

You can:
| Name | P2Y purinoceptor 11 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | P2RY11 |
| Synonym | P2Y11 P2Y11 receptor purinergic receptor P2Y, G-protein coupled, 11 |
| Disease | N/A |
| Length | 374 |
| Amino acid sequence | MAANVSGAKSCPANFLAAADDKLSGFQGDFLWPILVVEFLVAVASNGLALYRFSIRKQRPWHPAVVFSVQLAVSDLLCALTLPPLAAYLYPPKHWRYGEAACRLERFLFTCNLLGSVIFITCISLNRYLGIVHPFFARSHLRPKHAWAVSAAGWVLAALLAMPTLSFSHLKRPQQGAGNCSVARPEACIKCLGTADHGLAAYRAYSLVLAGLGCGLPLLLTLAAYGALGRAVLRSPGMTVAEKLRVAALVASGVALYASSYVPYHIMRVLNVDARRRWSTRCPSFADIAQATAALELGPYVGYQVMRGLMPLAFCVHPLLYMAAVPSLGCCCRHCPGYRDSWNPEDAKSTGQALPLNATAAPKPSEPQSRELSQ |
| UniProt | Q96G91 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | Q96G91 |
| 3D structure model | This predicted structure model is from GPCR-EXP Q96G91. |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL4867 |
| IUPHAR | 327 |
| DrugBank | BE0005499 |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 5501 | 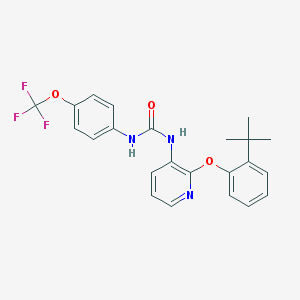 BPTU BPTU | C23H22F3N3O3 | 445.442 | 7 / 2 | 6.1 | No |
| 6822 | 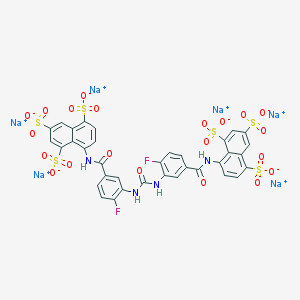 CHEMBL268784 CHEMBL268784 | C35H18F2N4Na6O21S6 | 1198.83 | 23 / 4 | N/A | No |
| 7532 | 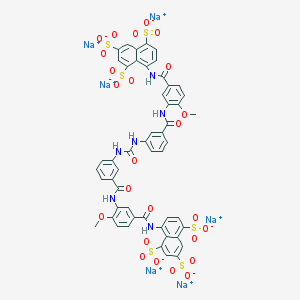 CHEMBL413910 CHEMBL413910 | C51H34N6Na6O25S6 | 1461.15 | 25 / 6 | N/A | No |
| 553329 | 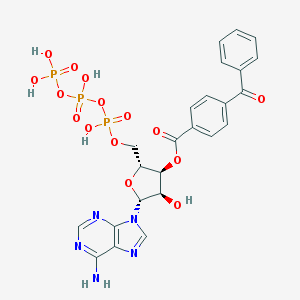 BzATP BzATP | C24H24N5O15P3 | 715.397 | 19 / 6 | -2.1 | No |
| 23817 | 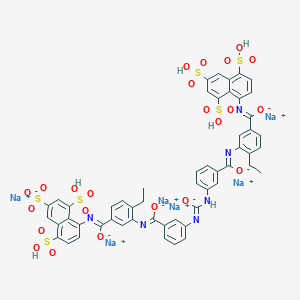 CID 136825090 CID 136825090 | C53H38N6Na6O23S6 | 1457.2 | 28 / 6 | N/A | No |
| 36209 | 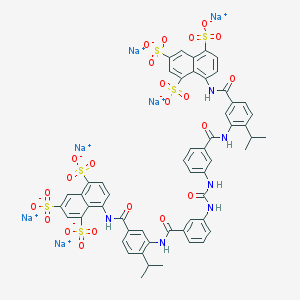 CHEMBL405500 CHEMBL405500 | C55H42N6Na6O23S6 | 1485.26 | 23 / 6 | N/A | No |
| 37986 | 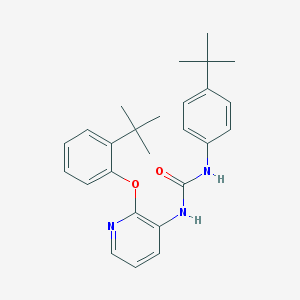 CHEMBL2333771 CHEMBL2333771 | C26H31N3O2 | 417.553 | 3 / 2 | 6.6 | No |
| 43304 | 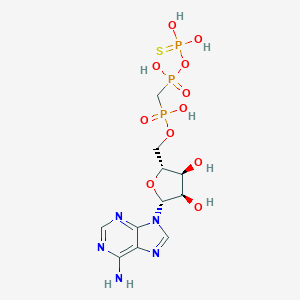 CHEMBL3289396 CHEMBL3289396 | C11H18N5O11P3S | 521.27 | 16 / 7 | -4.1 | No |
| 78871 | 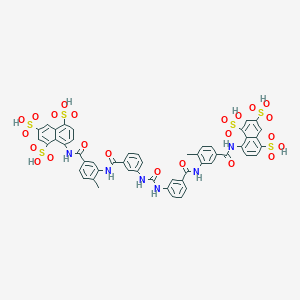 suramin suramin | C51H40N6O23S6 | 1297.26 | 23 / 12 | 1.5 | No |
| 82200 | 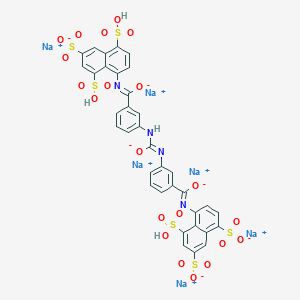 CID 136775315 CID 136775315 | C35H20N4Na6O21S6 | 1162.85 | 24 / 4 | N/A | No |
| 89818 | 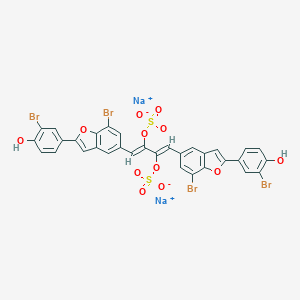 iso-iantheran A iso-iantheran A | C32H16Br4Na2O12S2 | 1022.18 | 12 / 2 | N/A | No |
| 102376 | 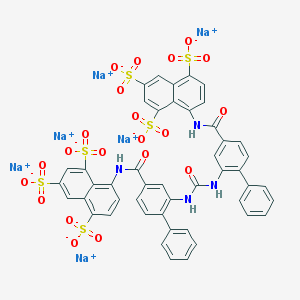 CHEMBL385555 CHEMBL385555 | C47H28N4Na6O21S6 | 1315.05 | 21 / 4 | N/A | No |
| 553768 | 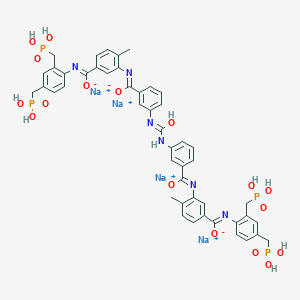 J-000176 J-000176 | C47H44N6Na4O17P4 | 1180.75 | 22 / 10 | N/A | No |
| 476598 | 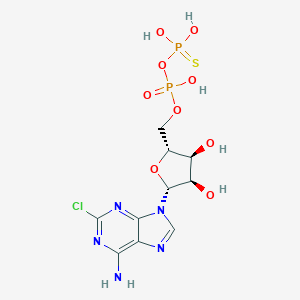 CHEMBL3634186 CHEMBL3634186 | C10H14ClN5O9P2S | 477.706 | 14 / 6 | -1.9 | No |
| 111326 | 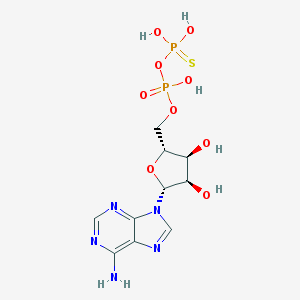 Adpbetas Adpbetas | C10H15N5O9P2S | 443.264 | 14 / 6 | -2.9 | No |
| 118337 | 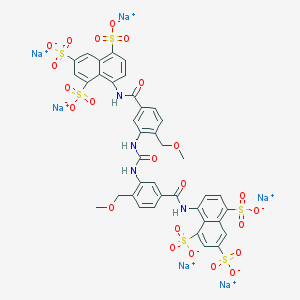 CHEMBL414577 CHEMBL414577 | C39H28N4Na6O23S6 | 1250.96 | 23 / 4 | N/A | No |
| 480928 | 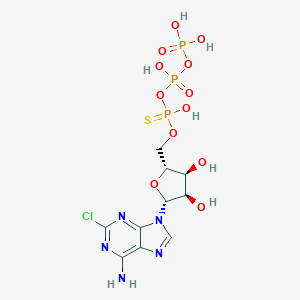 CHEMBL3634184 CHEMBL3634184 | C10H15ClN5O12P3S | 557.684 | 17 / 7 | -3.0 | No |
| 157237 | 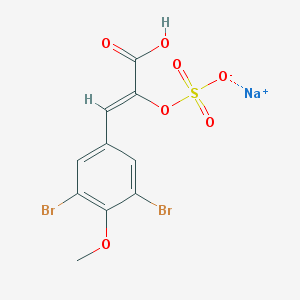 CHEMBL240925 CHEMBL240925 | C10H7Br2NaO7S | 454.017 | 7 / 1 | N/A | N/A |
| 157773 | 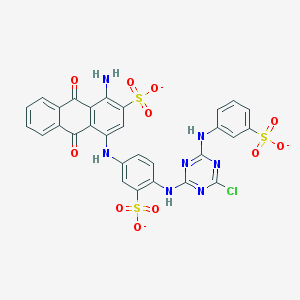 Basilen Blue Basilen Blue | C29H17ClN7O11S3-3 | 771.123 | 18 / 4 | 4.1 | No |
| 554072 | 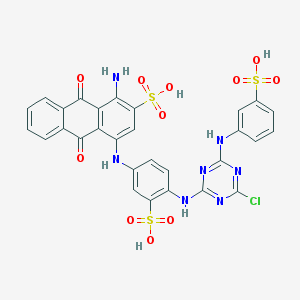 Affi gel blue Affi gel blue | C29H20ClN7O11S3 | 774.147 | 18 / 7 | 4.4 | No |
| 483063 | 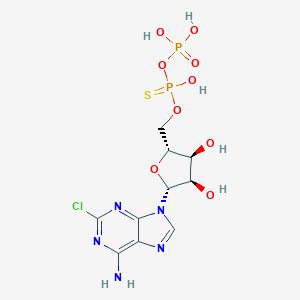 CHEMBL3634183 CHEMBL3634183 | C10H14ClN5O9P2S | 477.706 | 14 / 6 | -1.9 | No |
| 171184 | 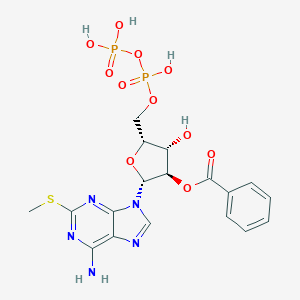 CHEMBL2386490 CHEMBL2386490 | C18H21N5O11P2S | 577.398 | 16 / 5 | -1.6 | No |
| 486847 | 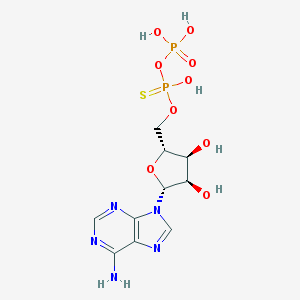 CHEMBL575257 CHEMBL575257 | C10H15N5O9P2S | 443.264 | 14 / 6 | -2.9 | No |
| 214762 | 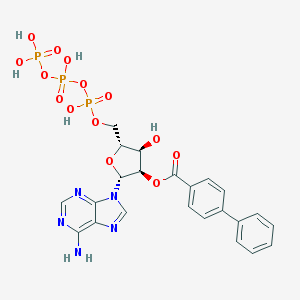 CHEMBL339386 CHEMBL339386 | C23H24N5O14P3 | 687.387 | 18 / 6 | -1.9 | No |
| 220435 | 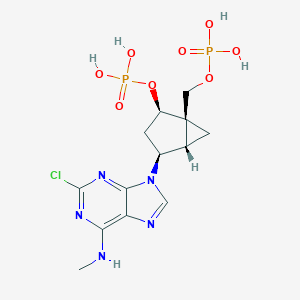 CHEMBL3085531 CHEMBL3085531 | C13H18ClN5O8P2 | 469.712 | 12 / 5 | -1.6 | No |
| 554400 | 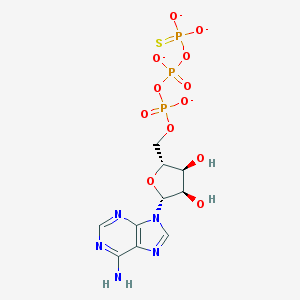 ATPgammaS ATPgammaS | C10H12N5O12P3S-4 | 519.21 | 17 / 3 | -4.3 | No |
| 226524 | 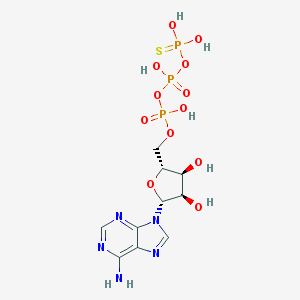 gamma-Thio-ATP gamma-Thio-ATP | C10H16N5O12P3S | 523.242 | 17 / 7 | -4.0 | No |
| 240383 | 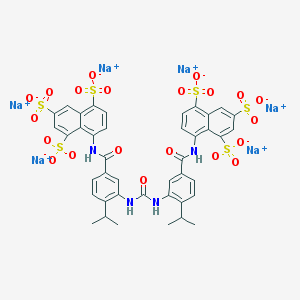 AC1L9PN3 AC1L9PN3 | C41H32N4Na6O21S6 | 1247.01 | 21 / 4 | N/A | No |
| 258405 | 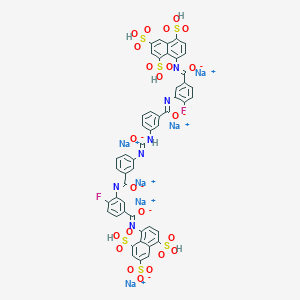 CID 136776946 CID 136776946 | C49H28F2N6Na6O23S6 | 1437.08 | 30 / 6 | N/A | No |
| 258878 | 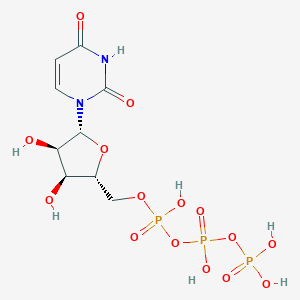 uridine 5'-triphosphate uridine 5'-triphosphate | C9H15N2O15P3 | 484.139 | 15 / 7 | -5.8 | No |
| 261908 | 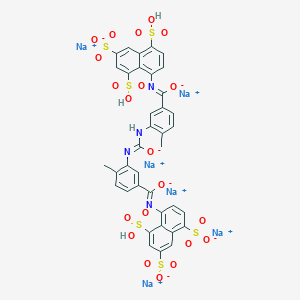 NF-058 NF-058 | C37H24N4Na6O21S6 | 1190.91 | 24 / 4 | N/A | No |
| 263499 | 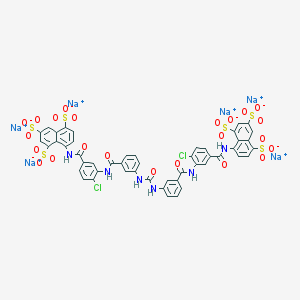 CHEMBL413180 CHEMBL413180 | C49H28Cl2N6Na6O23S6 | 1469.98 | 23 / 6 | N/A | No |
| 267058 | 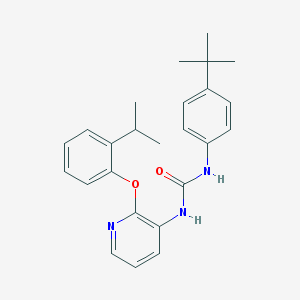 CHEMBL2333772 CHEMBL2333772 | C25H29N3O2 | 403.526 | 3 / 2 | 6.1 | No |
| 271348 | 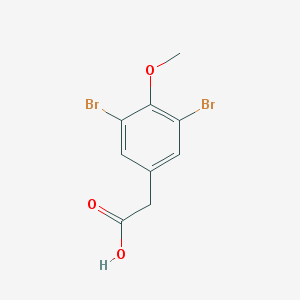 3,5-dibromo-4-methoxy-phenylacetic acid 3,5-dibromo-4-methoxy-phenylacetic acid | C9H8Br2O3 | 323.968 | 3 / 1 | 2.7 | Yes |
| 499146 | 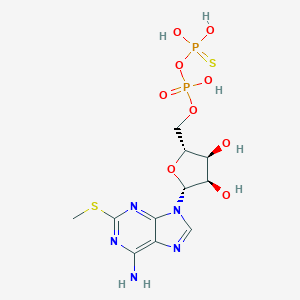 CHEMBL3634185 CHEMBL3634185 | C11H17N5O9P2S2 | 489.351 | 15 / 6 | -2.0 | No |
| 554826 | 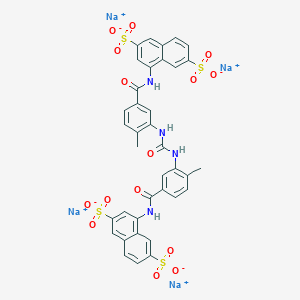 NF 340 NF 340 | C37H26N4Na4O15S4 | 986.827 | 15 / 4 | N/A | No |
| 502310 | 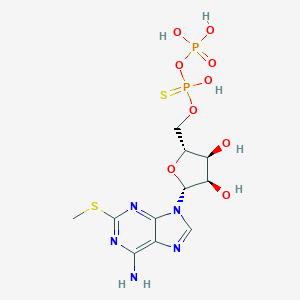 CHEMBL3634182 CHEMBL3634182 | C11H17N5O9P2S2 | 489.351 | 15 / 6 | -2.0 | No |
| 323923 | 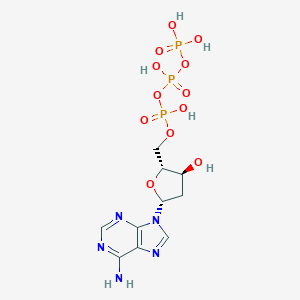 dATP dATP | C10H16N5O12P3 | 491.182 | 16 / 6 | -4.4 | No |
| 334225 | 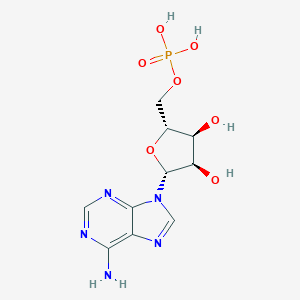 adenosine 5'-monophosphate adenosine 5'-monophosphate | C10H14N5O7P | 347.224 | 11 / 5 | -3.5 | No |
| 334479 | 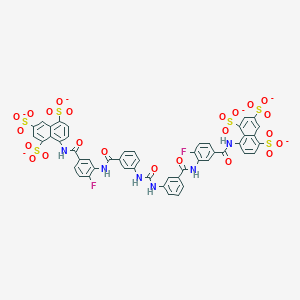 CHEMBL413145 CHEMBL413145 | C49H28F2N6O23S6-6 | 1299.14 | 25 / 6 | 0.5 | No |
| 554942 | 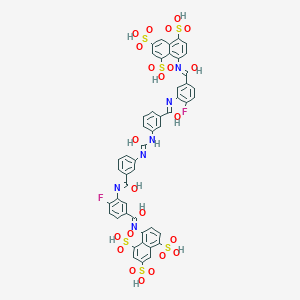 CID 136776947 CID 136776947 | C49H34F2N6O23S6 | 1305.19 | 30 / 12 | 3.7 | No |
| 335723 | 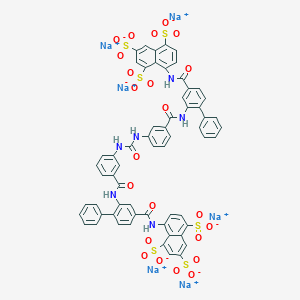 CHEMBL266646 CHEMBL266646 | C61H38N6Na6O23S6 | 1553.29 | 23 / 6 | N/A | No |
| 364554 | 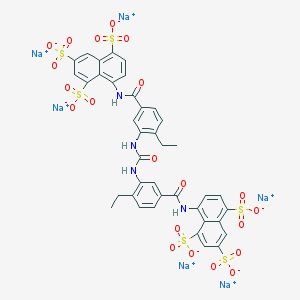 CHEMBL408014 CHEMBL408014 | C39H28N4Na6O21S6 | 1218.96 | 21 / 4 | N/A | No |
| 367908 | 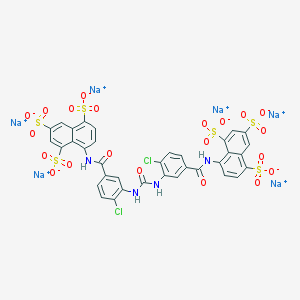 CHEMBL265513 CHEMBL265513 | C35H18Cl2N4Na6O21S6 | 1231.73 | 21 / 4 | N/A | No |
| 368120 | 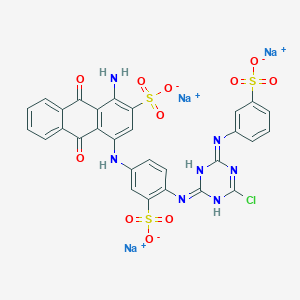 CID 136908877 CID 136908877 | C29H17ClN7Na3O11S3 | 840.092 | 15 / 4 | N/A | No |
| 368336 | 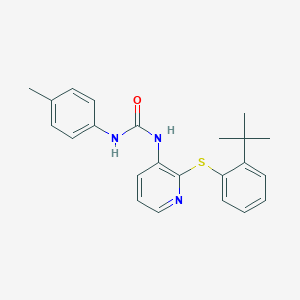 CHEMBL2333767 CHEMBL2333767 | C23H25N3OS | 391.533 | 3 / 2 | 5.9 | No |
| 376226 | 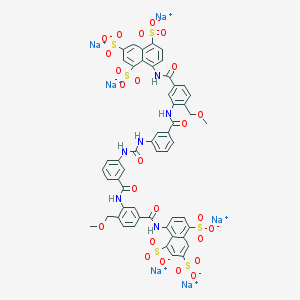 CHEMBL409304 CHEMBL409304 | C53H38N6Na6O25S6 | 1489.2 | 25 / 6 | N/A | No |
| 378618 | 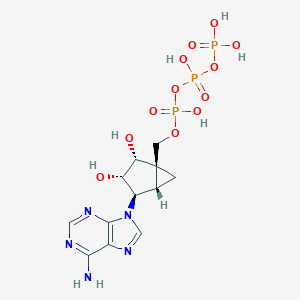 CHEMBL2111533 CHEMBL2111533 | C12H18N5O12P3 | 517.22 | 16 / 7 | -5.3 | No |
| 378622 | 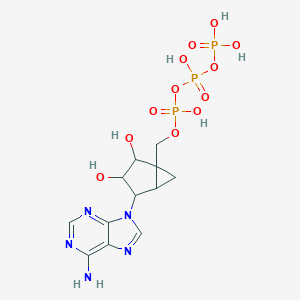 CHEMBL302077 CHEMBL302077 | C12H18N5O12P3 | 517.22 | 16 / 7 | -5.3 | No |
| 380758 | 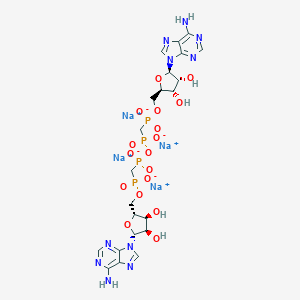 CHEMBL1802096 CHEMBL1802096 | C22H28N10Na4O17P4 | 920.373 | 25 / 6 | N/A | No |
| 382530 | 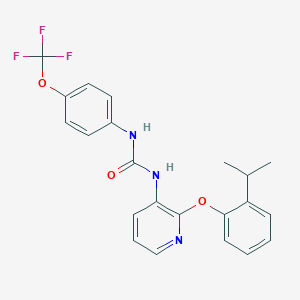 CHEMBL2333773 CHEMBL2333773 | C22H20F3N3O3 | 431.415 | 7 / 2 | 5.6 | No |
| 394145 | 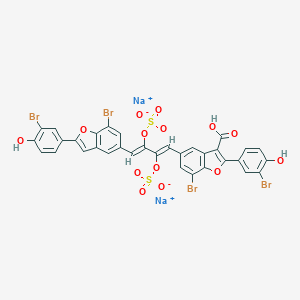 8-carboxy-iso-iantheran A 8-carboxy-iso-iantheran A | C33H16Br4Na2O14S2 | 1066.19 | 14 / 3 | N/A | No |
| 396471 | 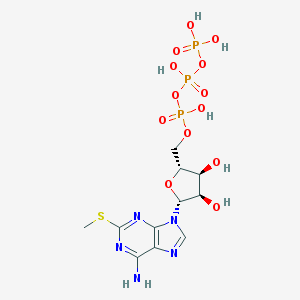 2-Mesatp 2-Mesatp | C11H18N5O13P3S | 553.268 | 18 / 7 | -4.9 | No |
| 399782 | 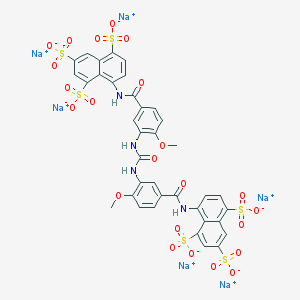 CHEMBL412037 CHEMBL412037 | C37H24N4Na6O23S6 | 1222.9 | 23 / 4 | N/A | No |
| 400824 | 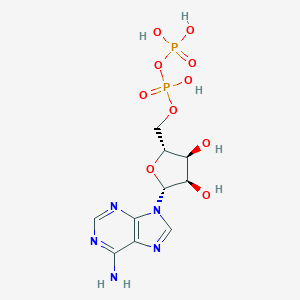 Adenosine 5'-diphosphate Adenosine 5'-diphosphate | C10H15N5O10P2 | 427.203 | 14 / 6 | -4.6 | No |
| 430564 | 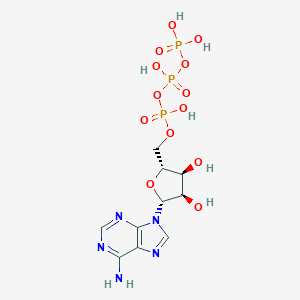 Adenosine triphosphate Adenosine triphosphate | C10H16N5O13P3 | 507.181 | 17 / 7 | -5.7 | No |
| 431361 | 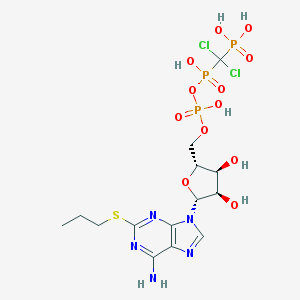 AR-C67085MX AR-C67085MX | C14H22Cl2N5O12P3S | 648.234 | 17 / 7 | -2.9 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218