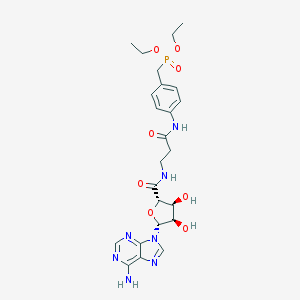 CHEMBL517017 CHEMBL517017 | C24H32N7O8P | 577.535 | 12 / 5 | -1.3 | No |
| 6219 | 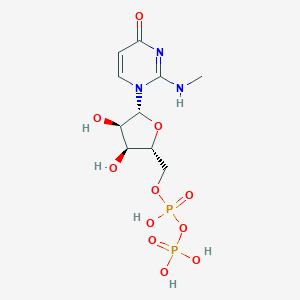 CHEMBL1767416 CHEMBL1767416 | C10H17N3O11P2 | 417.204 | 11 / 6 | -4.1 | No |
| 7015 | 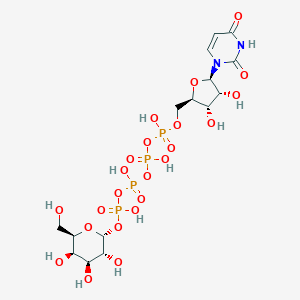 CHEMBL444212 CHEMBL444212 | C15H26N2O23P4 | 726.259 | 23 / 11 | -8.5 | No |
| 7019 | 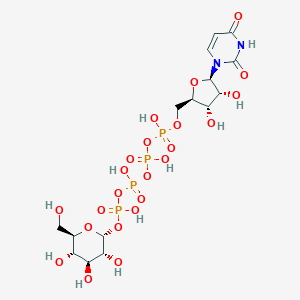 CHEMBL499138 CHEMBL499138 | C15H26N2O23P4 | 726.259 | 23 / 11 | -8.5 | No |
| 7020 | 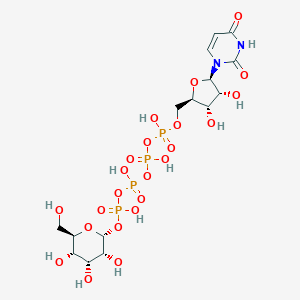 CHEMBL1784885 CHEMBL1784885 | C15H26N2O23P4 | 726.259 | 23 / 11 | -8.5 | No |
| 7024 | 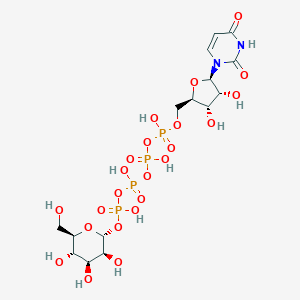 CHEMBL1784886 CHEMBL1784886 | C15H26N2O23P4 | 726.259 | 23 / 11 | -8.5 | No |
| 12386 | 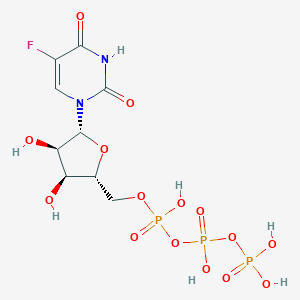 FUTP FUTP | C9H14FN2O15P3 | 502.13 | 16 / 7 | -5.8 | No |
| 12612 | 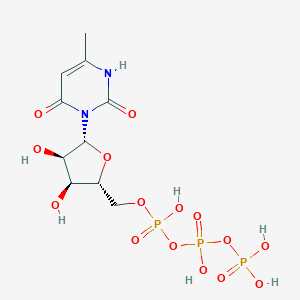 CHEMBL413841 CHEMBL413841 | C10H17N2O15P3 | 498.166 | 15 / 7 | -5.8 | No |
| 16610 | 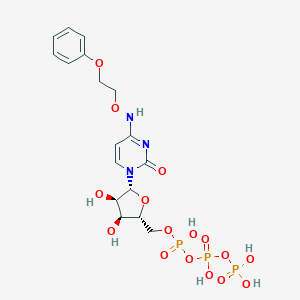 CHEMBL3261361 CHEMBL3261361 | C17H24N3O16P3 | 619.305 | 16 / 7 | -3.3 | No |
| 17592 | 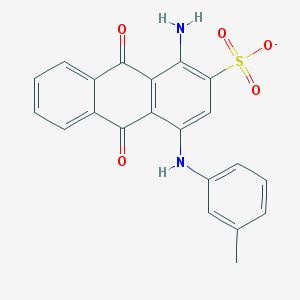 CHEMBL445413 CHEMBL445413 | C21H15N2O5S- | 407.42 | 7 / 2 | 3.9 | Yes |
| 18300 | 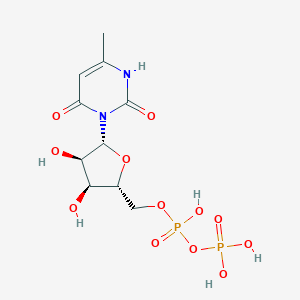 CHEMBL386096 CHEMBL386096 | C10H16N2O12P2 | 418.188 | 12 / 6 | -4.7 | No |
| 18896 | 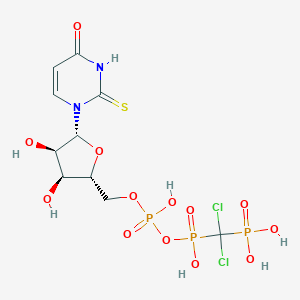 CHEMBL519835 CHEMBL519835 | C10H15Cl2N2O13P3S | 567.112 | 14 / 7 | -4.4 | No |
| 19508 | 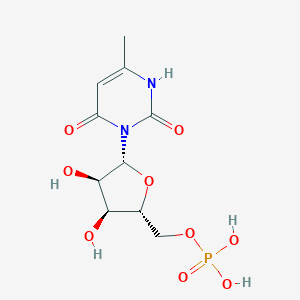 CHEMBL215263 CHEMBL215263 | C10H15N2O9P | 338.209 | 9 / 5 | -3.6 | Yes |
| 21440 | 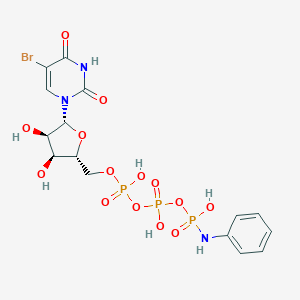 CHEMBL1765120 CHEMBL1765120 | C15H19BrN3O14P3 | 638.149 | 15 / 7 | -2.3 | No |
| 22713 | 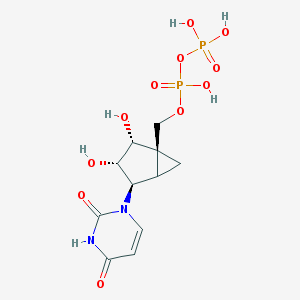 CHEMBL215811 CHEMBL215811 | C11H16N2O11P2 | 414.2 | 11 / 6 | -4.6 | No |
| 26413 | 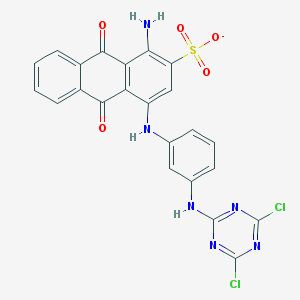 CHEMBL1672105 CHEMBL1672105 | C23H13Cl2N6O5S- | 556.354 | 11 / 3 | 5.3 | No |
| 26845 | 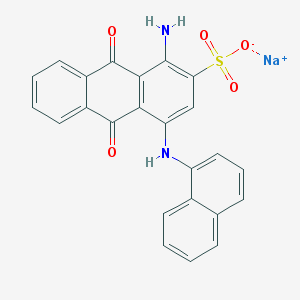 PSB 06126 PSB 06126 | C24H15N2NaO5S | 466.443 | 7 / 2 | N/A | N/A |
| 27552 | 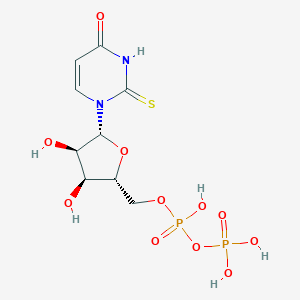 2-thio-UDP 2-thio-UDP | C9H14N2O11P2S | 420.222 | 12 / 6 | -4.1 | No |
| 30316 | 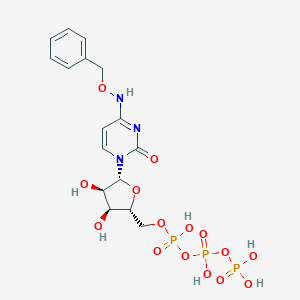 CHEMBL1784892 CHEMBL1784892 | C16H22N3O15P3 | 589.279 | 15 / 7 | -3.7 | No |
| 32086 | 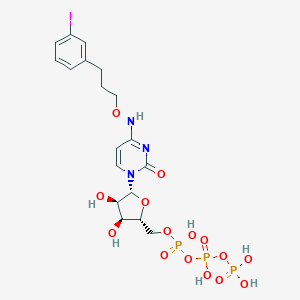 CHEMBL3261365 CHEMBL3261365 | C18H25IN3O15P3 | 743.23 | 15 / 7 | -3.1 | No |
| 32487 | 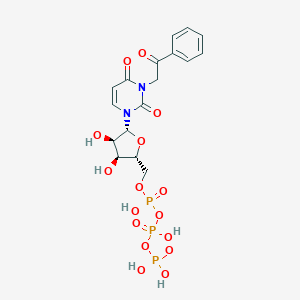 CHEMBL385488 CHEMBL385488 | C17H21N2O16P3 | 602.274 | 16 / 6 | -4.5 | No |
| 33912 | 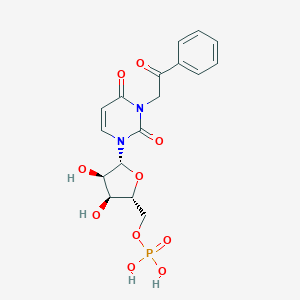 CHEMBL215381 CHEMBL215381 | C17H19N2O10P | 442.317 | 10 / 4 | -2.4 | Yes |
| 34088 | 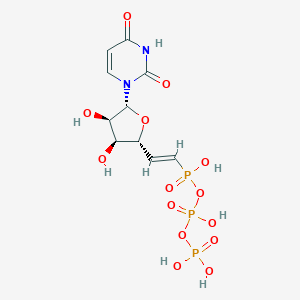 CHEMBL523850 CHEMBL523850 | C10H15N2O14P3 | 480.151 | 14 / 7 | -6.1 | No |
| 38814 | 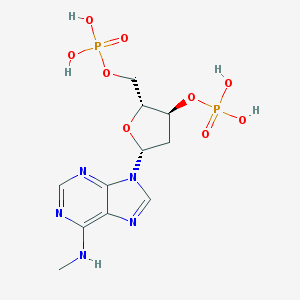 CHEMBL129841 CHEMBL129841 | C11H17N5O9P2 | 425.231 | 13 / 5 | -2.9 | No |
| 42141 | 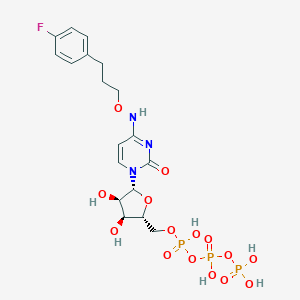 CHEMBL3261363 CHEMBL3261363 | C18H25FN3O15P3 | 635.324 | 16 / 7 | -3.6 | No |
| 42290 | 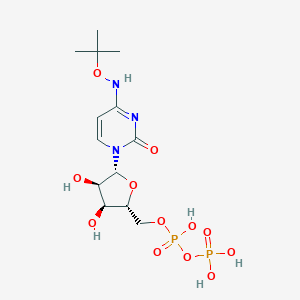 CHEMBL1084296 CHEMBL1084296 | C13H23N3O12P2 | 475.284 | 12 / 6 | -3.1 | No |
| 42487 | 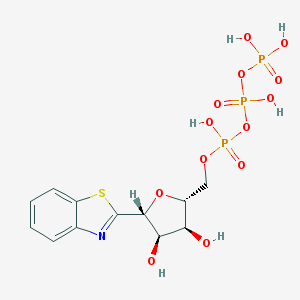 CHEMBL241481 CHEMBL241481 | C12H16NO13P3S | 507.235 | 15 / 6 | -3.3 | No |
| 44384 | 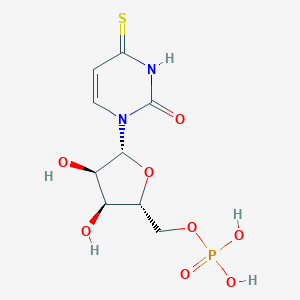 CHEMBL1230331 CHEMBL1230331 | C9H13N2O8PS | 340.243 | 9 / 5 | -3.0 | Yes |
| 47985 | 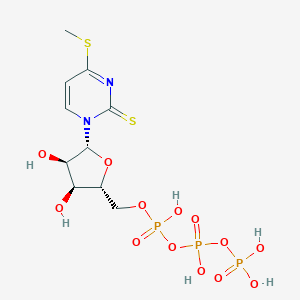 CHEMBL482472 CHEMBL482472 | C10H17N2O13P3S2 | 530.288 | 15 / 6 | -4.3 | No |
| 49427 | 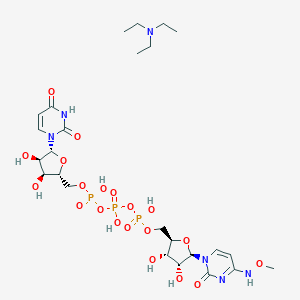 CHEMBL1083256 CHEMBL1083256 | C25H43N6O20P3 | 840.562 | 21 / 9 | N/A | No |
| 49773 | 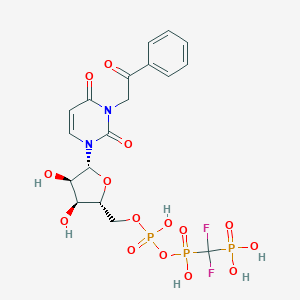 CHEMBL1765118 CHEMBL1765118 | C18H21F2N2O15P3 | 636.283 | 17 / 6 | -3.9 | No |
| 53515 | 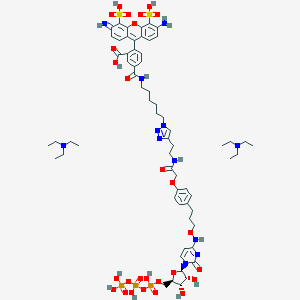 CHEMBL3261377 CHEMBL3261377 | C63H89N12O27P3S2 | 1603.5 | 33 / 14 | N/A | No |
| 53631 | 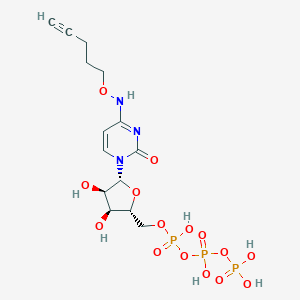 CHEMBL3261373 CHEMBL3261373 | C14H22N3O15P3 | 565.257 | 15 / 7 | -4.2 | No |
| 58254 | 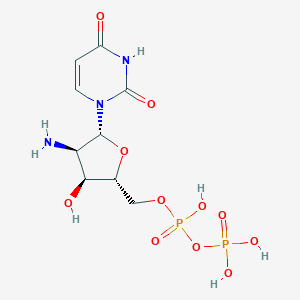 CHEMBL215911 CHEMBL215911 | C9H15N3O11P2 | 403.177 | 12 / 6 | -7.3 | No |
| 58527 | 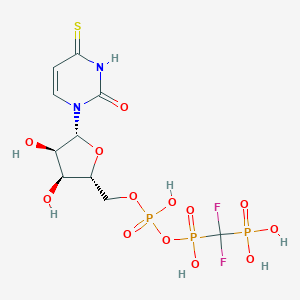 CHEMBL1765119 CHEMBL1765119 | C10H15F2N2O13P3S | 534.209 | 16 / 7 | -4.6 | No |
| 60007 | 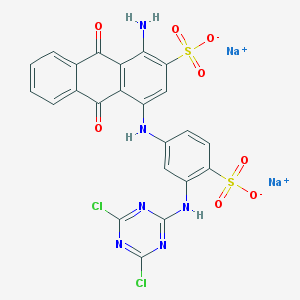 Azure KX Azure KX | C23H12Cl2N6Na2O8S2 | 681.383 | 14 / 3 | N/A | No |
| 62392 | 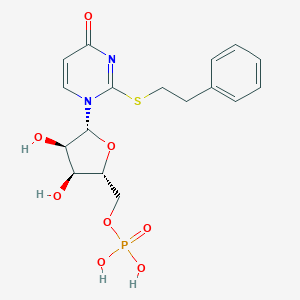 CHEMBL1767421 CHEMBL1767421 | C17H21N2O8PS | 444.395 | 9 / 4 | 0.1 | Yes |
| 68552 | 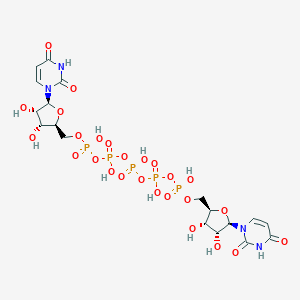 CHEMBL2113401 CHEMBL2113401 | C18H27N4O26P5 | 870.285 | 26 / 11 | -10.2 | No |
| 73968 | 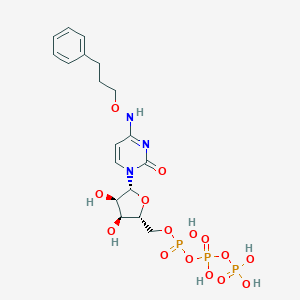 MRS-4062 MRS-4062 | C18H26N3O15P3 | 617.333 | 15 / 7 | -3.7 | No |
| 75983 | 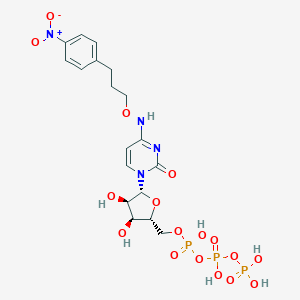 CHEMBL3261368 CHEMBL3261368 | C18H25N4O17P3 | 662.33 | 17 / 7 | -3.9 | No |
| 76441 | 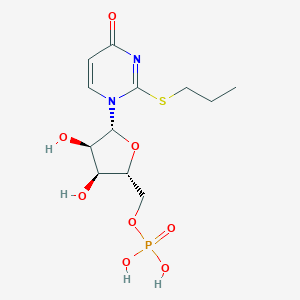 CHEMBL1767419 CHEMBL1767419 | C12H19N2O8PS | 382.324 | 9 / 4 | -1.0 | Yes |
| 559692 | 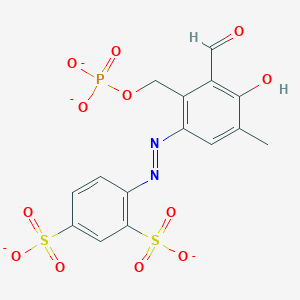 CHEMBL477339 CHEMBL477339 | C15H11N2O12PS2-4 | 506.349 | 14 / 1 | -1.1 | No |
| 77910 | 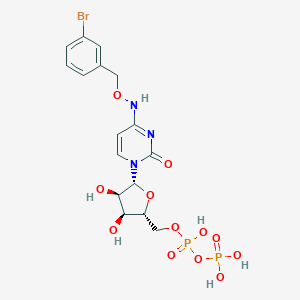 CHEMBL3220046 CHEMBL3220046 | C16H20BrN3O12P2 | 588.197 | 12 / 6 | -2.7 | No |
| 78875 | 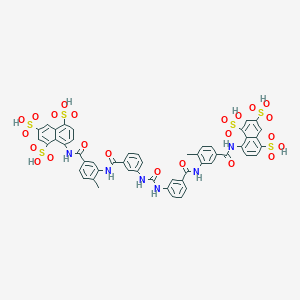 suramin suramin | C51H40N6O23S6 | 1297.26 | 23 / 12 | 1.5 | No |
| 79696 | 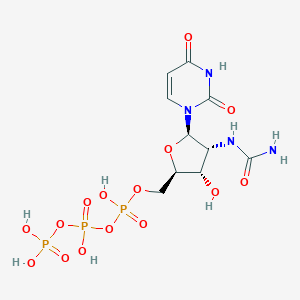 CHEMBL484295 CHEMBL484295 | C10H17N4O15P3 | 526.18 | 15 / 8 | -6.4 | No |
| 82497 | 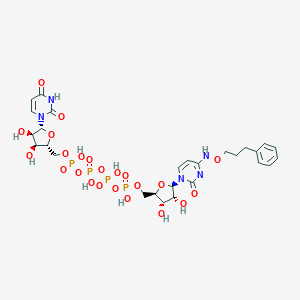 CHEMBL3261379 CHEMBL3261379 | C27H37N5O23P4 | 923.5 | 23 / 10 | -6.7 | No |
| 83771 | 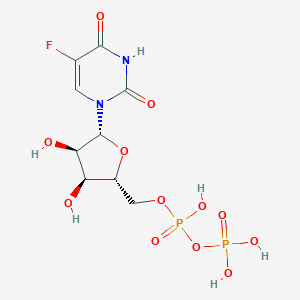 5-fluorouridine diphosphate 5-fluorouridine diphosphate | C9H13FN2O12P2 | 422.151 | 13 / 6 | -4.7 | No |
| 84171 | 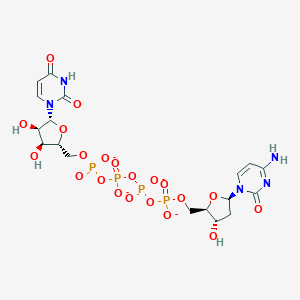 CHEMBL1767407 CHEMBL1767407 | C18H23N5O21P4-4 | 769.291 | 21 / 5 | -8.8 | No |
| 84175 | 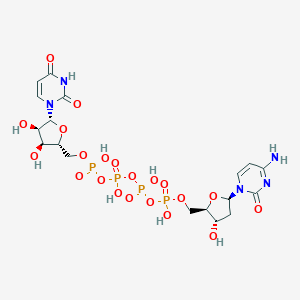 denufosol denufosol | C18H27N5O21P4 | 773.323 | 21 / 9 | -8.4 | No |
| 85065 | 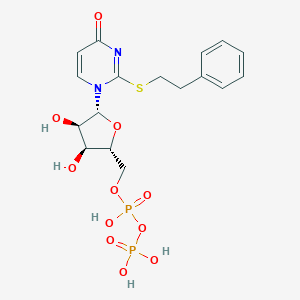 CHEMBL1767413 CHEMBL1767413 | C17H22N2O11P2S | 524.374 | 12 / 5 | -1.9 | No |
| 90106 | 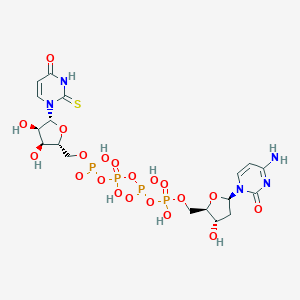 CHEMBL509434 CHEMBL509434 | C18H27N5O20P4S | 789.384 | 21 / 9 | -7.8 | No |
| 91001 | 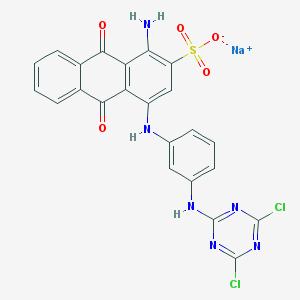 PSB-10211 PSB-10211 | C23H13Cl2N6NaO5S | 579.344 | 11 / 3 | N/A | No |
| 91124 | 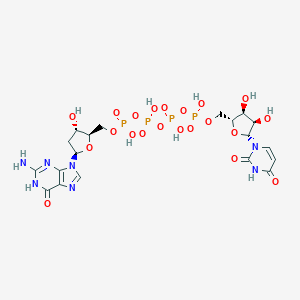 CHEMBL505403 CHEMBL505403 | C19H27N7O21P4 | 813.348 | 23 / 10 | -8.6 | No |
| 94098 | 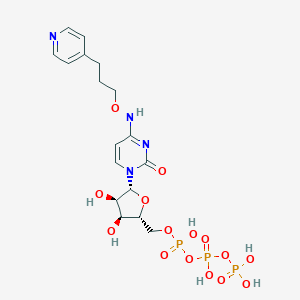 CHEMBL3261371 CHEMBL3261371 | C17H25N4O15P3 | 618.321 | 16 / 7 | -4.8 | No |
| 97497 | 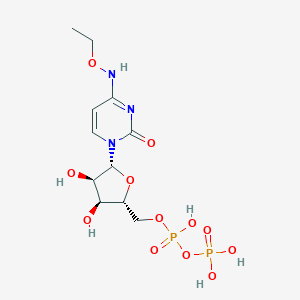 CHEMBL1084295 CHEMBL1084295 | C11H19N3O12P2 | 447.23 | 12 / 6 | -3.7 | No |
| 98051 | 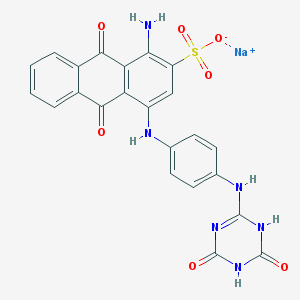 CHEMBL1672106 CHEMBL1672106 | C23H15N6NaO7S | 542.458 | 9 / 5 | N/A | No |
| 99040 | 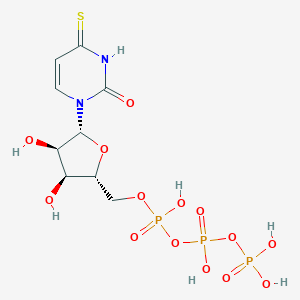 4-Thio-utp 4-Thio-utp | C9H15N2O14P3S | 500.2 | 15 / 7 | -5.2 | No |
| 100341 | 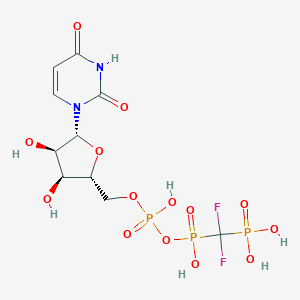 CHEMBL1765117 CHEMBL1765117 | C10H15F2N2O14P3 | 518.148 | 16 / 7 | -5.2 | No |
| 104006 | 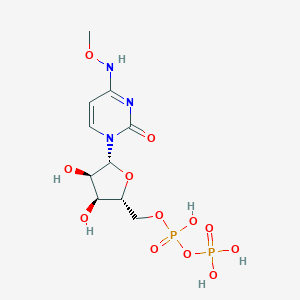 CHEMBL1084020 CHEMBL1084020 | C10H17N3O12P2 | 433.203 | 12 / 6 | -4.1 | No |
| 553797 | 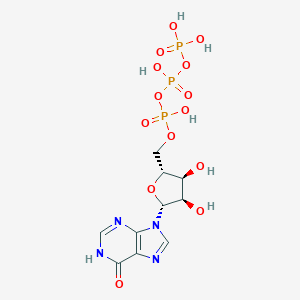 inosine triphosphate inosine triphosphate | C10H15N4O14P3 | 508.165 | 16 / 7 | -5.1 | No |
| 109888 | 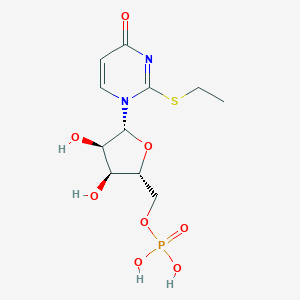 CHEMBL1767418 CHEMBL1767418 | C11H17N2O8PS | 368.297 | 9 / 4 | -1.5 | Yes |
| 110374 | 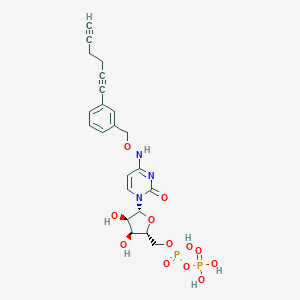 CHEMBL3220051 CHEMBL3220051 | C22H25N3O12P2 | 585.399 | 12 / 6 | -2.2 | No |
| 110820 | 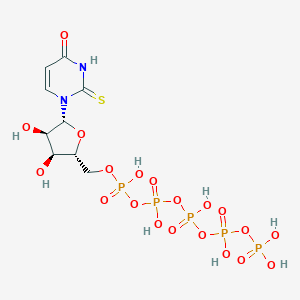 CHEMBL482683 CHEMBL482683 | C9H17N2O20P5S | 660.158 | 21 / 9 | -7.4 | No |
| 110828 | 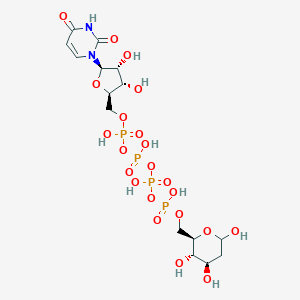 CHEMBL454230 CHEMBL454230 | C15H26N2O22P4 | 710.26 | 22 / 10 | -8.1 | No |
| 113312 | 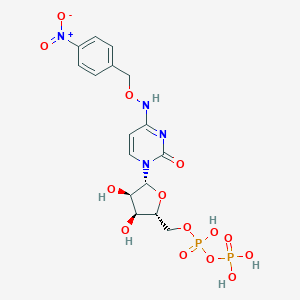 CHEMBL3220050 CHEMBL3220050 | C16H20N4O14P2 | 554.298 | 14 / 6 | -3.6 | No |
| 115802 | 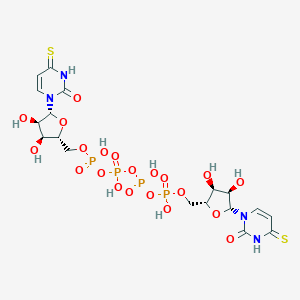 CHEMBL503798 CHEMBL503798 | C18H26N4O21P4S2 | 822.428 | 23 / 10 | -7.9 | No |
| 120595 | 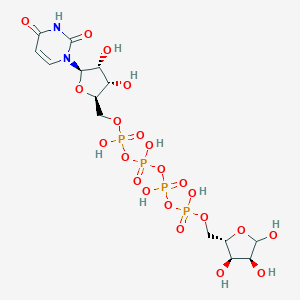 CHEMBL448985 CHEMBL448985 | C14H24N2O22P4 | 696.233 | 22 / 10 | -8.4 | No |
| 122940 | 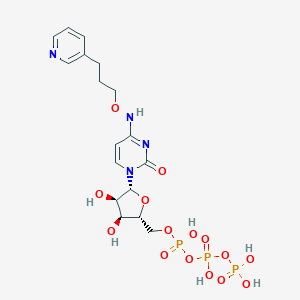 CHEMBL3261370 CHEMBL3261370 | C17H25N4O15P3 | 618.321 | 16 / 7 | -4.8 | No |
| 126433 | 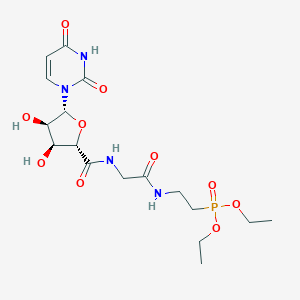 CHEMBL469244 CHEMBL469244 | C17H27N4O10P | 478.395 | 10 / 5 | -3.0 | Yes |
| 130390 | 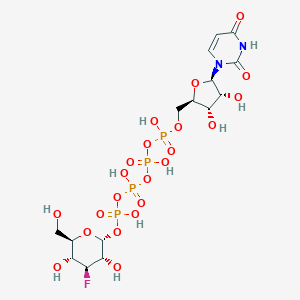 CHEMBL1784904 CHEMBL1784904 | C15H25FN2O22P4 | 728.25 | 23 / 10 | -7.5 | No |
| 130703 | 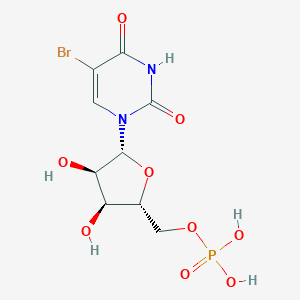 5-Brump 5-Brump | C9H12BrN2O9P | 403.078 | 9 / 5 | -3.0 | Yes |
| 135255 | 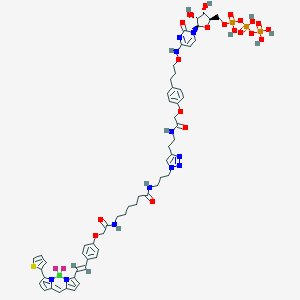 CHEMBL3261378 CHEMBL3261378 | C56H67BF2N11O20P3S | 1388.0 | 26 / 10 | N/A | No |
| 139972 | 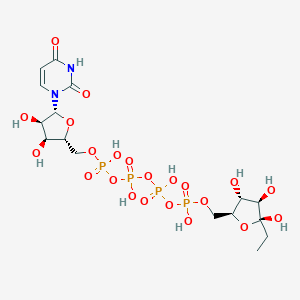 CHEMBL500840 CHEMBL500840 | C16H28N2O22P4 | 724.287 | 22 / 10 | -8.1 | No |
| 142532 | 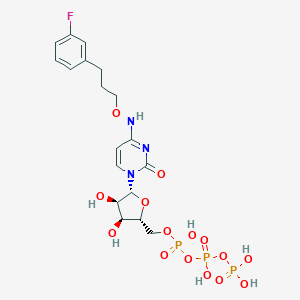 CHEMBL3261362 CHEMBL3261362 | C18H25FN3O15P3 | 635.324 | 16 / 7 | -3.6 | No |
| 143288 | 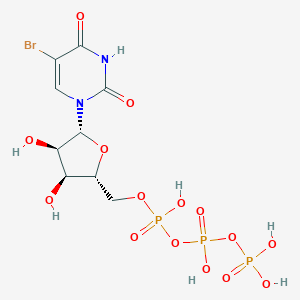 5-Bromo-utp 5-Bromo-utp | C9H14BrN2O15P3 | 563.035 | 15 / 7 | -5.2 | No |
| 143719 | 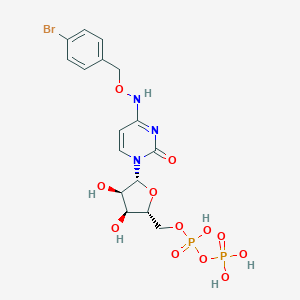 CHEMBL3220048 CHEMBL3220048 | C16H20BrN3O12P2 | 588.197 | 12 / 6 | -2.7 | No |
| 149453 | 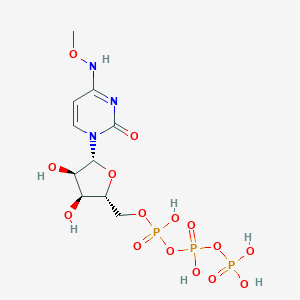 CHEMBL1198849 CHEMBL1198849 | C10H18N3O15P3 | 513.181 | 15 / 7 | -5.2 | No |
| 150993 | 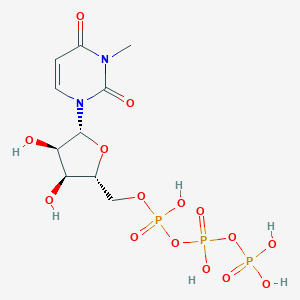 CHEMBL215783 CHEMBL215783 | C10H17N2O15P3 | 498.166 | 15 / 6 | -6.0 | No |
| 153113 | 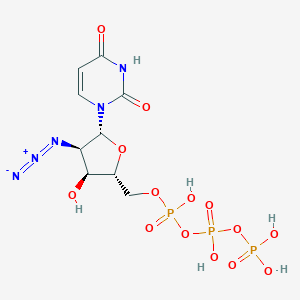 CHEMBL1784888 CHEMBL1784888 | C9H14N5O14P3 | 509.153 | 16 / 6 | -4.6 | No |
| 155360 | 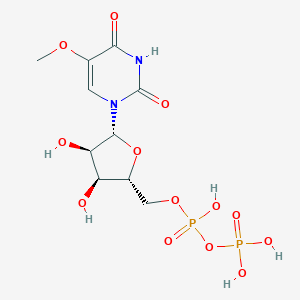 CHEMBL590738 CHEMBL590738 | C10H16N2O13P2 | 434.187 | 13 / 6 | -4.9 | No |
| 156672 | 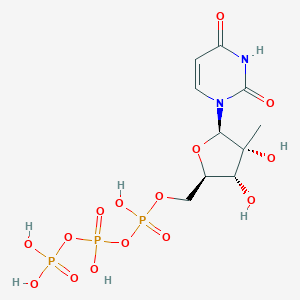 CHEMBL521487 CHEMBL521487 | C10H17N2O15P3 | 498.166 | 15 / 7 | -6.0 | No |
| 156695 | 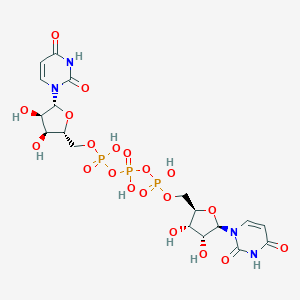 CHEMBL239151 CHEMBL239151 | C18H25N4O20P3 | 710.327 | 20 / 9 | -8.0 | No |
| 156700 | 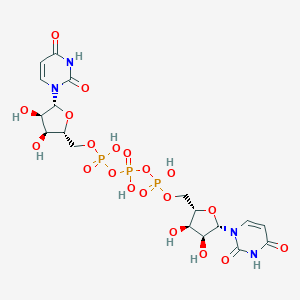 CHEMBL1162193 CHEMBL1162193 | C18H25N4O20P3 | 710.327 | 20 / 9 | -8.0 | No |
| 156882 | 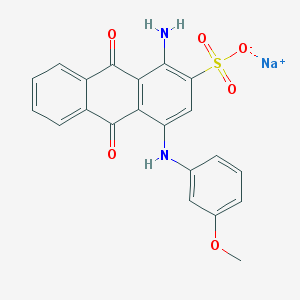 78510-28-8 78510-28-8 | C21H15N2NaO6S | 446.409 | 8 / 2 | N/A | N/A |
| 157774 | 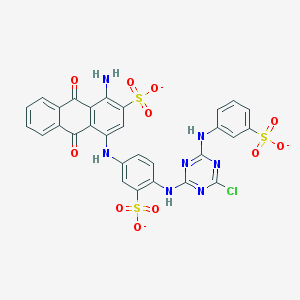 Basilen Blue Basilen Blue | C29H17ClN7O11S3-3 | 771.123 | 18 / 4 | 4.1 | No |
| 161816 | 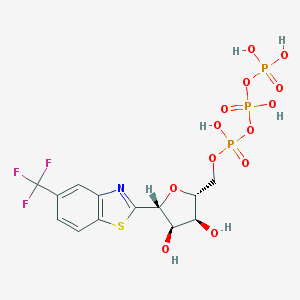 CHEMBL391317 CHEMBL391317 | C13H15F3NO13P3S | 575.233 | 18 / 6 | -2.4 | No |
| 162299 | 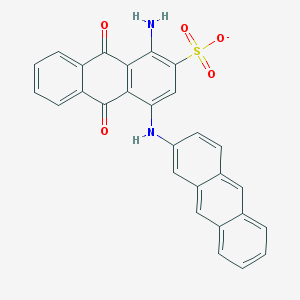 CHEMBL608559 CHEMBL608559 | C28H17N2O5S- | 493.513 | 7 / 2 | 6.0 | No |
| 167577 | 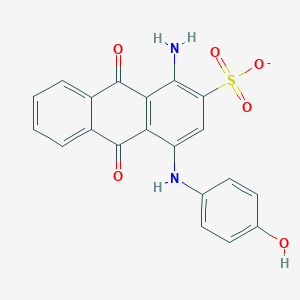 CHEMBL496030 CHEMBL496030 | C20H13N2O6S- | 409.392 | 8 / 3 | 3.1 | Yes |
| 169590 | 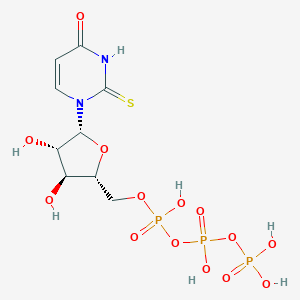 CHEMBL484494 CHEMBL484494 | C9H15N2O14P3S | 500.2 | 15 / 7 | -5.2 | No |
| 169594 | 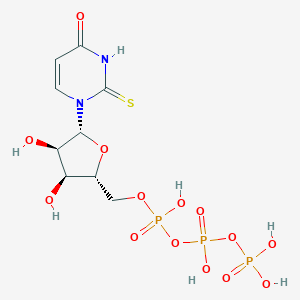 2-thioUTP 2-thioUTP | C9H15N2O14P3S | 500.2 | 15 / 7 | -5.2 | No |
| 170304 | 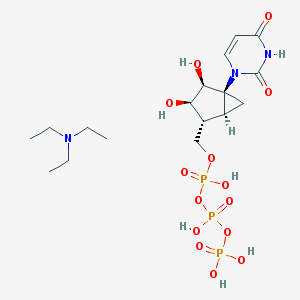 CHEMBL1083148 CHEMBL1083148 | C17H32N3O14P3 | 595.371 | 15 / 7 | N/A | No |
| 171530 | 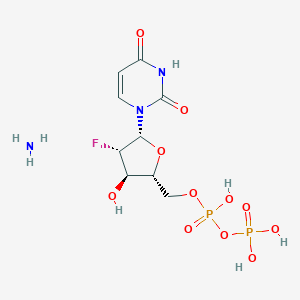 CHEMBL1161886 CHEMBL1161886 | C9H16FN3O11P2 | 423.183 | 13 / 6 | N/A | No |
| 172855 | 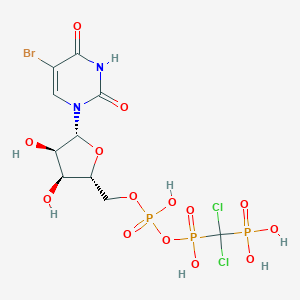 CHEMBL217467 CHEMBL217467 | C10H14BrCl2N2O14P3 | 629.947 | 14 / 7 | -4.1 | No |
| 175123 | 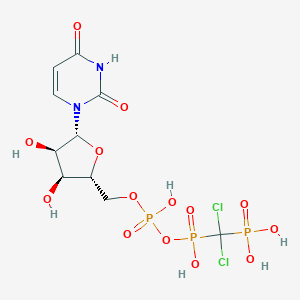 CHEMBL1765114 CHEMBL1765114 | C10H15Cl2N2O14P3 | 551.051 | 14 / 7 | -4.7 | No |
| 178349 | 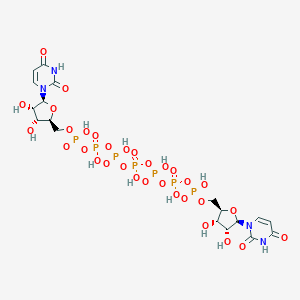 CHEMBL2113398 CHEMBL2113398 | C18H29N4O32P7 | 1030.24 | 32 / 13 | -12.4 | No |
| 182123 | 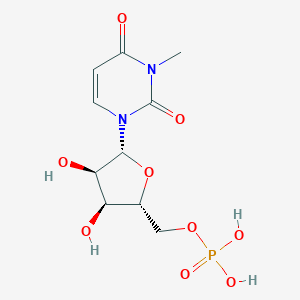 CHEMBL215940 CHEMBL215940 | C10H15N2O9P | 338.209 | 9 / 4 | -3.8 | Yes |
| 184866 | 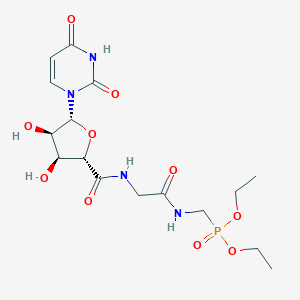 CHEMBL468354 CHEMBL468354 | C16H25N4O10P | 464.368 | 10 / 5 | -2.9 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218