

You can:
| Name | C-C chemokine receptor type 6 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | CCR6 |
| Synonym | DRY6 G-protein coupled receptor 29 GPR-CY4 GPR29 GPRCY4 [ Show all ] |
| Disease | N/A |
| Length | 374 |
| Amino acid sequence | MSGESMNFSDVFDSSEDYFVSVNTSYYSVDSEMLLCSLQEVRQFSRLFVPIAYSLICVFGLLGNILVVITFAFYKKARSMTDVYLLNMAIADILFVLTLPFWAVSHATGAWVFSNATCKLLKGIYAINFNCGMLLLTCISMDRYIAIVQATKSFRLRSRTLPRSKIICLVVWGLSVIISSSTFVFNQKYNTQGSDVCEPKYQTVSEPIRWKLLMLGLELLFGFFIPLMFMIFCYTFIVKTLVQAQNSKRHKAIRVIIAVVLVFLACQIPHNMVLLVTAANLGKMNRSCQSEKLIGYTKTVTEVLAFLHCCLNPVLYAFIGQKFRNYFLKILKDLWCVRRKYKSSGFSCAGRYSENISRQTSETADNDNASSFTM |
| UniProt | P51684 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | P51684 |
| 3D structure model | This predicted structure model is from GPCR-EXP P51684. |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL4423 |
| IUPHAR | N/A |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 382 | 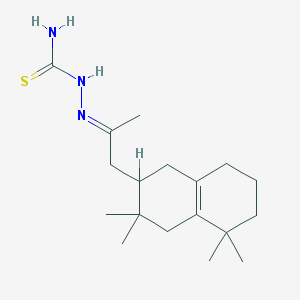 MLS000526436 MLS000526436 | C18H31N3S | 321.527 | 2 / 2 | 3.7 | Yes |
| 384 | 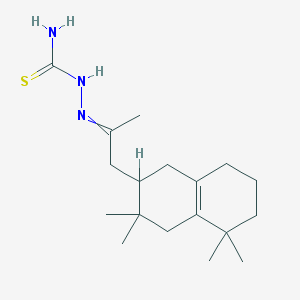 AC1MEFGH AC1MEFGH | C18H31N3S | 321.527 | 2 / 2 | 3.7 | Yes |
| 1875 | 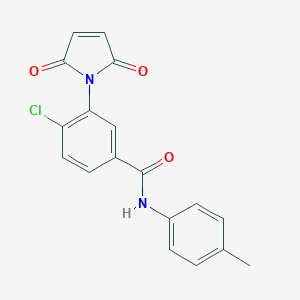 MLS000574219 MLS000574219 | C18H13ClN2O3 | 340.763 | 3 / 1 | 2.9 | Yes |
| 1894 | 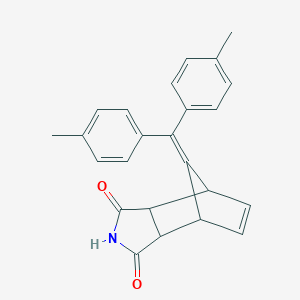 MLS000537663 MLS000537663 | C24H21NO2 | 355.437 | 2 / 1 | 4.1 | Yes |
| 3274 | 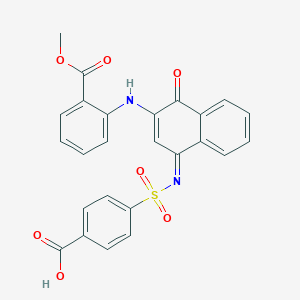 MLS000948142 MLS000948142 | C25H18N2O7S | 490.486 | 9 / 2 | 4.5 | Yes |
| 3275 | 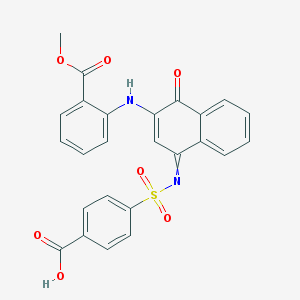 AC1LVDP6 AC1LVDP6 | C25H18N2O7S | 490.486 | 9 / 2 | 4.5 | Yes |
| 5343 | 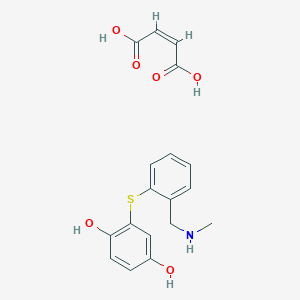 CHEMBL1359359 CHEMBL1359359 | C18H19NO6S | 377.411 | 8 / 5 | N/A | N/A |
| 8596 | 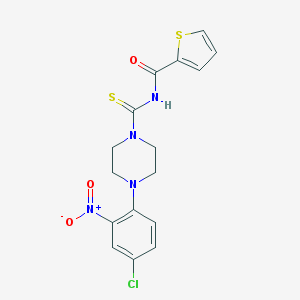 MLS000573225 MLS000573225 | C16H15ClN4O3S2 | 410.891 | 6 / 1 | 3.7 | Yes |
| 557549 | 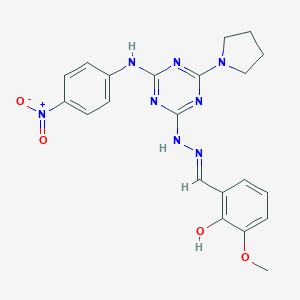 MLS000689796 MLS000689796 | C21H22N8O4 | 450.459 | 11 / 3 | 4.2 | No |
| 9718 | 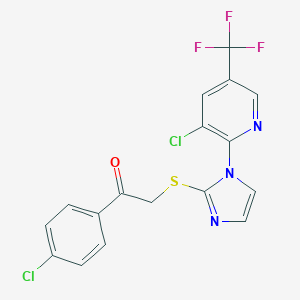 MLS000326742 MLS000326742 | C17H10Cl2F3N3OS | 432.242 | 7 / 0 | 5.3 | No |
| 9944 | 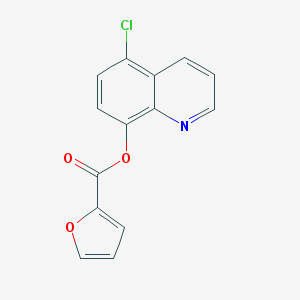 SMR000120908 SMR000120908 | C14H8ClNO3 | 273.672 | 4 / 0 | 3.6 | Yes |
| 12851 | 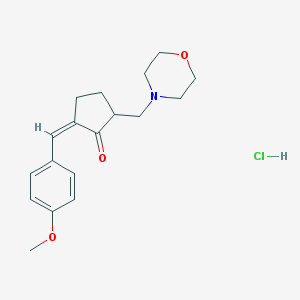 MLS002701690 MLS002701690 | C18H24ClNO3 | 337.844 | 4 / 1 | N/A | N/A |
| 13286 | 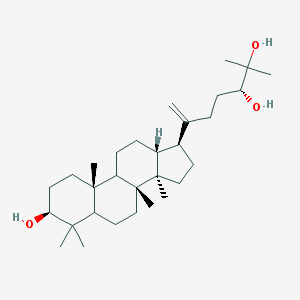 Aglaitriol Aglaitriol | C30H52O3 | 460.743 | 3 / 3 | 7.7 | No |
| 442182 | 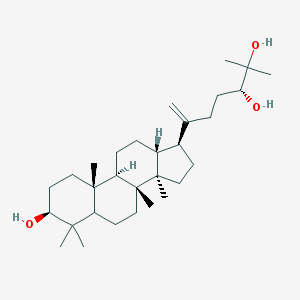 CHEMBL3392059 CHEMBL3392059 | C30H52O3 | 460.743 | 3 / 3 | 7.7 | No |
| 13302 | 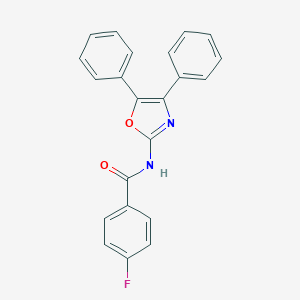 N-(4,5-diphenyl-1,3-oxazol-2-yl)-4-fluorobenzamide N-(4,5-diphenyl-1,3-oxazol-2-yl)-4-fluorobenzamide | C22H15FN2O2 | 358.372 | 4 / 1 | 4.9 | Yes |
| 557720 | 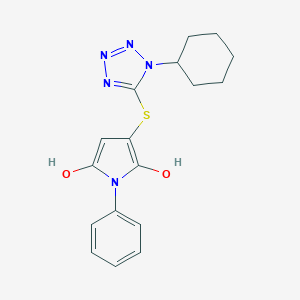 BDBM79970 BDBM79970 | C17H19N5O2S | 357.432 | 6 / 2 | 4.0 | Yes |
| 16892 | 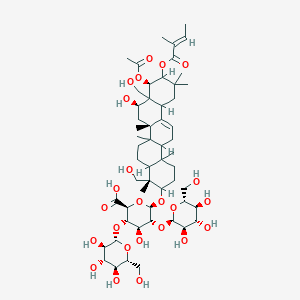 beta-Escin beta-Escin | C55H86O24 | 1131.27 | 24 / 13 | 0.7 | No |
| 557763 | 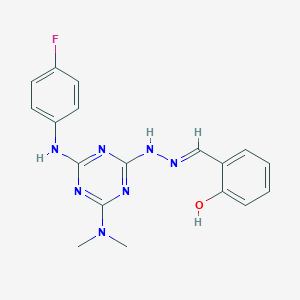 AC1O0HXG AC1O0HXG | C18H18FN7O | 367.388 | 9 / 3 | 4.0 | Yes |
| 17924 | 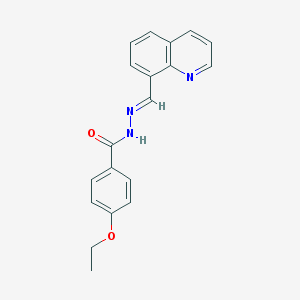 MLS000570968 MLS000570968 | C19H17N3O2 | 319.364 | 4 / 1 | 3.4 | Yes |
| 17925 | 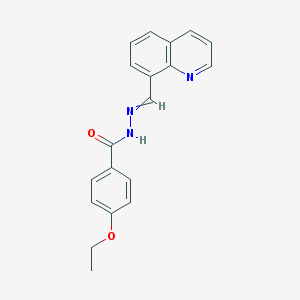 4-ethoxy-N-(quinolin-8-ylmethylideneamino)benzamide 4-ethoxy-N-(quinolin-8-ylmethylideneamino)benzamide | C19H17N3O2 | 319.364 | 4 / 1 | 3.4 | Yes |
| 18159 | 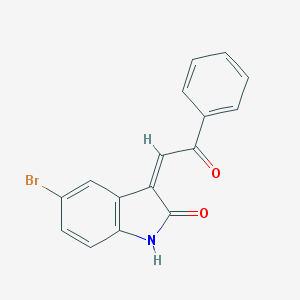 MLS002702136 MLS002702136 | C16H10BrNO2 | 328.165 | 2 / 1 | 3.1 | Yes |
| 19055 | 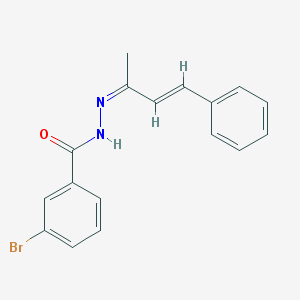 MLS000709305 MLS000709305 | C17H15BrN2O | 343.224 | 2 / 1 | 4.2 | Yes |
| 19056 | 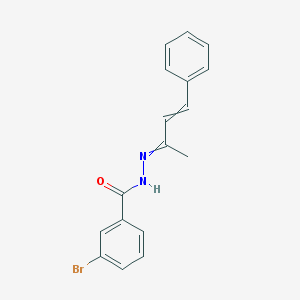 AC1LEVXZ AC1LEVXZ | C17H15BrN2O | 343.224 | 2 / 1 | 4.2 | Yes |
| 19754 | 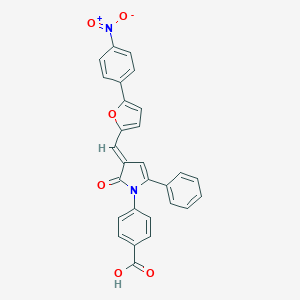 4E1RCat 4E1RCat | C28H18N2O6 | 478.46 | 6 / 1 | 5.3 | No |
| 20704 | 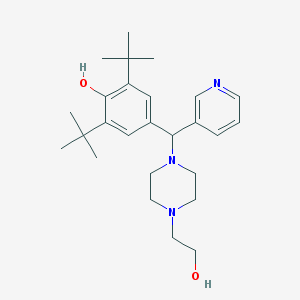 MLS000054481 MLS000054481 | C26H39N3O2 | 425.617 | 5 / 2 | 4.5 | Yes |
| 21061 | 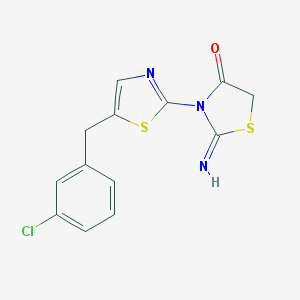 MLS000707891 MLS000707891 | C13H10ClN3OS2 | 323.813 | 5 / 1 | 3.8 | Yes |
| 557948 | 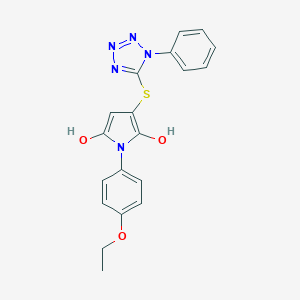 BDBM54738 BDBM54738 | C19H17N5O3S | 395.437 | 7 / 2 | 4.3 | Yes |
| 22234 | 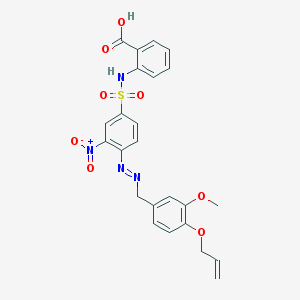 BDBM80056 BDBM80056 | C24H22N4O8S | 526.52 | 11 / 2 | 4.7 | No |
| 22777 | 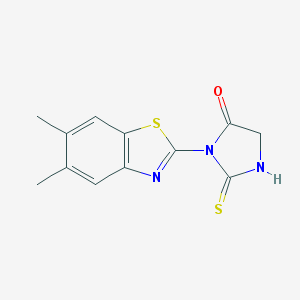 3-(5,6-dimethyl-1,3-benzothiazol-2-yl)-2-thioxoimidazolidin-4-one 3-(5,6-dimethyl-1,3-benzothiazol-2-yl)-2-thioxoimidazolidin-4-one | C12H11N3OS2 | 277.36 | 4 / 1 | 2.8 | Yes |
| 24759 | 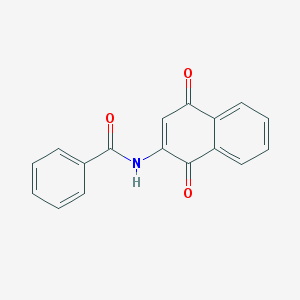 ppm-18 ppm-18 | C17H11NO3 | 277.279 | 3 / 1 | 2.9 | Yes |
| 27688 | 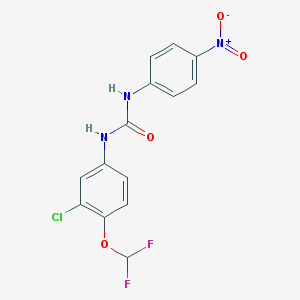 MLS000581434 MLS000581434 | C14H10ClF2N3O4 | 357.698 | 6 / 2 | 4.1 | Yes |
| 30671 | 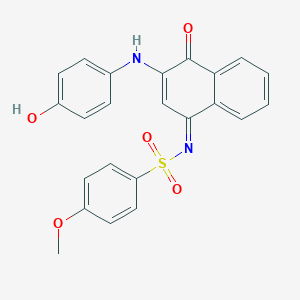 MLS000948021 MLS000948021 | C23H18N2O5S | 434.466 | 7 / 2 | 3.9 | Yes |
| 30672 | 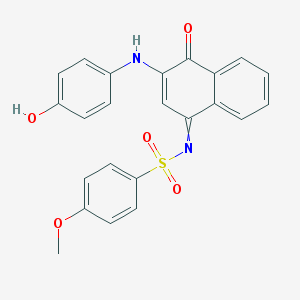 GNF-Pf-2153 GNF-Pf-2153 | C23H18N2O5S | 434.466 | 7 / 2 | 3.9 | Yes |
| 536783 | 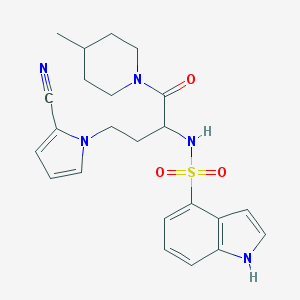 CHEMBL3889627 CHEMBL3889627 | C23H27N5O3S | 453.561 | 5 / 2 | 2.6 | Yes |
| 31315 | 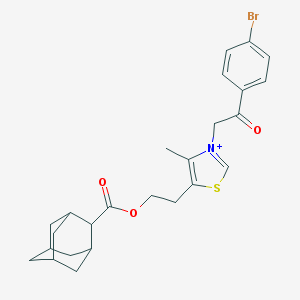 MLS000533789 MLS000533789 | C25H29BrNO3S+ | 503.475 | 4 / 0 | 6.9 | No |
| 33221 | 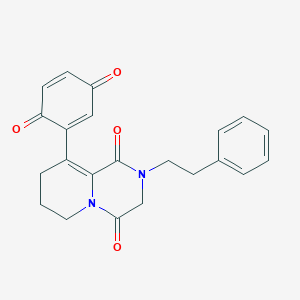 MLS000705957 MLS000705957 | C22H20N2O4 | 376.412 | 4 / 0 | 1.3 | Yes |
| 33573 | 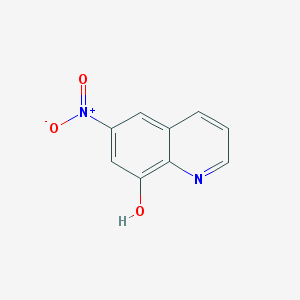 6-nitroquinolin-8-ol 6-nitroquinolin-8-ol | C9H6N2O3 | 190.158 | 4 / 1 | 1.5 | Yes |
| 34688 | 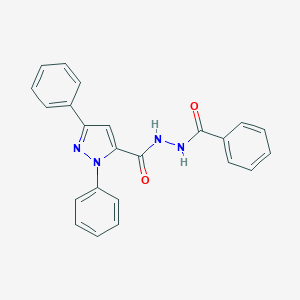 MLS000388665 MLS000388665 | C23H18N4O2 | 382.423 | 3 / 2 | 4.3 | Yes |
| 467451 | 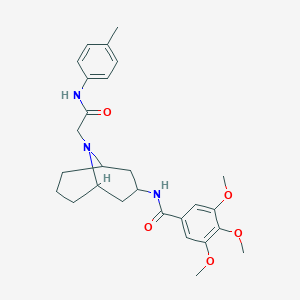 CHEMBL3187346 CHEMBL3187346 | C27H35N3O5 | 481.593 | 6 / 2 | 4.0 | Yes |
| 42276 | 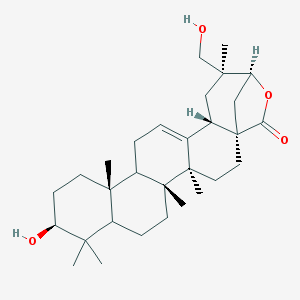 Machaerogenin Machaerogenin | C30H46O4 | 470.694 | 4 / 2 | 5.7 | No |
| 44105 | 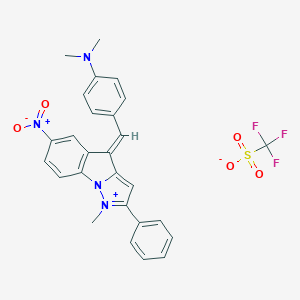 NSC-720622 NSC-720622 | C27H23F3N4O5S | 572.559 | 9 / 0 | N/A | No |
| 45816 | 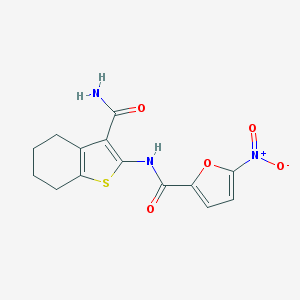 BAS 06848191 BAS 06848191 | C14H13N3O5S | 335.334 | 6 / 2 | 3.0 | Yes |
| 558740 | 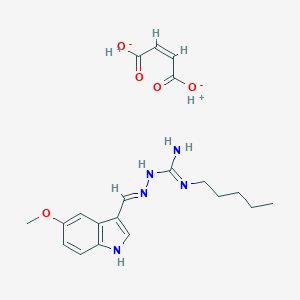 Tegaserod maleate Tegaserod maleate | C20H27N5O5 | 417.466 | 7 / 5 | N/A | N/A |
| 469073 | 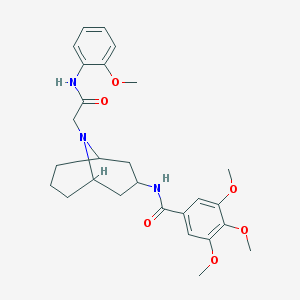 CHEMBL3188149 CHEMBL3188149 | C27H35N3O6 | 497.592 | 7 / 2 | 3.6 | Yes |
| 50334 | 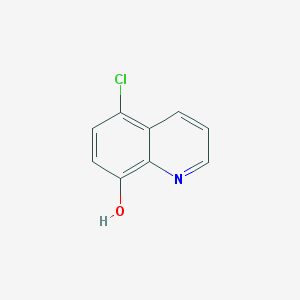 5-CHLORO-8-HYDROXYQUINOLINE 5-CHLORO-8-HYDROXYQUINOLINE | C9H6ClNO | 179.603 | 2 / 1 | 2.9 | Yes |
| 558851 | 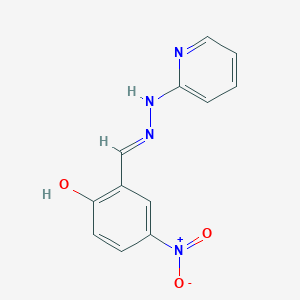 2-hydroxy-5-nitrobenzaldehyde pyridin-2-ylhydrazone 2-hydroxy-5-nitrobenzaldehyde pyridin-2-ylhydrazone | C12H10N4O3 | 258.237 | 6 / 2 | 2.3 | Yes |
| 558899 | 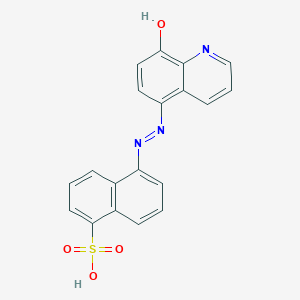 MLS001197249 MLS001197249 | C19H13N3O4S | 379.39 | 7 / 2 | 3.6 | Yes |
| 53009 | 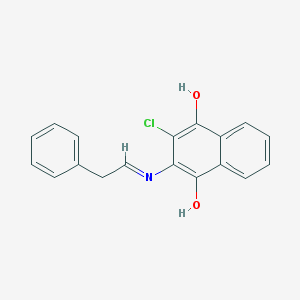 cid_237490 cid_237490 | C18H14ClNO2 | 311.765 | 3 / 2 | 4.3 | Yes |
| 53241 | 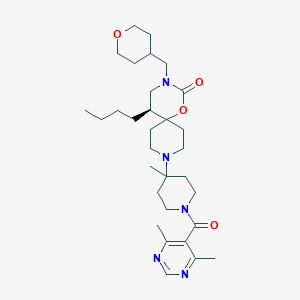 PharmaGSID_48521 PharmaGSID_48521 | C31H49N5O4 | 555.764 | 7 / 0 | 4.1 | No |
| 54562 | 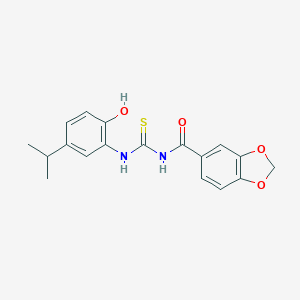 MLS001035753 MLS001035753 | C18H18N2O4S | 358.412 | 5 / 3 | 4.1 | Yes |
| 55295 | 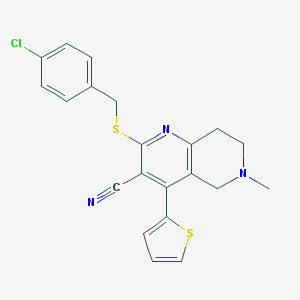 SMR000160396 SMR000160396 | C21H18ClN3S2 | 411.966 | 5 / 0 | 4.8 | Yes |
| 55737 | 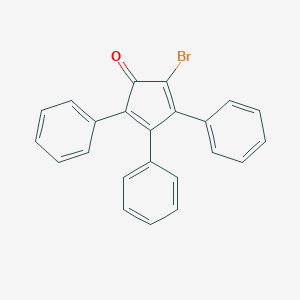 2-Bromo-3,4,5-triphenyl-2,4-cyclopentadien-1-one 2-Bromo-3,4,5-triphenyl-2,4-cyclopentadien-1-one | C23H15BrO | 387.276 | 1 / 0 | 5.7 | No |
| 469871 | 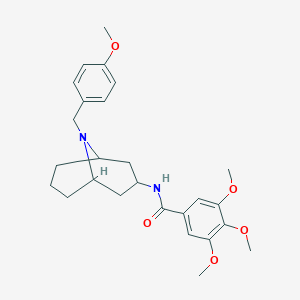 CHEMBL3185809 CHEMBL3185809 | C26H34N2O5 | 454.567 | 6 / 1 | 4.1 | Yes |
| 559057 | 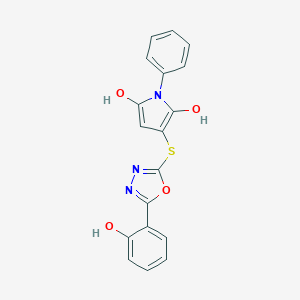 BDBM79969 BDBM79969 | C18H13N3O4S | 367.379 | 7 / 3 | 3.8 | Yes |
| 59214 | 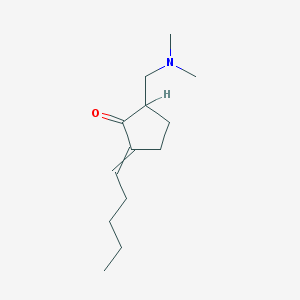 NCI60_008501 NCI60_008501 | C13H23NO | 209.333 | 2 / 0 | 2.8 | Yes |
| 60933 | 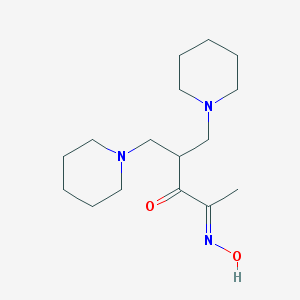 AC1OCRI6 AC1OCRI6 | C16H29N3O2 | 295.427 | 5 / 1 | 2.1 | Yes |
| 62389 | 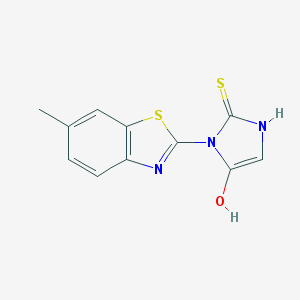 cid_896091 cid_896091 | C11H9N3OS2 | 263.333 | 4 / 2 | 2.5 | Yes |
| 63063 | 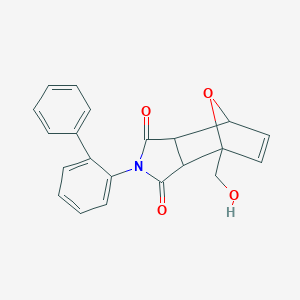 AC1MEYUL AC1MEYUL | C21H17NO4 | 347.37 | 4 / 1 | 1.5 | Yes |
| 66490 | 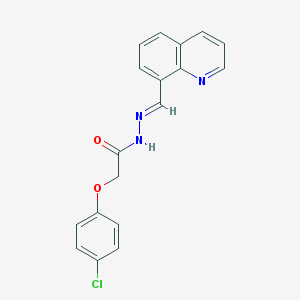 MLS001125716 MLS001125716 | C18H14ClN3O2 | 339.779 | 4 / 1 | 3.8 | Yes |
| 66491 | 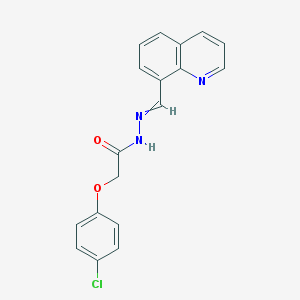 2-(4-chlorophenoxy)-N-(quinolin-8-ylmethylideneamino)acetamide 2-(4-chlorophenoxy)-N-(quinolin-8-ylmethylideneamino)acetamide | C18H14ClN3O2 | 339.779 | 4 / 1 | 3.8 | Yes |
| 67030 | 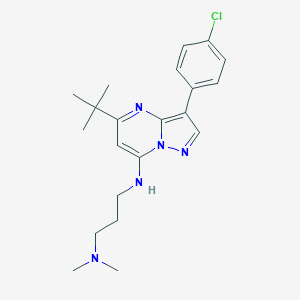 MLS000762858 MLS000762858 | C21H28ClN5 | 385.94 | 4 / 1 | 5.0 | Yes |
| 68014 | 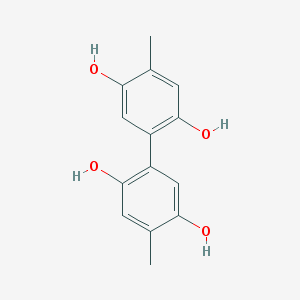 NSC2805 NSC2805 | C14H14O4 | 246.262 | 4 / 4 | 2.9 | Yes |
| 68986 | 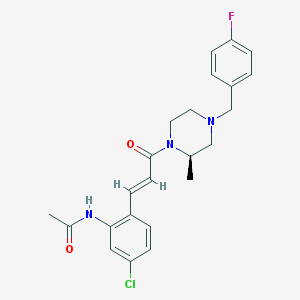 CHEMBL198949 CHEMBL198949 | C23H25ClFN3O2 | 429.92 | 4 / 1 | 3.5 | Yes |
| 70175 | 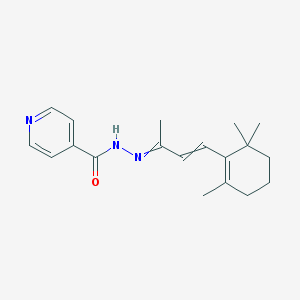 AC1L65UY AC1L65UY | C19H25N3O | 311.429 | 3 / 1 | 3.3 | Yes |
| 70176 | 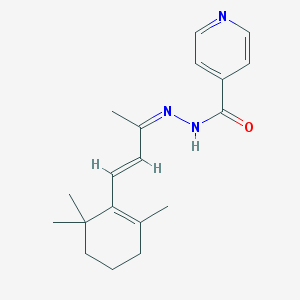 MLS000711749 MLS000711749 | C19H25N3O | 311.429 | 3 / 1 | 3.3 | Yes |
| 71301 | 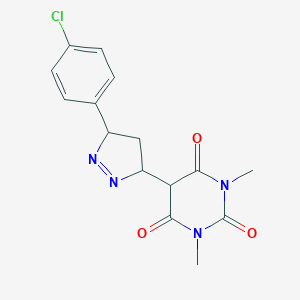 BDBM72680 BDBM72680 | C15H15ClN4O3 | 334.76 | 5 / 0 | 1.7 | Yes |
| 81862 | 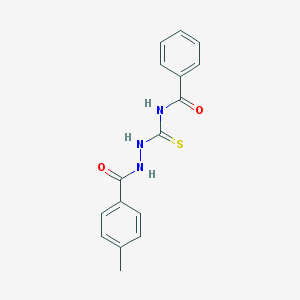 MLS001180103 MLS001180103 | C16H15N3O2S | 313.375 | 3 / 3 | 3.5 | Yes |
| 84712 | 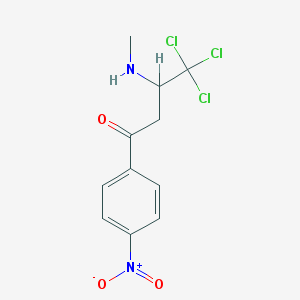 4,4,4-trichloro-3-(methylamino)-1-(4-nitrophenyl)butan-1-one 4,4,4-trichloro-3-(methylamino)-1-(4-nitrophenyl)butan-1-one | C11H11Cl3N2O3 | 325.57 | 4 / 1 | 2.9 | Yes |
| 89295 | 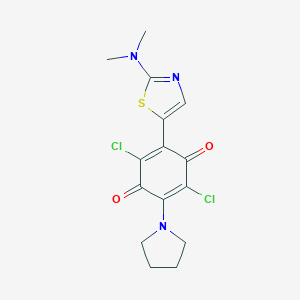 MLS000548208 MLS000548208 | C15H15Cl2N3O2S | 372.264 | 6 / 0 | 3.7 | Yes |
| 91765 | 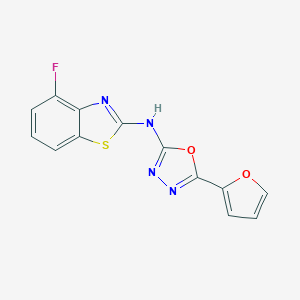 SMR000027223 SMR000027223 | C13H7FN4O2S | 302.283 | 8 / 1 | 3.3 | Yes |
| 474306 | 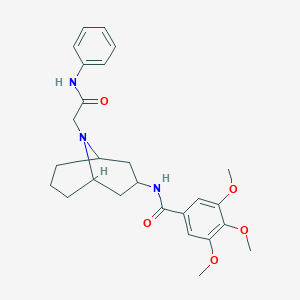 SMR000006668 SMR000006668 | C26H33N3O5 | 467.566 | 6 / 2 | 3.6 | Yes |
| 93040 | 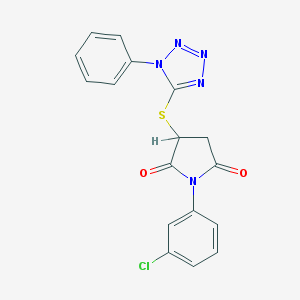 1-(3-chlorophenyl)-3-[(1-phenyl-1H-tetrazol-5-yl)sulfanyl]pyrrolidine-2,5-dione 1-(3-chlorophenyl)-3-[(1-phenyl-1H-tetrazol-5-yl)sulfanyl]pyrrolidine-2,5-dione | C17H12ClN5O2S | 385.826 | 6 / 0 | 3.3 | Yes |
| 95087 | 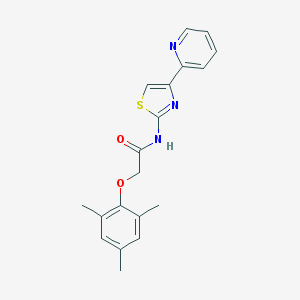 MLS001007696 MLS001007696 | C19H19N3O2S | 353.44 | 5 / 1 | 3.9 | Yes |
| 95192 | 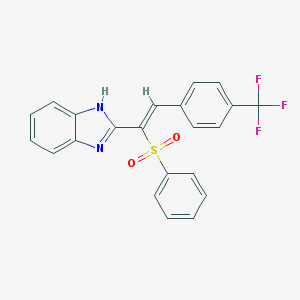 MLS002635606 MLS002635606 | C22H15F3N2O2S | 428.429 | 6 / 1 | 5.3 | No |
| 95632 | 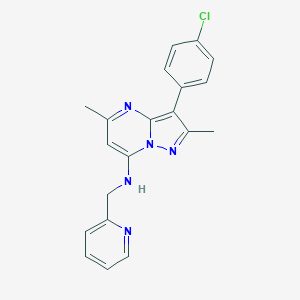 3-(4-chlorophenyl)-2,5-dimethyl-N-(pyridin-2-ylmethyl)pyrazolo[1,5-a]pyrimidin-7-amine 3-(4-chlorophenyl)-2,5-dimethyl-N-(pyridin-2-ylmethyl)pyrazolo[1,5-a]pyrimidin-7-amine | C20H18ClN5 | 363.849 | 4 / 1 | 4.2 | Yes |
| 102560 | 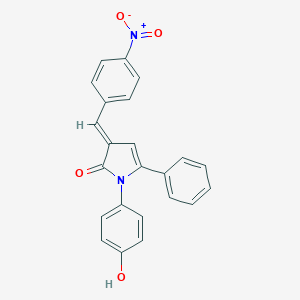 SMR000131790 SMR000131790 | C23H16N2O4 | 384.391 | 4 / 1 | 4.6 | Yes |
| 102690 | 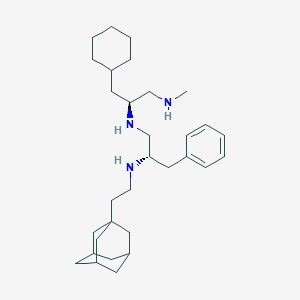 MLS001242636 MLS001242636 | C31H51N3 | 465.77 | 3 / 3 | 7.9 | No |
| 103216 | 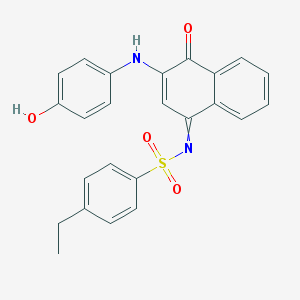 ZINC18037497 ZINC18037497 | C24H20N2O4S | 432.494 | 6 / 2 | 4.7 | Yes |
| 103217 | 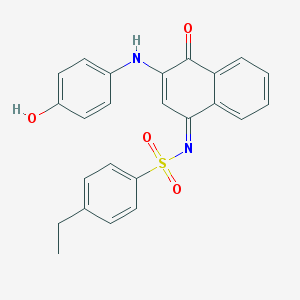 MLS000948011 MLS000948011 | C24H20N2O4S | 432.494 | 6 / 2 | 4.7 | Yes |
| 108294 | 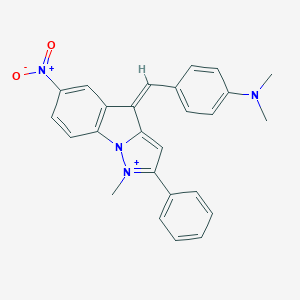 AC1NV8AN AC1NV8AN | C26H23N4O2+ | 423.496 | 3 / 0 | 5.8 | No |
| 110015 | 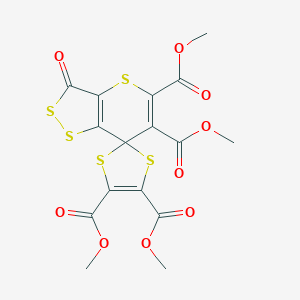 NSC686349 NSC686349 | C16H12O9S5 | 508.563 | 14 / 0 | 2.1 | No |
| 110620 | 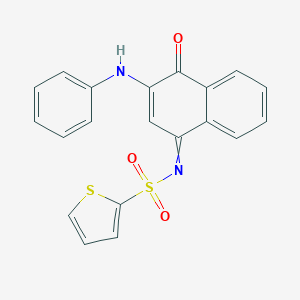 MLS000374486 MLS000374486 | C20H14N2O3S2 | 394.463 | 6 / 1 | 4.3 | Yes |
| 110622 | 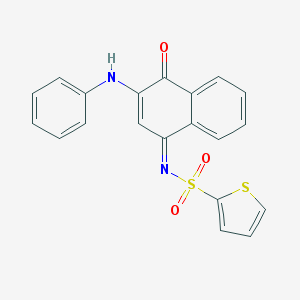 MLS000374486 MLS000374486 | C20H14N2O3S2 | 394.463 | 6 / 1 | 4.3 | Yes |
| 111096 | 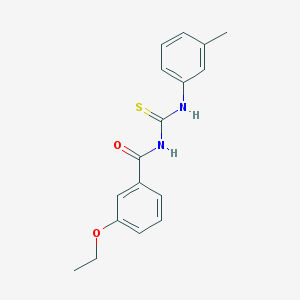 3-ethoxy-N-[(3-methylphenyl)carbamothioyl]benzamide 3-ethoxy-N-[(3-methylphenyl)carbamothioyl]benzamide | C17H18N2O2S | 314.403 | 3 / 2 | 4.2 | Yes |
| 111536 | 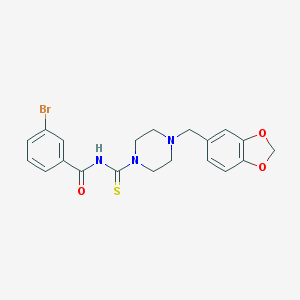 MLS000392397 MLS000392397 | C20H20BrN3O3S | 462.362 | 5 / 1 | 3.4 | Yes |
| 112136 | 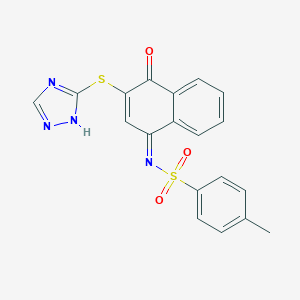 MLS000554446 MLS000554446 | C19H14N4O3S2 | 410.466 | 7 / 1 | 3.9 | Yes |
| 112138 | 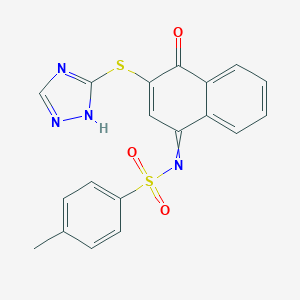 GNF-Pf-1591 GNF-Pf-1591 | C19H14N4O3S2 | 410.466 | 7 / 1 | 3.9 | Yes |
| 114248 | 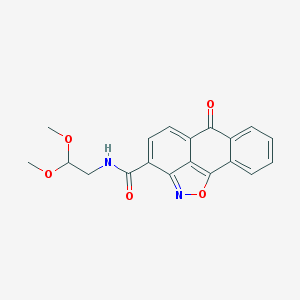 MLS001029500 MLS001029500 | C19H16N2O5 | 352.346 | 6 / 1 | 2.0 | Yes |
| 115566 | 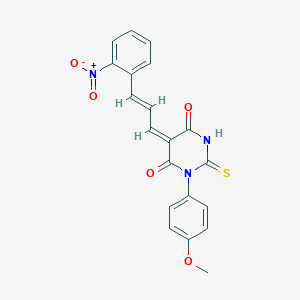 MLS000723835 MLS000723835 | C20H15N3O5S | 409.416 | 6 / 1 | 3.7 | Yes |
| 115567 | 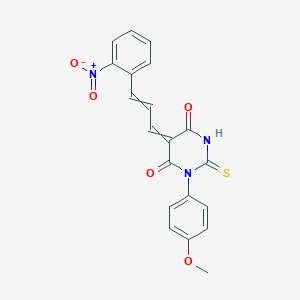 AC1LQTX7 AC1LQTX7 | C20H15N3O5S | 409.416 | 6 / 1 | 3.7 | Yes |
| 115746 | 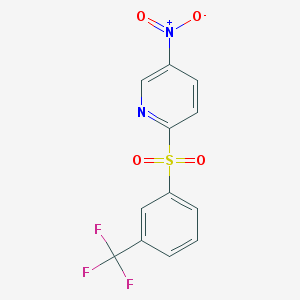 5-nitro-2-{[3-(trifluoromethyl)phenyl]sulfonyl}pyridine 5-nitro-2-{[3-(trifluoromethyl)phenyl]sulfonyl}pyridine | C12H7F3N2O4S | 332.253 | 8 / 0 | 2.7 | Yes |
| 116097 | 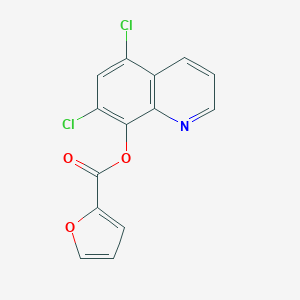 5,7-dichloroquinolin-8-yl furan-2-carboxylate 5,7-dichloroquinolin-8-yl furan-2-carboxylate | C14H7Cl2NO3 | 308.114 | 4 / 0 | 4.3 | Yes |
| 116270 | 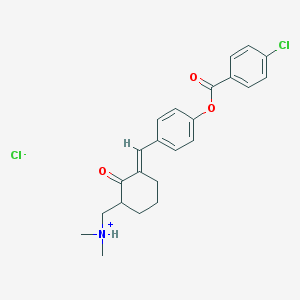 CID 10694119 CID 10694119 | C23H25Cl2NO3 | 434.357 | 4 / 1 | N/A | N/A |
| 116529 | 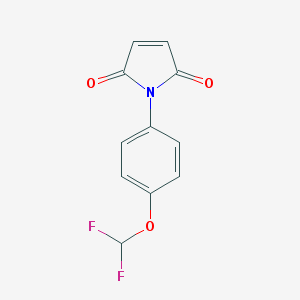 SMR000178049 SMR000178049 | C11H7F2NO3 | 239.178 | 5 / 0 | 2.0 | Yes |
| 117263 | 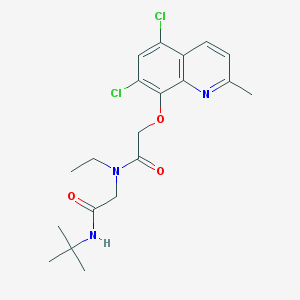 MLS001019021 MLS001019021 | C20H25Cl2N3O3 | 426.338 | 4 / 1 | 4.0 | Yes |
| 118993 | 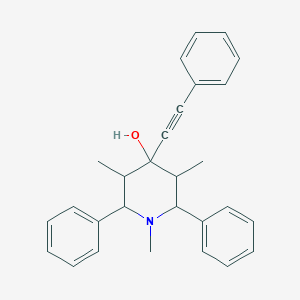 SMR000176604 SMR000176604 | C28H29NO | 395.546 | 2 / 1 | 5.7 | No |
| 119760 | 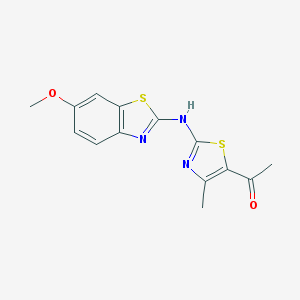 SMR000027225 SMR000027225 | C14H13N3O2S2 | 319.397 | 7 / 1 | 4.0 | Yes |
| 120121 | 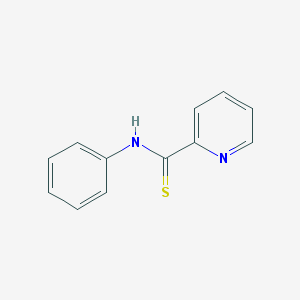 n-phenylpyridine-2-carbothioamide n-phenylpyridine-2-carbothioamide | C12H10N2S | 214.286 | 2 / 1 | 2.6 | Yes |
| 120246 | 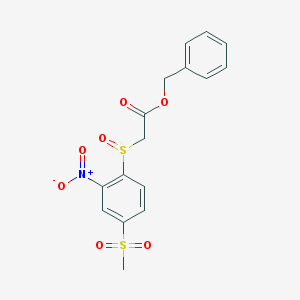 MLS000662068 MLS000662068 | C16H15NO7S2 | 397.416 | 8 / 0 | 1.6 | Yes |
| 124115 | 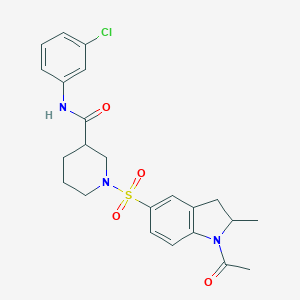 MLS001122797 MLS001122797 | C23H26ClN3O4S | 475.988 | 5 / 1 | 2.9 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218