

You can:
| Name | Mas-related G-protein coupled receptor member X1 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | MRGPRX1 |
| Synonym | Sensory neuron-specific G-protein coupled receptor 4 Sensory neuron-specific G-protein coupled receptor 3/4 MRGX1 MRGPRX1 Mrgprc11 [ Show all ] |
| Disease | N/A |
| Length | 322 |
| Amino acid sequence | MDPTISTLDTELTPINGTEETLCYKQTLSLTVLTCIVSLVGLTGNAVVLWLLGCRMRRNAFSIYILNLAAADFLFLSGRLIYSLLSFISIPHTISKILYPVMMFSYFAGLSFLSAVSTERCLSVLWPIWYRCHRPTHLSAVVCVLLWALSLLRSILEWMLCGFLFSGADSAWCQTSDFITVAWLIFLCVVLCGSSLVLLIRILCGSRKIPLTRLYVTILLTVLVFLLCGLPFGIQFFLFLWIHVDREVLFCHVHLVSIFLSALNSSANPIIYFFVGSFRQRQNRQNLKLVLQRALQDASEVDEGGGQLPEEILELSGSRLEQ |
| UniProt | Q96LB2 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | Q96LB2 |
| 3D structure model | This predicted structure model is from GPCR-EXP Q96LB2. |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL5850 |
| IUPHAR | 156 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 521495 | 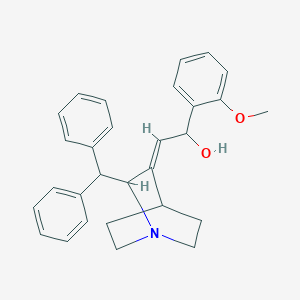 CHEMBL3730717 CHEMBL3730717 | C29H31NO2 | 425.572 | 3 / 1 | 4.9 | Yes |
| 2189 | 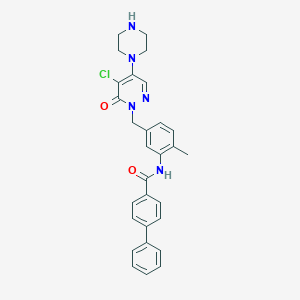 CHEMBL458002 CHEMBL458002 | C29H28ClN5O2 | 514.026 | 5 / 2 | 4.3 | No |
| 5936 | 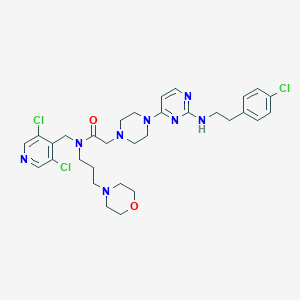 CHEMBL1762707 CHEMBL1762707 | C31H39Cl3N8O2 | 662.057 | 9 / 1 | 4.6 | No |
| 442531 | 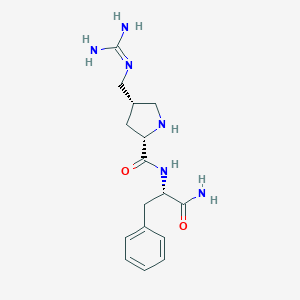 CHEMBL3343991 CHEMBL3343991 | C16H24N6O2 | 332.408 | 4 / 5 | -1.0 | Yes |
| 29144 | 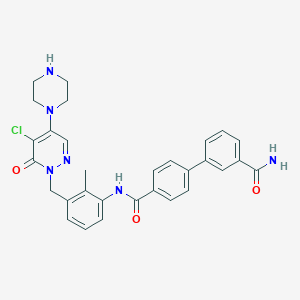 CHEMBL474474 CHEMBL474474 | C30H29ClN6O3 | 557.051 | 6 / 3 | 3.2 | No |
| 522498 | 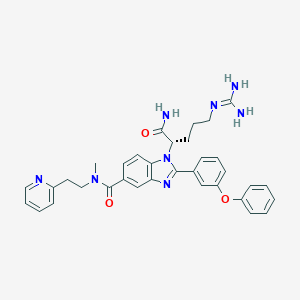 CHEMBL3719266 CHEMBL3719266 | C34H36N8O3 | 604.715 | 6 / 3 | 3.3 | No |
| 33689 | 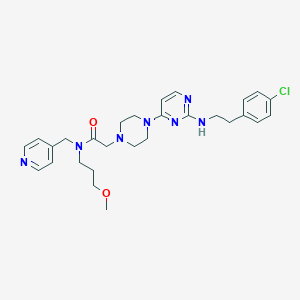 CHEMBL1762702 CHEMBL1762702 | C28H36ClN7O2 | 538.093 | 8 / 1 | 3.5 | No |
| 40890 | 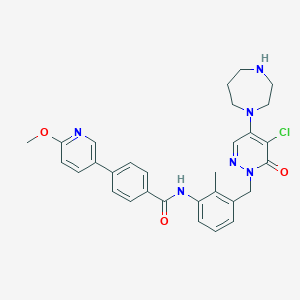 CHEMBL475077 CHEMBL475077 | C30H31ClN6O3 | 559.067 | 7 / 2 | 3.9 | No |
| 443622 | 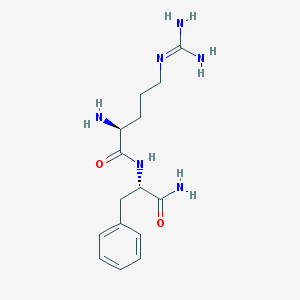 Rfamide Rfamide | C15H24N6O2 | 320.397 | 4 / 5 | -2.0 | Yes |
| 523186 | 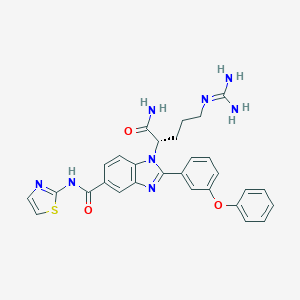 CHEMBL3718857 CHEMBL3718857 | C29H28N8O3S | 568.656 | 7 / 4 | 3.1 | No |
| 523548 | 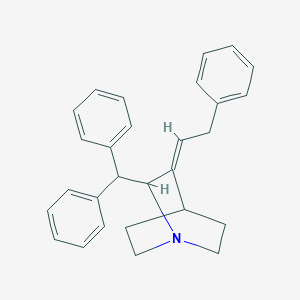 CHEMBL3731811 CHEMBL3731811 | C28H29N | 379.547 | 1 / 0 | 6.1 | No |
| 523647 | 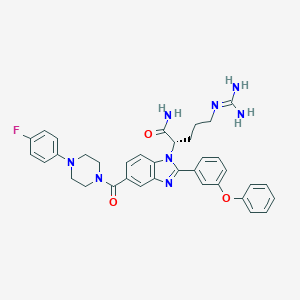 CHEMBL3717645 CHEMBL3717645 | C36H37FN8O3 | 648.743 | 7 / 3 | 4.0 | No |
| 559594 | 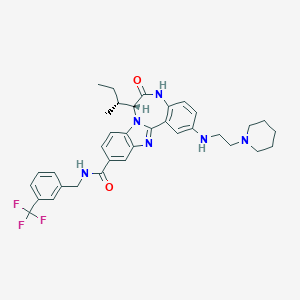 CHEMBL505263 CHEMBL505263 | C35H39F3N6O2 | 632.732 | 8 / 3 | 6.4 | No |
| 559597 | 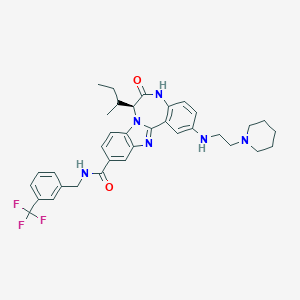 CHEMBL505263 CHEMBL505263 | C35H39F3N6O2 | 632.732 | 8 / 3 | 6.4 | No |
| 77223 | 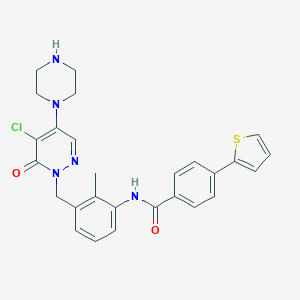 CHEMBL477431 CHEMBL477431 | C27H26ClN5O2S | 520.048 | 6 / 2 | 4.0 | No |
| 559862 | 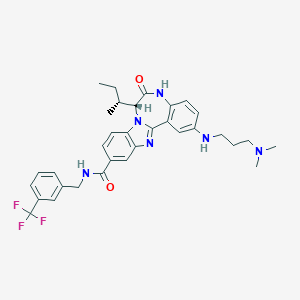 CHEMBL502132 CHEMBL502132 | C33H37F3N6O2 | 606.694 | 8 / 3 | 5.9 | No |
| 559864 | 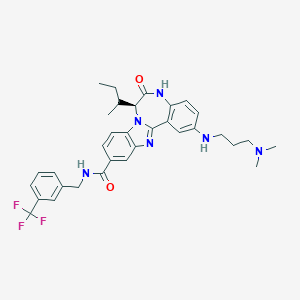 CHEMBL502132 CHEMBL502132 | C33H37F3N6O2 | 606.694 | 8 / 3 | 5.9 | No |
| 560176 | 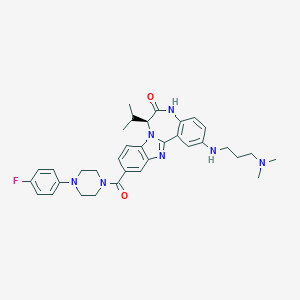 CHEMBL501027 CHEMBL501027 | C34H40FN7O2 | 597.739 | 7 / 2 | 5.1 | No |
| 560178 | 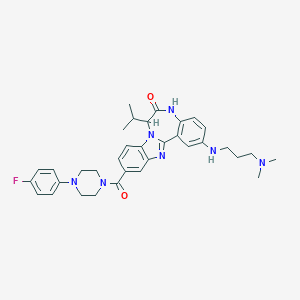 CHEMBL501027 CHEMBL501027 | C34H40FN7O2 | 597.739 | 7 / 2 | 5.1 | No |
| 524418 | 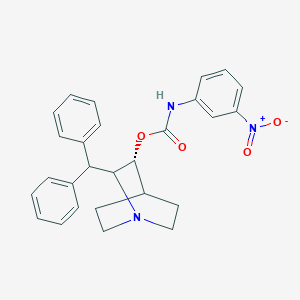 CHEMBL3732603 CHEMBL3732603 | C27H27N3O4 | 457.53 | 5 / 1 | 5.4 | No |
| 560403 | 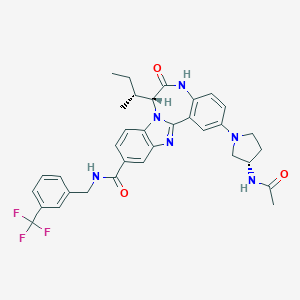 CHEMBL503202 CHEMBL503202 | C34H35F3N6O3 | 632.688 | 8 / 3 | 5.2 | No |
| 560404 | 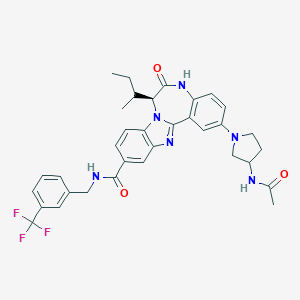 CHEMBL503202 CHEMBL503202 | C34H35F3N6O3 | 632.688 | 8 / 3 | 5.2 | No |
| 99762 | 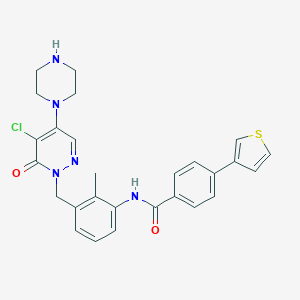 CHEMBL477432 CHEMBL477432 | C27H26ClN5O2S | 520.048 | 6 / 2 | 4.0 | No |
| 445678 | 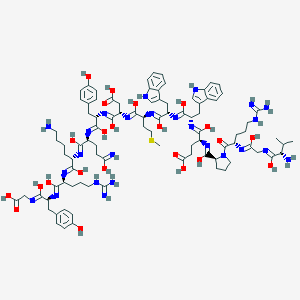 BAM(8-22) BAM(8-22) | C91H127N25O23S | 1971.23 | 42 / 30 | 1.4 | No |
| 560509 | 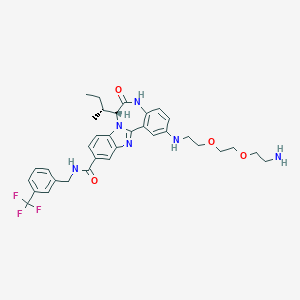 CHEMBL466116 CHEMBL466116 | C34H39F3N6O4 | 652.719 | 10 / 4 | 4.3 | No |
| 560510 | 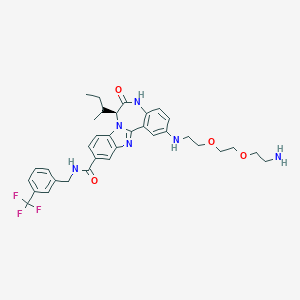 CHEMBL466116 CHEMBL466116 | C34H39F3N6O4 | 652.719 | 10 / 4 | 4.3 | No |
| 524562 | 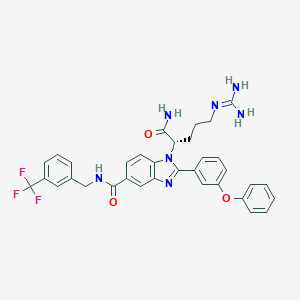 CHEMBL3717941 CHEMBL3717941 | C34H32F3N7O3 | 643.671 | 8 / 4 | 4.5 | No |
| 524642 | 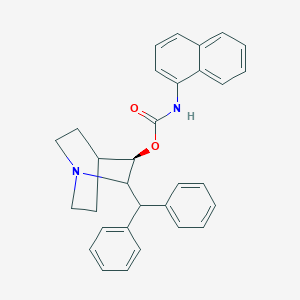 CHEMBL3730405 CHEMBL3730405 | C31H30N2O2 | 462.593 | 3 / 1 | 6.9 | No |
| 107377 | 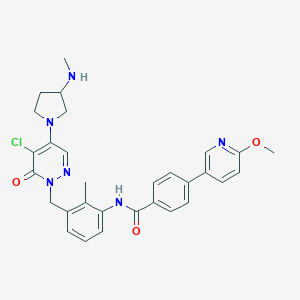 CHEMBL474470 CHEMBL474470 | C30H31ClN6O3 | 559.067 | 7 / 2 | 4.0 | No |
| 524769 | 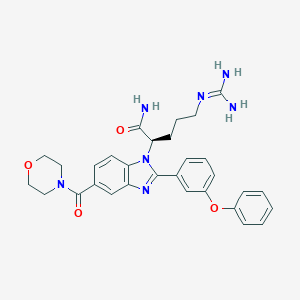 CHEMBL3719296 CHEMBL3719296 | C30H33N7O4 | 555.639 | 6 / 3 | 2.0 | No |
| 126953 | 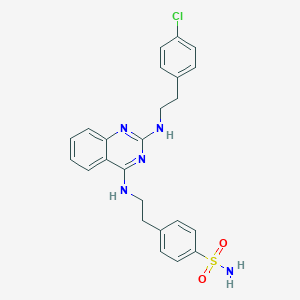 CHEMBL1762695 CHEMBL1762695 | C24H24ClN5O2S | 481.999 | 7 / 3 | 5.3 | No |
| 132276 | 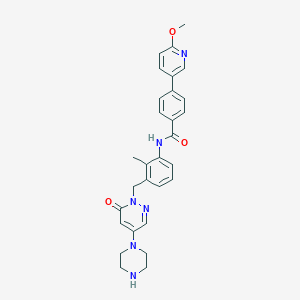 1103458-91-8 1103458-91-8 | C29H30N6O3 | 510.598 | 7 / 2 | 2.7 | No |
| 525366 | 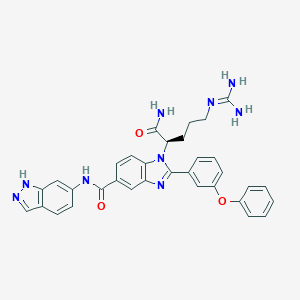 CHEMBL3716995 CHEMBL3716995 | C33H31N9O3 | 601.671 | 6 / 5 | 3.3 | No |
| 525388 | 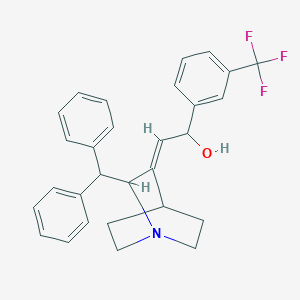 CHEMBL3730500 CHEMBL3730500 | C29H28F3NO | 463.544 | 5 / 1 | 5.9 | No |
| 133866 | 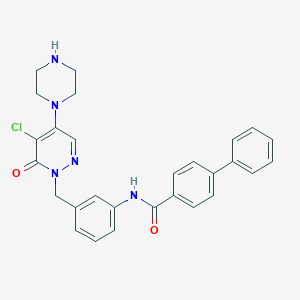 CHEMBL458001 CHEMBL458001 | C28H26ClN5O2 | 499.999 | 5 / 2 | 3.9 | Yes |
| 133944 | 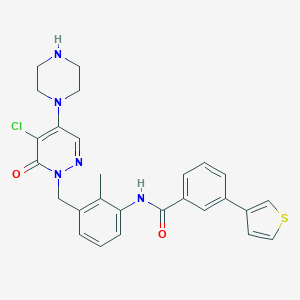 CHEMBL476619 CHEMBL476619 | C27H26ClN5O2S | 520.048 | 6 / 2 | 4.0 | No |
| 142470 | 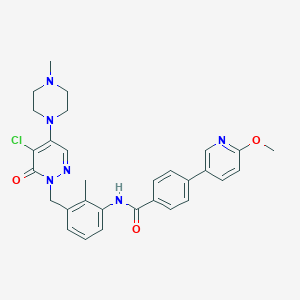 CHEMBL475282 CHEMBL475282 | C30H31ClN6O3 | 559.067 | 7 / 1 | 4.0 | No |
| 447648 | 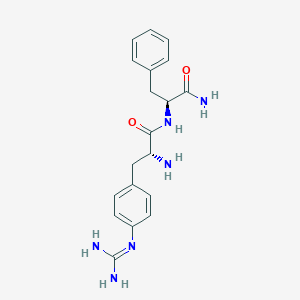 CHEMBL3343994 CHEMBL3343994 | C19H24N6O2 | 368.441 | 4 / 5 | -0.4 | Yes |
| 447651 | 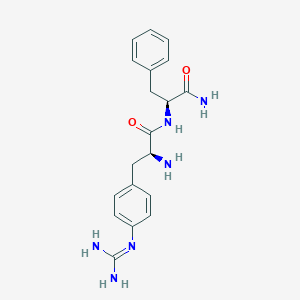 CHEMBL3343993 CHEMBL3343993 | C19H24N6O2 | 368.441 | 4 / 5 | -0.4 | Yes |
| 182220 | 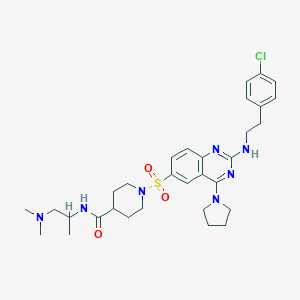 CHEMBL1762693 CHEMBL1762693 | C31H42ClN7O3S | 628.233 | 9 / 2 | 4.8 | No |
| 184565 | 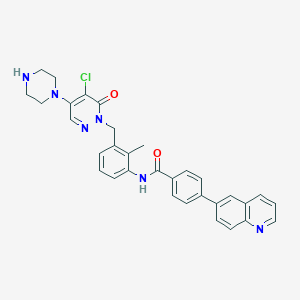 CHEMBL449189 CHEMBL449189 | C32H29ClN6O2 | 565.074 | 6 / 2 | 4.6 | No |
| 200942 | 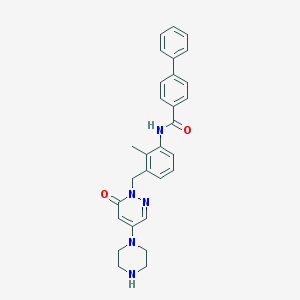 CHEMBL477008 CHEMBL477008 | C29H29N5O2 | 479.584 | 5 / 2 | 3.5 | Yes |
| 206159 | 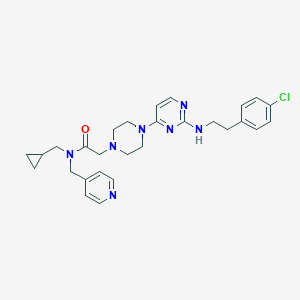 CHEMBL1762704 CHEMBL1762704 | C28H34ClN7O | 520.078 | 7 / 1 | 4.1 | No |
| 206204 | 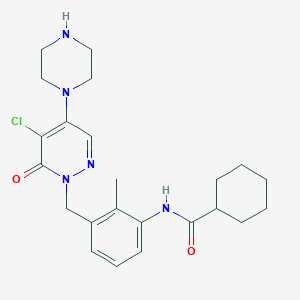 CHEMBL456481 CHEMBL456481 | C23H30ClN5O2 | 443.976 | 5 / 2 | 3.1 | Yes |
| 208646 | 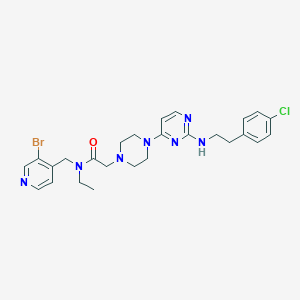 CHEMBL1762705 CHEMBL1762705 | C26H31BrClN7O | 572.936 | 7 / 1 | 4.4 | No |
| 216658 | 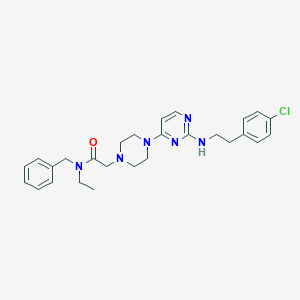 CHEMBL1762706 CHEMBL1762706 | C27H33ClN6O | 493.052 | 6 / 1 | 4.8 | Yes |
| 227233 | 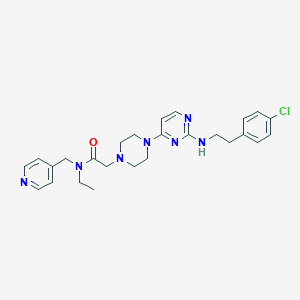 CHEMBL1762703 CHEMBL1762703 | C26H32ClN7O | 494.04 | 7 / 1 | 3.7 | Yes |
| 236254 | 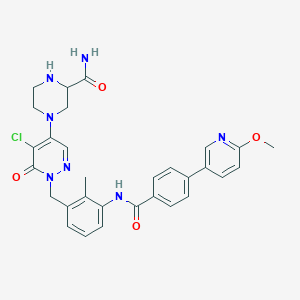 CHEMBL500240 CHEMBL500240 | C30H30ClN7O4 | 588.065 | 8 / 3 | 2.5 | No |
| 240899 | 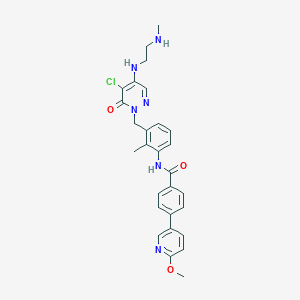 CHEMBL476627 CHEMBL476627 | C28H29ClN6O3 | 533.029 | 7 / 3 | 3.6 | No |
| 250610 | 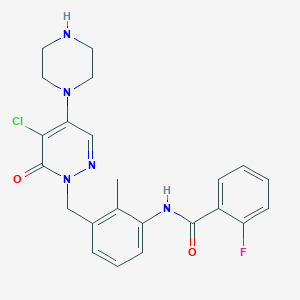 CHEMBL514267 CHEMBL514267 | C23H23ClFN5O2 | 455.918 | 6 / 2 | 2.8 | Yes |
| 251834 | 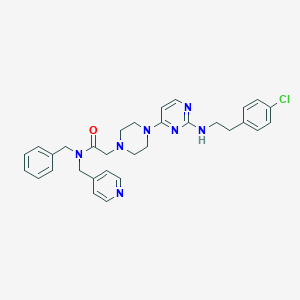 CHEMBL1762699 CHEMBL1762699 | C31H34ClN7O | 556.111 | 7 / 1 | 4.8 | No |
| 253731 | 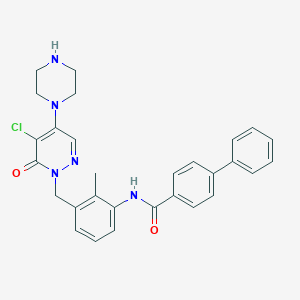 CHEMBL458220 CHEMBL458220 | C29H28ClN5O2 | 514.026 | 5 / 2 | 4.3 | No |
| 528779 | 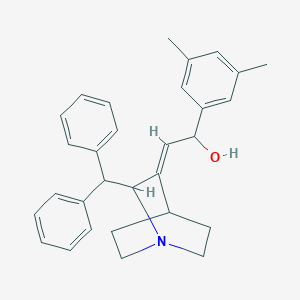 CHEMBL3730179 CHEMBL3730179 | C30H33NO | 423.6 | 2 / 1 | 5.7 | No |
| 528783 | 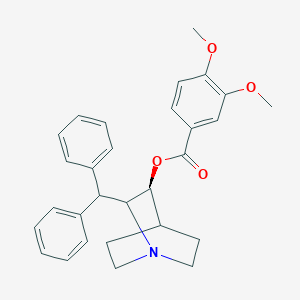 CHEMBL3727438 CHEMBL3727438 | C29H31NO4 | 457.57 | 5 / 0 | 5.8 | No |
| 256237 | 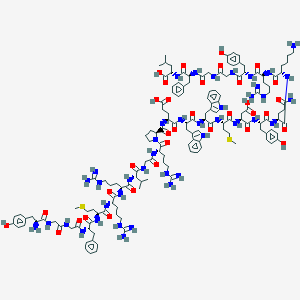 78355-50-7 78355-50-7 | C147H207N41O34S2 | 3156.65 | 42 / 46 | -5.2 | No |
| 260548 | 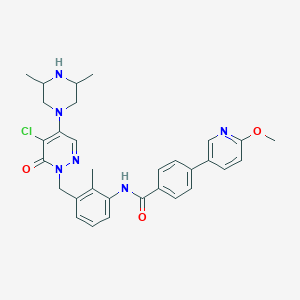 CHEMBL450183 CHEMBL450183 | C31H33ClN6O3 | 573.094 | 7 / 2 | 4.4 | No |
| 529225 | 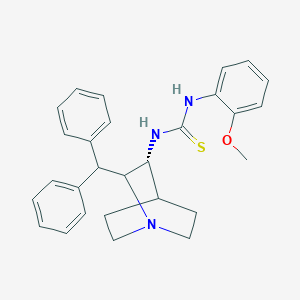 CHEMBL3728009 CHEMBL3728009 | C28H31N3OS | 457.636 | 3 / 2 | 5.6 | No |
| 279294 | 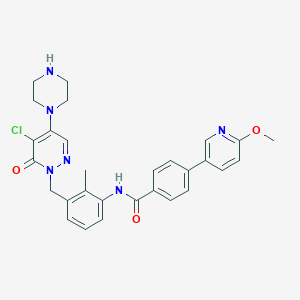 CHEMBL474473 CHEMBL474473 | C29H29ClN6O3 | 545.04 | 7 / 2 | 3.5 | No |
| 289723 | 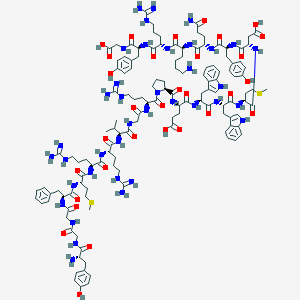 Bam 22P Bam 22P | C130H184N38O31S2 | 2839.26 | 39 / 43 | -7.0 | No |
| 289724 | 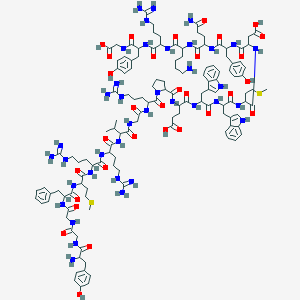 Bam-22P Bam-22P | C130H184N38O31S2 | 2839.26 | 39 / 43 | -7.0 | No |
| 314947 | 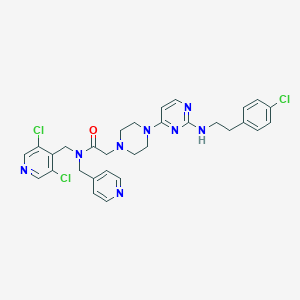 CHEMBL1762700 CHEMBL1762700 | C30H31Cl3N8O | 625.983 | 8 / 1 | 5.0 | No |
| 530409 | 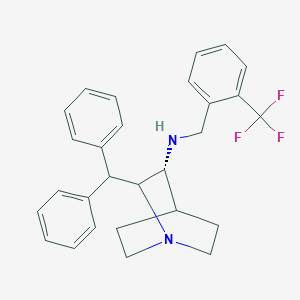 CHEMBL3728505 CHEMBL3728505 | C28H29F3N2 | 450.549 | 5 / 1 | 6.3 | No |
| 322256 | 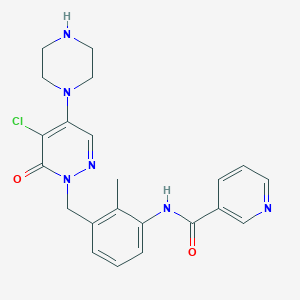 CHEMBL457788 CHEMBL457788 | C22H23ClN6O2 | 438.916 | 6 / 2 | 1.6 | Yes |
| 530810 | 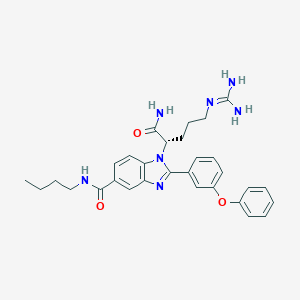 CHEMBL3715864 CHEMBL3715864 | C30H35N7O3 | 541.656 | 5 / 4 | 3.4 | No |
| 530834 | 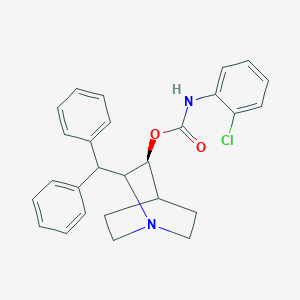 CHEMBL3733177 CHEMBL3733177 | C27H27ClN2O2 | 446.975 | 3 / 1 | 6.2 | No |
| 530837 | 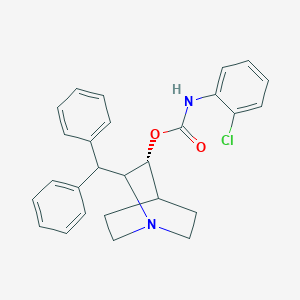 CHEMBL3730599 CHEMBL3730599 | C27H27ClN2O2 | 446.975 | 3 / 1 | 6.2 | No |
| 333016 | 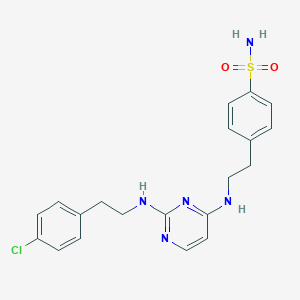 CHEMBL1762696 CHEMBL1762696 | C20H22ClN5O2S | 431.939 | 7 / 3 | 3.9 | Yes |
| 336121 | 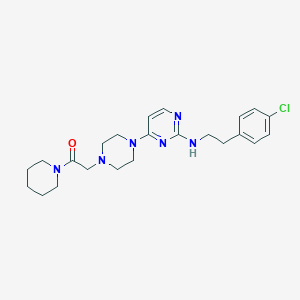 CHEMBL1762697 CHEMBL1762697 | C23H31ClN6O | 442.992 | 6 / 1 | 3.7 | Yes |
| 455207 | 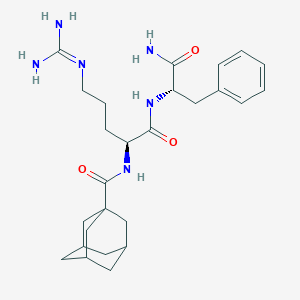 RF 9 RF 9 | C26H38N6O3 | 482.629 | 4 / 5 | 1.4 | Yes |
| 567947 | 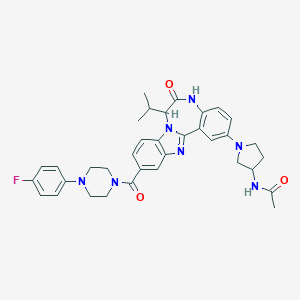 CHEMBL504032 CHEMBL504032 | C35H38FN7O3 | 623.733 | 7 / 2 | 4.3 | No |
| 567948 | 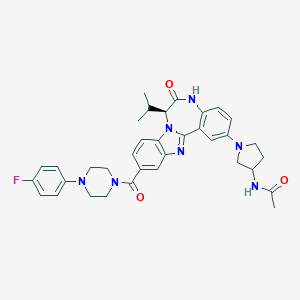 CHEMBL504032 CHEMBL504032 | C35H38FN7O3 | 623.733 | 7 / 2 | 4.3 | No |
| 342309 | 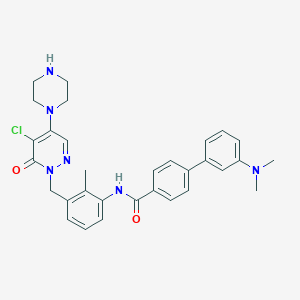 CHEMBL474272 CHEMBL474272 | C31H33ClN6O2 | 557.095 | 6 / 2 | 4.4 | No |
| 531139 | 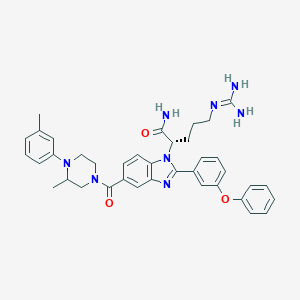 CHEMBL3716029 CHEMBL3716029 | C38H42N8O3 | 658.807 | 6 / 3 | 4.7 | No |
| 343874 | 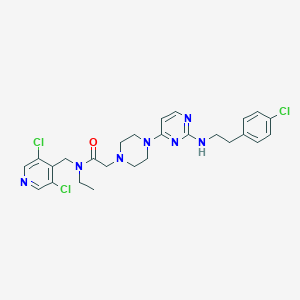 CHEMBL1760000 CHEMBL1760000 | C26H30Cl3N7O | 562.924 | 7 / 1 | 4.9 | No |
| 531243 | 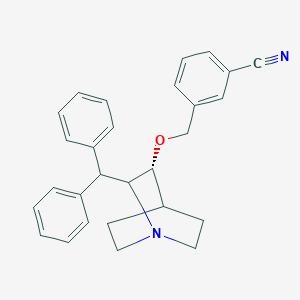 CHEMBL3729794 CHEMBL3729794 | C28H28N2O | 408.545 | 3 / 0 | 5.4 | No |
| 353888 | 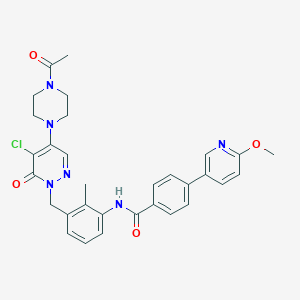 CHEMBL499944 CHEMBL499944 | C31H31ClN6O4 | 587.077 | 7 / 1 | 3.5 | No |
| 531822 | 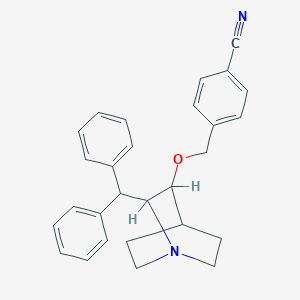 CHEMBL145103 CHEMBL145103 | C28H28N2O | 408.545 | 3 / 0 | 5.4 | No |
| 531854 | 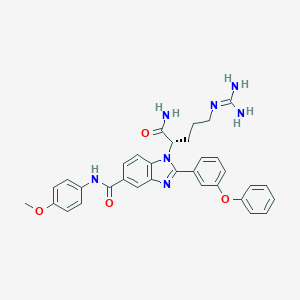 CHEMBL3716921 CHEMBL3716921 | C33H33N7O4 | 591.672 | 6 / 4 | 3.7 | No |
| 532044 | 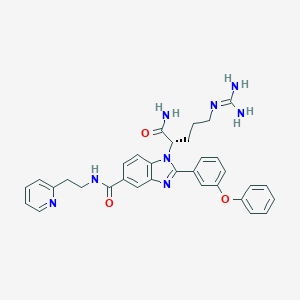 CHEMBL3717681 CHEMBL3717681 | C33H34N8O3 | 590.688 | 6 / 4 | 3.1 | No |
| 555154 | 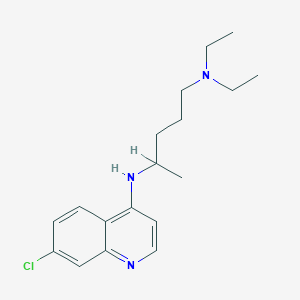 chloroquine chloroquine | C18H26ClN3 | 319.877 | 3 / 1 | 4.6 | Yes |
| 378364 | 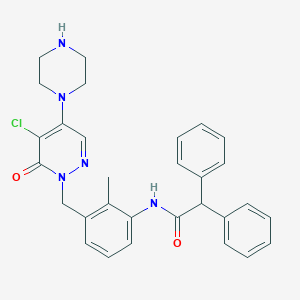 CHEMBL457789 CHEMBL457789 | C30H30ClN5O2 | 528.053 | 5 / 2 | 4.3 | No |
| 532326 | 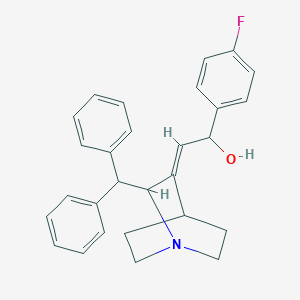 CHEMBL3727861 CHEMBL3727861 | C28H28FNO | 413.536 | 3 / 1 | 5.1 | No |
| 457077 | 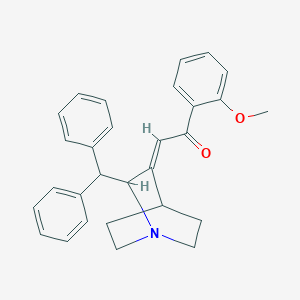 887109-77-5 887109-77-5 | C29H29NO2 | 423.556 | 3 / 0 | 5.3 | No |
| 388439 | 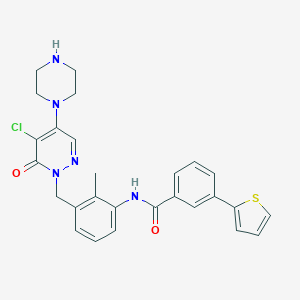 CHEMBL477643 CHEMBL477643 | C27H26ClN5O2S | 520.048 | 6 / 2 | 4.0 | No |
| 532547 | 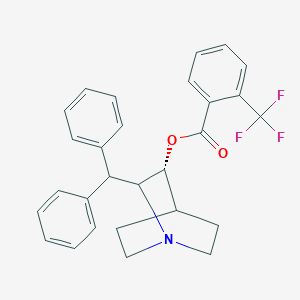 CHEMBL3731844 CHEMBL3731844 | C28H26F3NO2 | 465.516 | 6 / 0 | 6.8 | No |
| 405930 | 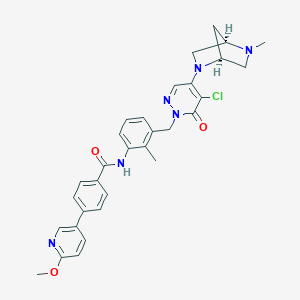 CHEMBL448975 CHEMBL448975 | C31H31ClN6O3 | 571.078 | 7 / 1 | 4.3 | No |
| 458060 | 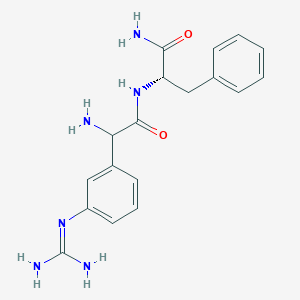 CHEMBL3343992 CHEMBL3343992 | C18H22N6O2 | 354.414 | 4 / 5 | -0.3 | Yes |
| 413377 | 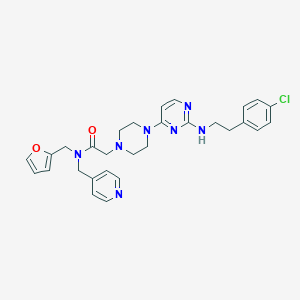 CHEMBL1762698 CHEMBL1762698 | C29H32ClN7O2 | 546.072 | 8 / 1 | 3.9 | No |
| 421317 | 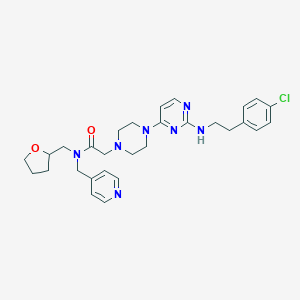 CHEMBL1762701 CHEMBL1762701 | C29H36ClN7O2 | 550.104 | 8 / 1 | 3.7 | No |
| 421435 | 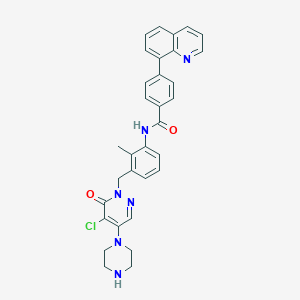 CHEMBL451657 CHEMBL451657 | C32H29ClN6O2 | 565.074 | 6 / 2 | 4.6 | No |
| 458450 | 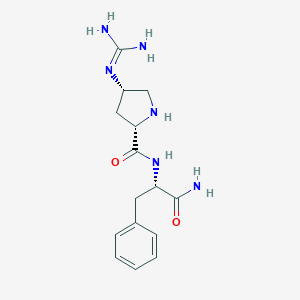 CHEMBL3343987 CHEMBL3343987 | C15H22N6O2 | 318.381 | 4 / 5 | -1.3 | Yes |
| 422881 | 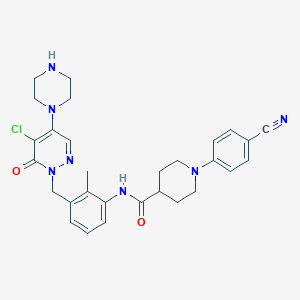 CHEMBL517029 CHEMBL517029 | C29H32ClN7O2 | 546.072 | 7 / 2 | 3.0 | No |
| 570572 | 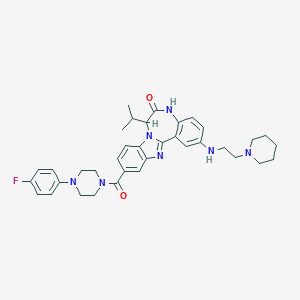 CHEMBL474761 CHEMBL474761 | C36H42FN7O2 | 623.777 | 7 / 2 | 5.6 | No |
| 570573 | 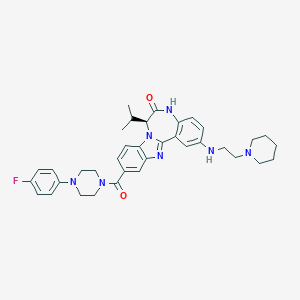 CHEMBL474761 CHEMBL474761 | C36H42FN7O2 | 623.777 | 7 / 2 | 5.6 | No |
| 458962 | 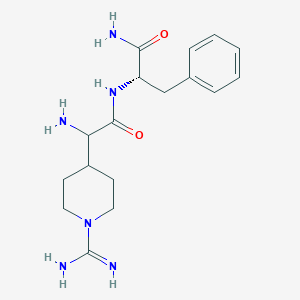 CHEMBL3343988 CHEMBL3343988 | C17H26N6O2 | 346.435 | 4 / 5 | -0.6 | Yes |
| 436806 | 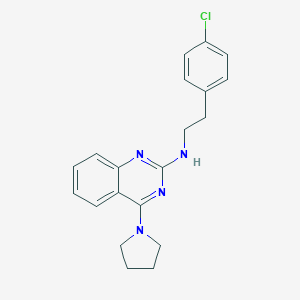 CHEMBL1762694 CHEMBL1762694 | C20H21ClN4 | 352.866 | 4 / 1 | 5.4 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218