

You can:
| Name | Histamine H3 receptor |
|---|---|
| Species | Rattus norvegicus (Rat) |
| Gene | Hrh3 |
| Synonym | GPCR97 H3 receptor H3R HH3R |
| Disease | N/A for non-human GPCRs |
| Length | 445 |
| Amino acid sequence | MERAPPDGLMNASGTLAGEAAAAGGARGFSAAWTAVLAALMALLIVATVLGNALVMLAFVADSSLRTQNNFFLLNLAISDFLVGAFCIPLYVPYVLTGRWTFGRGLCKLWLVVDYLLCASSVFNIVLISYDRFLSVTRAVSYRAQQGDTRRAVRKMALVWVLAFLLYGPAILSWEYLSGGSSIPEGHCYAEFFYNWYFLITASTLEFFTPFLSVTFFNLSIYLNIQRRTRLRLDGGREAGPEPPPDAQPSPPPAPPSCWGCWPKGHGEAMPLHRYGVGEAGPGVEAGEAALGGGSGGGAAASPTSSSGSSSRGTERPRSLKRGSKPSASSASLEKRMKMVSQSITQRFRLSRDKKVAKSLAIIVSIFGLCWAPYTLLMIIRAACHGRCIPDYWYETSFWLLWANSAVNPVLYPLCHYSFRRAFTKLLCPQKLKVQPHGSLEQCWK |
| UniProt | Q9QYN8 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL4124 |
| IUPHAR | 264 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 617 | 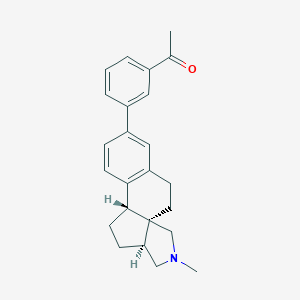 CHEMBL561028 CHEMBL561028 | C24H27NO | 345.486 | 2 / 0 | 4.5 | Yes |
| 618 | 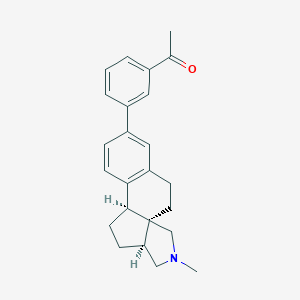 CHEMBL565206 CHEMBL565206 | C24H27NO | 345.486 | 2 / 0 | 4.5 | Yes |
| 738 | 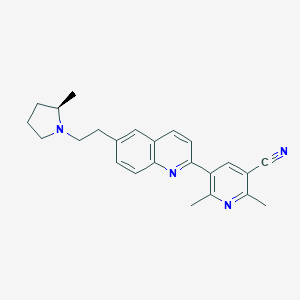 CHEMBL1085560 CHEMBL1085560 | C24H26N4 | 370.5 | 4 / 0 | 4.5 | Yes |
| 787 | 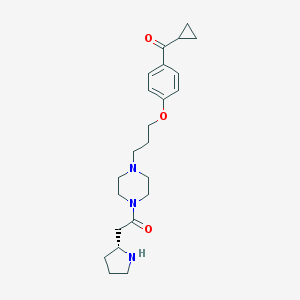 CHEMBL60435 CHEMBL60435 | C23H33N3O3 | 399.535 | 5 / 1 | 1.8 | Yes |
| 1058 | 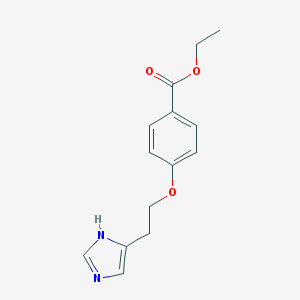 CHEMBL420892 CHEMBL420892 | C14H16N2O3 | 260.293 | 4 / 1 | 2.2 | Yes |
| 1785 | 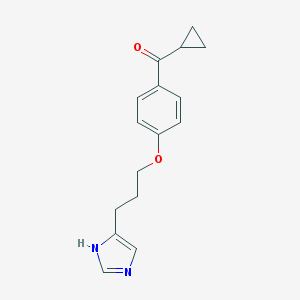 ciproxifan ciproxifan | C16H18N2O2 | 270.332 | 3 / 1 | 2.5 | Yes |
| 2204 | 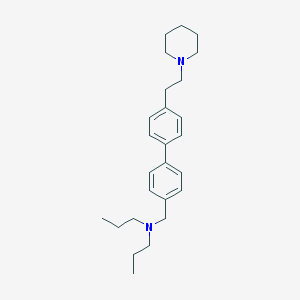 CHEMBL458722 CHEMBL458722 | C26H38N2 | 378.604 | 2 / 0 | 6.3 | No |
| 2936 | 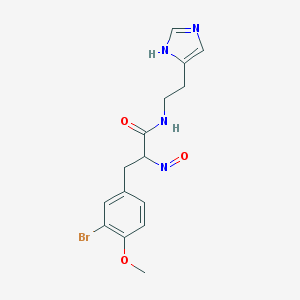 BDBM50070217 BDBM50070217 | C15H17BrN4O3 | 381.23 | 5 / 2 | 1.9 | Yes |
| 2958 | 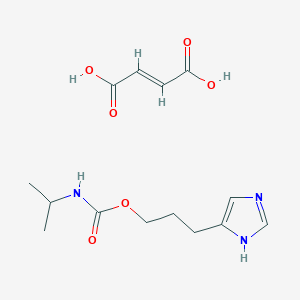 CID 44370594 CID 44370594 | C14H21N3O6 | 327.337 | 7 / 4 | N/A | N/A |
| 3019 | 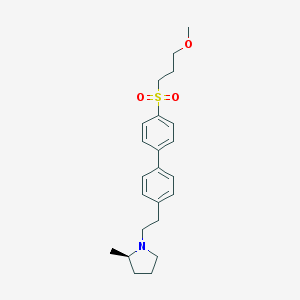 UNII-10521U3EQA UNII-10521U3EQA | C23H31NO3S | 401.565 | 4 / 0 | 4.2 | Yes |
| 441832 | 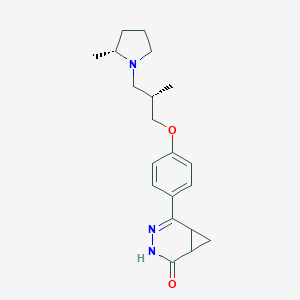 CHEMBL3417505 CHEMBL3417505 | C20H27N3O2 | 341.455 | 4 / 1 | 2.8 | Yes |
| 3777 | 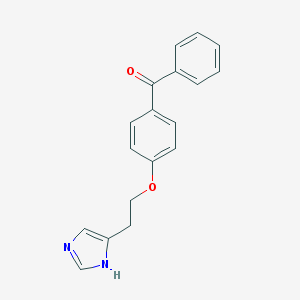 CHEMBL125751 CHEMBL125751 | C18H16N2O2 | 292.338 | 3 / 1 | 3.4 | Yes |
| 4092 | 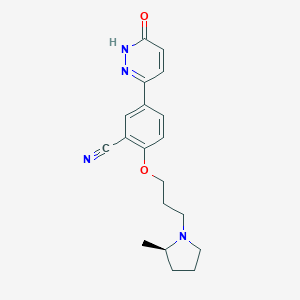 CHEMBL1829483 CHEMBL1829483 | C19H22N4O2 | 338.411 | 5 / 1 | 2.3 | Yes |
| 4378 | 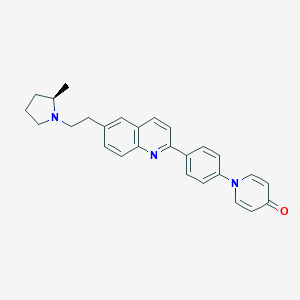 CHEMBL1086514 CHEMBL1086514 | C27H27N3O | 409.533 | 4 / 0 | 5.0 | Yes |
| 4403 | 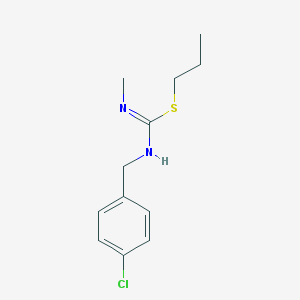 VUF-5649 VUF-5649 | C12H17ClN2S | 256.792 | 2 / 1 | 3.7 | Yes |
| 4799 | 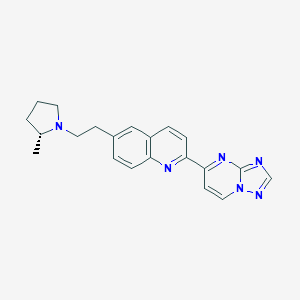 CHEMBL1084616 CHEMBL1084616 | C21H22N6 | 358.449 | 5 / 0 | 3.3 | Yes |
| 4970 | 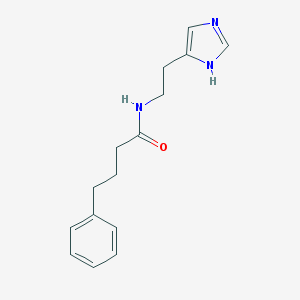 CHEMBL12494 CHEMBL12494 | C15H19N3O | 257.337 | 2 / 2 | 1.9 | Yes |
| 5296 | 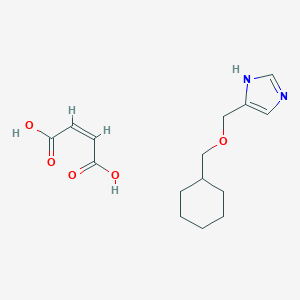 CID 44323707 CID 44323707 | C15H22N2O5 | 310.35 | 6 / 3 | N/A | N/A |
| 5445 | 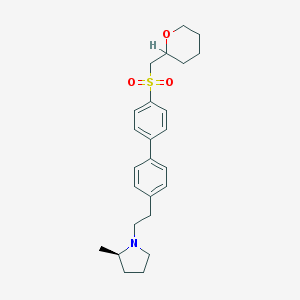 CHEMBL1934527 CHEMBL1934527 | C25H33NO3S | 427.603 | 4 / 0 | 4.8 | Yes |
| 5778 | 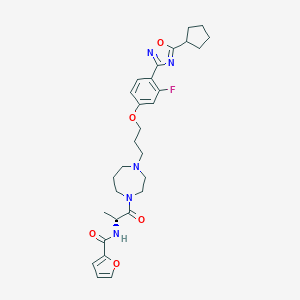 CHEMBL161202 CHEMBL161202 | C29H36FN5O5 | 553.635 | 9 / 1 | 4.4 | No |
| 5985 | 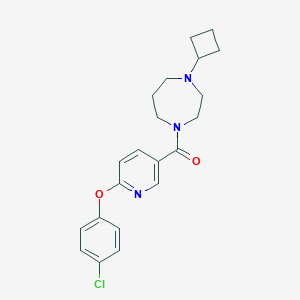 CHEMBL1171886 CHEMBL1171886 | C21H24ClN3O2 | 385.892 | 4 / 0 | 3.9 | Yes |
| 6498 | 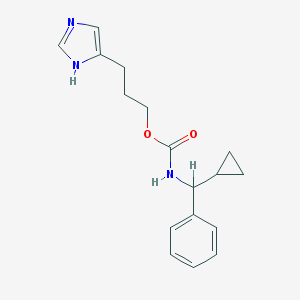 CHEMBL128585 CHEMBL128585 | C17H21N3O2 | 299.374 | 3 / 2 | 2.7 | Yes |
| 7198 | 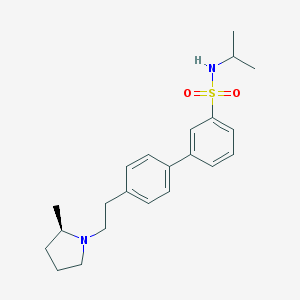 CHEMBL563696 CHEMBL563696 | C22H30N2O2S | 386.554 | 4 / 1 | 4.5 | Yes |
| 7199 | 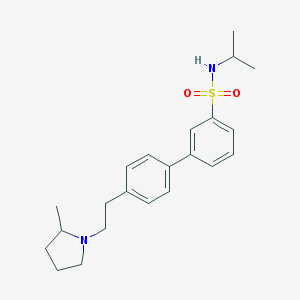 CHEMBL563696 CHEMBL563696 | C22H30N2O2S | 386.554 | 4 / 1 | 4.5 | Yes |
| 7498 | 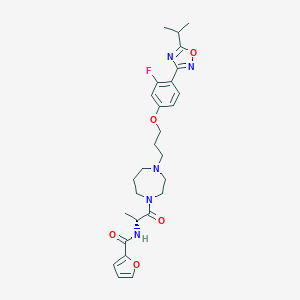 CHEMBL158056 CHEMBL158056 | C27H34FN5O5 | 527.597 | 9 / 1 | 3.9 | No |
| 8103 | 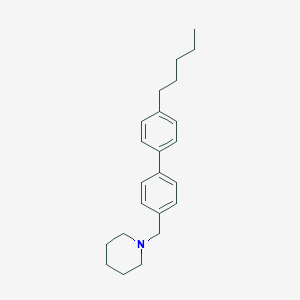 CHEMBL1934066 CHEMBL1934066 | C23H31N | 321.508 | 1 / 0 | 6.7 | No |
| 8180 | 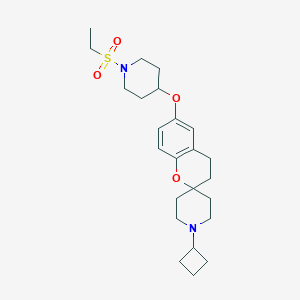 CHEMBL2013044 CHEMBL2013044 | C24H36N2O4S | 448.622 | 6 / 0 | 3.8 | Yes |
| 8414 | 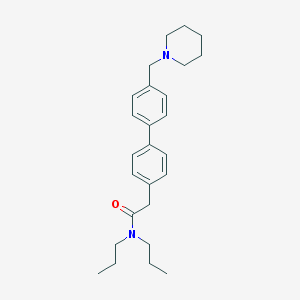 CHEMBL458507 CHEMBL458507 | C26H36N2O | 392.587 | 2 / 0 | 5.4 | No |
| 8418 | 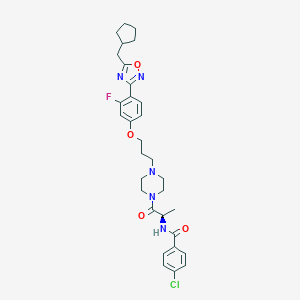 CHEMBL346711 CHEMBL346711 | C31H37ClFN5O4 | 598.116 | 8 / 1 | 6.1 | No |
| 8522 | 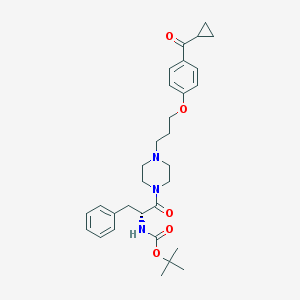 CHEMBL60143 CHEMBL60143 | C31H41N3O5 | 535.685 | 6 / 1 | 4.4 | No |
| 8553 | 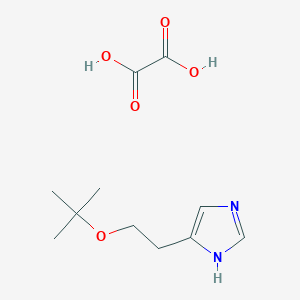 CID 44323688 CID 44323688 | C11H18N2O5 | 258.274 | 6 / 3 | N/A | N/A |
| 9012 | 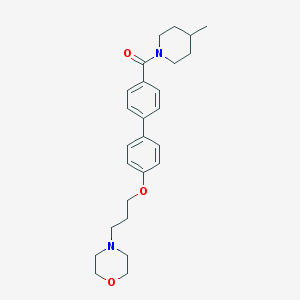 CHEMBL27545 CHEMBL27545 | C26H34N2O3 | 422.569 | 4 / 0 | 4.3 | Yes |
| 9062 | 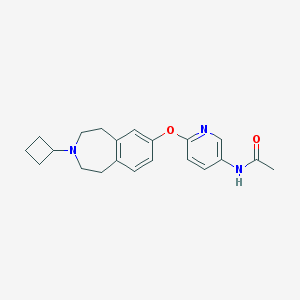 CHEMBL3092827 CHEMBL3092827 | C21H25N3O2 | 351.45 | 4 / 1 | 3.1 | Yes |
| 9454 | 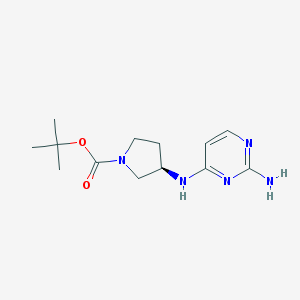 SCHEMBL3778815 SCHEMBL3778815 | C13H21N5O2 | 279.344 | 6 / 2 | 1.3 | Yes |
| 9807 | 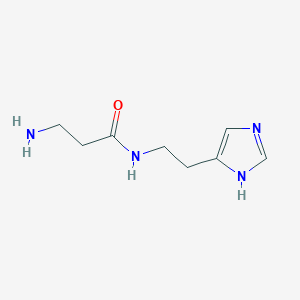 Carcinine Carcinine | C8H14N4O | 182.227 | 3 / 3 | -1.5 | Yes |
| 9905 | 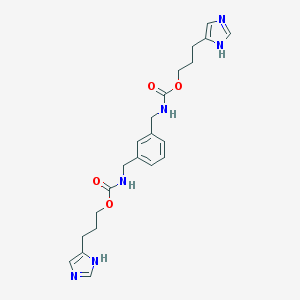 CHEMBL415989 CHEMBL415989 | C22H28N6O4 | 440.504 | 6 / 4 | 1.9 | Yes |
| 10555 | 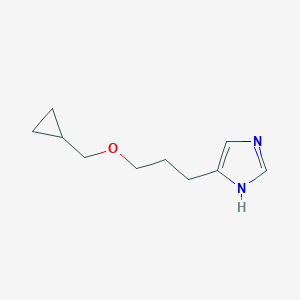 SCHEMBL7976815 SCHEMBL7976815 | C10H16N2O | 180.251 | 2 / 1 | 1.3 | Yes |
| 11206 | 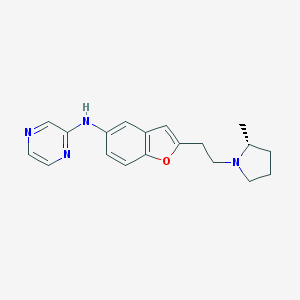 CHEMBL196467 CHEMBL196467 | C19H22N4O | 322.412 | 5 / 1 | 3.3 | Yes |
| 11655 | 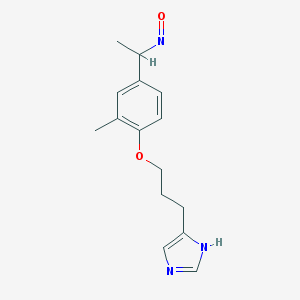 BDBM50091375 BDBM50091375 | C15H19N3O2 | 273.336 | 4 / 1 | 2.5 | Yes |
| 11796 | 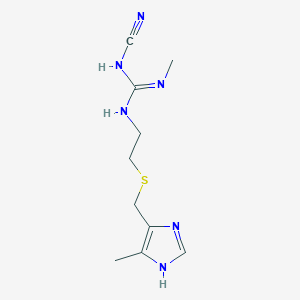 cimetidine cimetidine | C10H16N6S | 252.34 | 4 / 3 | 0.4 | Yes |
| 11873 | 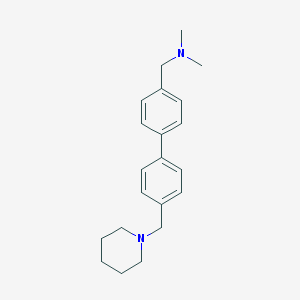 CHEMBL457006 CHEMBL457006 | C21H28N2 | 308.469 | 2 / 0 | 4.1 | Yes |
| 12339 | 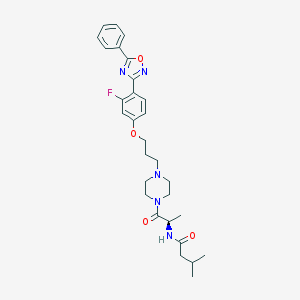 CHEMBL161565 CHEMBL161565 | C29H36FN5O4 | 537.636 | 8 / 1 | 4.2 | No |
| 12354 | 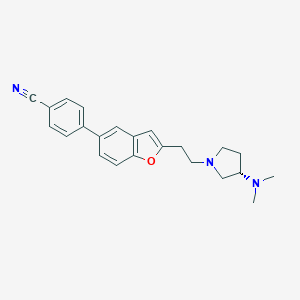 CHEMBL159281 CHEMBL159281 | C23H25N3O | 359.473 | 4 / 0 | 4.2 | Yes |
| 12356 | 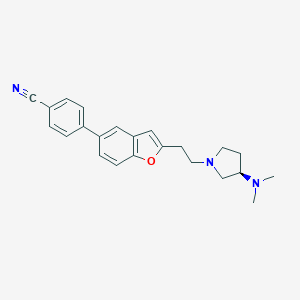 CHEMBL159648 CHEMBL159648 | C23H25N3O | 359.473 | 4 / 0 | 4.2 | Yes |
| 12688 | 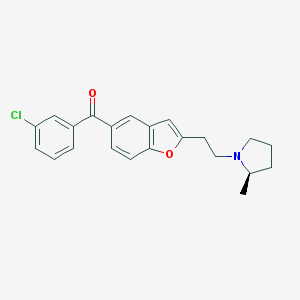 CHEMBL179263 CHEMBL179263 | C22H22ClNO2 | 367.873 | 3 / 0 | 5.5 | No |
| 12856 | 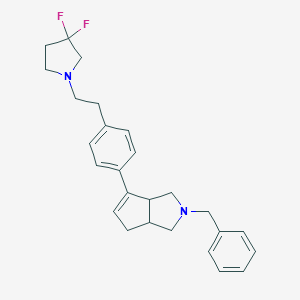 CHEMBL402296 CHEMBL402296 | C26H30F2N2 | 408.537 | 4 / 0 | 5.0 | Yes |
| 13019 | 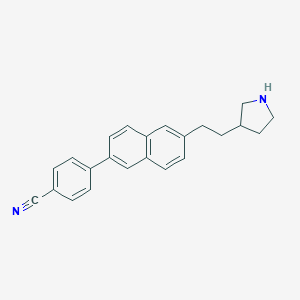 CHEMBL246679 CHEMBL246679 | C23H22N2 | 326.443 | 2 / 1 | 5.1 | No |
| 13439 | 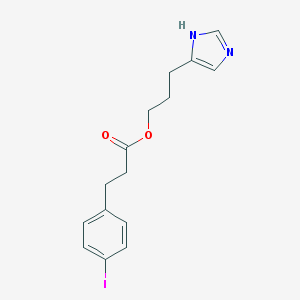 CHEMBL423473 CHEMBL423473 | C15H17IN2O2 | 384.217 | 3 / 1 | 3.1 | Yes |
| 13496 | 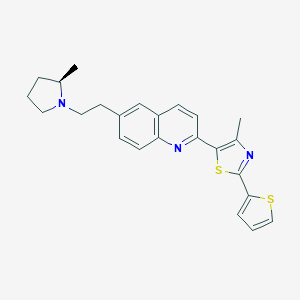 CHEMBL1086292 CHEMBL1086292 | C24H25N3S2 | 419.605 | 5 / 0 | 5.9 | No |
| 14558 | 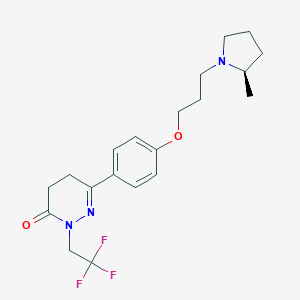 CHEMBL1935105 CHEMBL1935105 | C20H26F3N3O2 | 397.442 | 7 / 0 | 3.5 | Yes |
| 14646 | 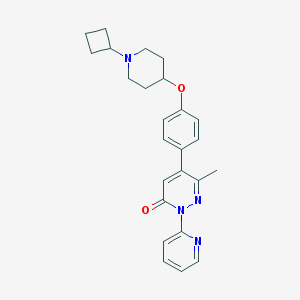 CHEMBL1945848 CHEMBL1945848 | C25H28N4O2 | 416.525 | 5 / 0 | 3.7 | Yes |
| 14750 | 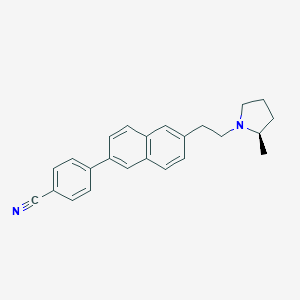 CHEMBL248155 CHEMBL248155 | C24H24N2 | 340.47 | 2 / 0 | 5.7 | No |
| 14753 | 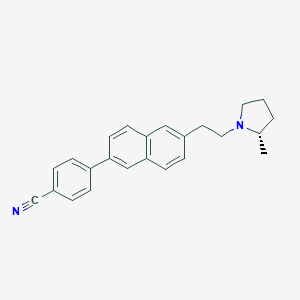 CHEMBL247505 CHEMBL247505 | C24H24N2 | 340.47 | 2 / 0 | 5.7 | No |
| 14754 | 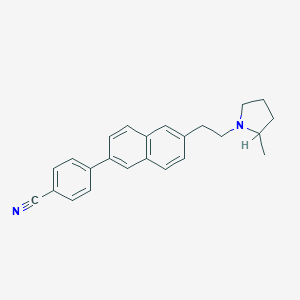 CHEMBL395190 CHEMBL395190 | C24H24N2 | 340.47 | 2 / 0 | 5.7 | No |
| 14809 | 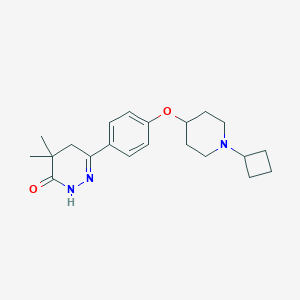 CHEMBL1950743 CHEMBL1950743 | C21H29N3O2 | 355.482 | 4 / 1 | 3.3 | Yes |
| 15146 | 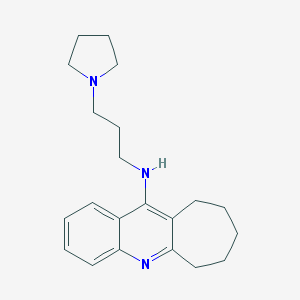 CHEMBL66388 CHEMBL66388 | C21H29N3 | 323.484 | 3 / 1 | 4.8 | Yes |
| 15410 | 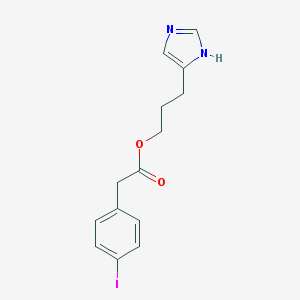 CHEMBL277227 CHEMBL277227 | C14H15IN2O2 | 370.19 | 3 / 1 | 2.8 | Yes |
| 15583 | 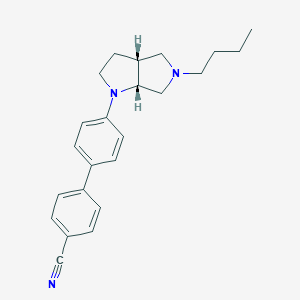 CHEMBL559445 CHEMBL559445 | C23H27N3 | 345.49 | 3 / 0 | 4.9 | Yes |
| 15762 | 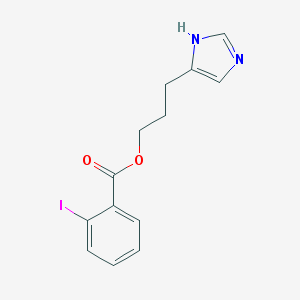 CHEMBL20191 CHEMBL20191 | C13H13IN2O2 | 356.163 | 3 / 1 | 3.3 | Yes |
| 15849 | 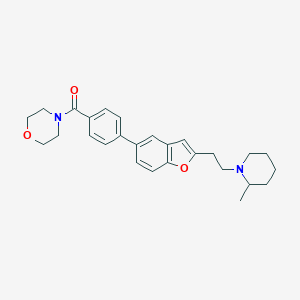 CHEMBL161130 CHEMBL161130 | C27H32N2O3 | 432.564 | 4 / 0 | 4.6 | Yes |
| 16010 | 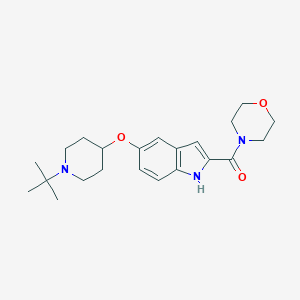 CHEMBL495611 CHEMBL495611 | C22H31N3O3 | 385.508 | 4 / 1 | 3.0 | Yes |
| 16011 | 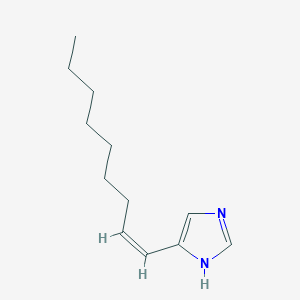 CHEMBL61729 CHEMBL61729 | C12H20N2 | 192.306 | 1 / 1 | 4.2 | Yes |
| 16013 | 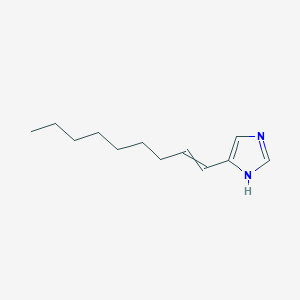 (1H-imidazol-4-yl)-1-nonene (1H-imidazol-4-yl)-1-nonene | C12H20N2 | 192.306 | 1 / 1 | 4.2 | Yes |
| 16285 | 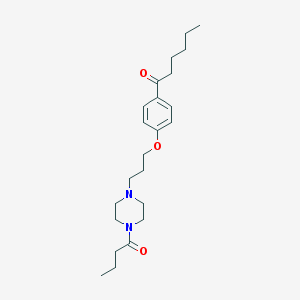 CHEMBL64479 CHEMBL64479 | C23H36N2O3 | 388.552 | 4 / 0 | 4.0 | Yes |
| 16639 | 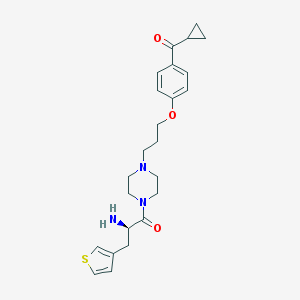 CHEMBL60559 CHEMBL60559 | C24H31N3O3S | 441.59 | 6 / 1 | 2.5 | Yes |
| 16978 | 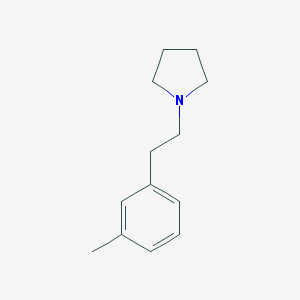 CHEMBL257919 CHEMBL257919 | C13H19N | 189.302 | 1 / 0 | 3.1 | Yes |
| 17317 | 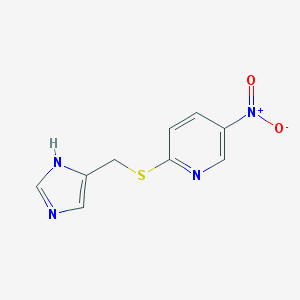 CHEMBL112576 CHEMBL112576 | C9H8N4O2S | 236.249 | 5 / 1 | 1.2 | Yes |
| 17458 | 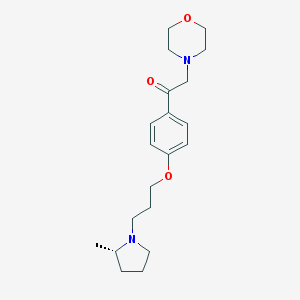 CHEMBL1951056 CHEMBL1951056 | C20H30N2O3 | 346.471 | 5 / 0 | 2.6 | Yes |
| 17463 | 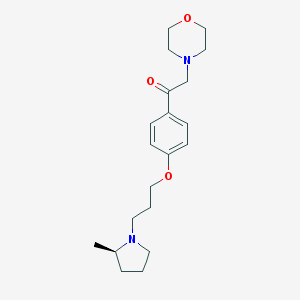 CHEMBL1951055 CHEMBL1951055 | C20H30N2O3 | 346.471 | 5 / 0 | 2.6 | Yes |
| 17464 | 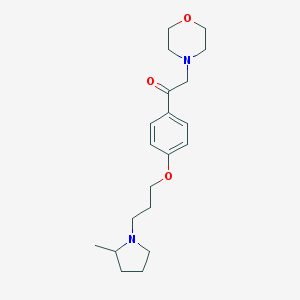 SCHEMBL931469 SCHEMBL931469 | C20H30N2O3 | 346.471 | 5 / 0 | 2.6 | Yes |
| 17990 | 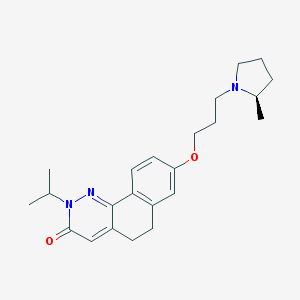 CHEMBL2031471 CHEMBL2031471 | C23H31N3O2 | 381.52 | 4 / 0 | 3.8 | Yes |
| 18036 | 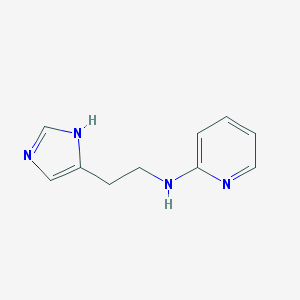 CHEMBL324890 CHEMBL324890 | C10H12N4 | 188.234 | 3 / 2 | 1.3 | Yes |
| 18045 | 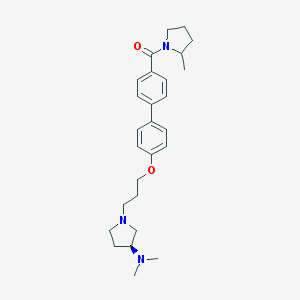 CHEMBL27835 CHEMBL27835 | C27H37N3O2 | 435.612 | 4 / 0 | 4.5 | Yes |
| 18106 | 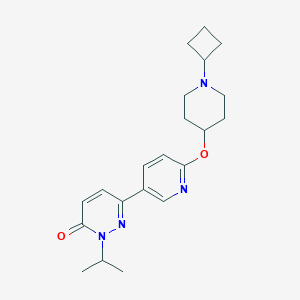 CHEMBL1950740 CHEMBL1950740 | C21H28N4O2 | 368.481 | 5 / 0 | 3.0 | Yes |
| 18110 | 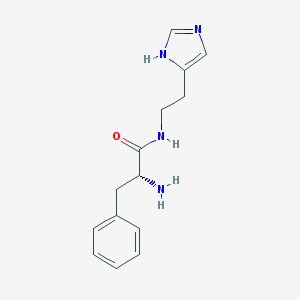 CHEMBL18884 CHEMBL18884 | C14H18N4O | 258.325 | 3 / 3 | 0.8 | Yes |
| 18111 | 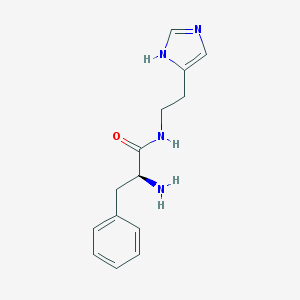 AC1OYVT5 AC1OYVT5 | C14H18N4O | 258.325 | 3 / 3 | 0.8 | Yes |
| 18453 | 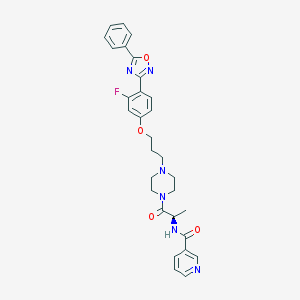 CHEMBL161279 CHEMBL161279 | C30H31FN6O4 | 558.614 | 9 / 1 | 3.5 | No |
| 18527 | 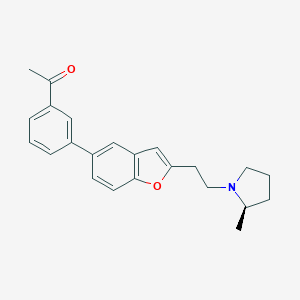 CHEMBL178853 CHEMBL178853 | C23H25NO2 | 347.458 | 3 / 0 | 4.9 | Yes |
| 18741 | 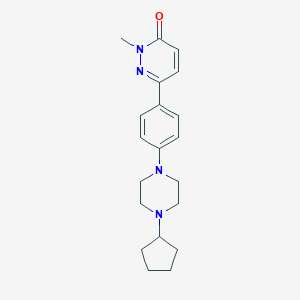 CHEMBL1823405 CHEMBL1823405 | C20H26N4O | 338.455 | 4 / 0 | 2.6 | Yes |
| 19045 | 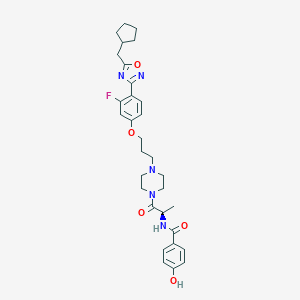 CHEMBL347909 CHEMBL347909 | C31H38FN5O5 | 579.673 | 9 / 2 | 5.1 | No |
| 19167 | 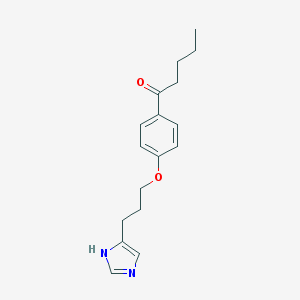 CHEMBL127901 CHEMBL127901 | C17H22N2O2 | 286.375 | 3 / 1 | 3.4 | Yes |
| 19414 | 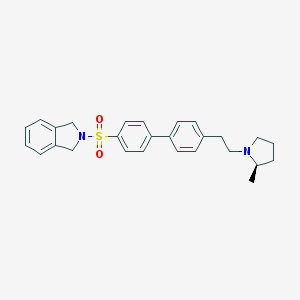 CHEMBL564803 CHEMBL564803 | C27H30N2O2S | 446.609 | 4 / 0 | 5.0 | Yes |
| 19415 | 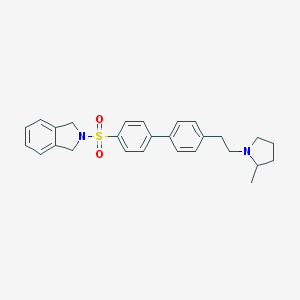 CHEMBL564803 CHEMBL564803 | C27H30N2O2S | 446.609 | 4 / 0 | 5.0 | Yes |
| 19723 | 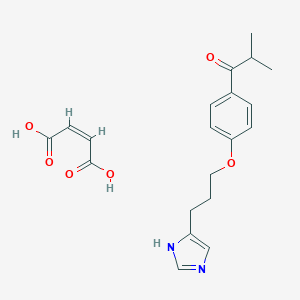 CHEMBL131021 CHEMBL131021 | C20H24N2O6 | 388.42 | 7 / 3 | N/A | N/A |
| 19925 | 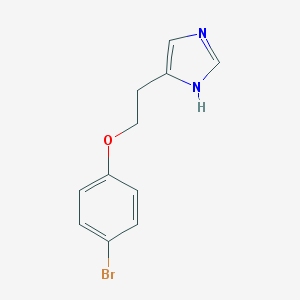 CHEMBL124864 CHEMBL124864 | C11H11BrN2O | 267.126 | 2 / 1 | 2.6 | Yes |
| 19972 | 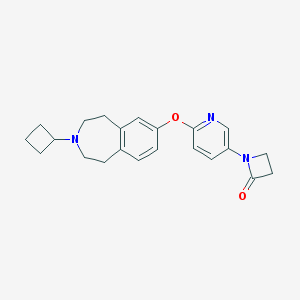 CHEMBL3092823 CHEMBL3092823 | C22H25N3O2 | 363.461 | 4 / 0 | 3.0 | Yes |
| 20262 | 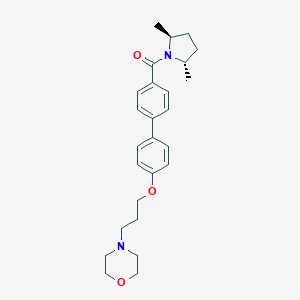 CHEMBL25126 CHEMBL25126 | C26H34N2O3 | 422.569 | 4 / 0 | 4.4 | Yes |
| 20293 | 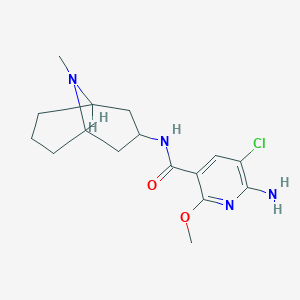 CHEMBL135410 CHEMBL135410 | C16H23ClN4O2 | 338.836 | 5 / 2 | 2.3 | Yes |
| 20479 | 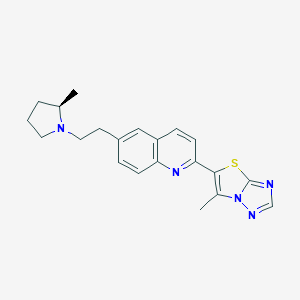 CHEMBL1084383 CHEMBL1084383 | C21H23N5S | 377.51 | 5 / 0 | 4.4 | Yes |
| 20749 | 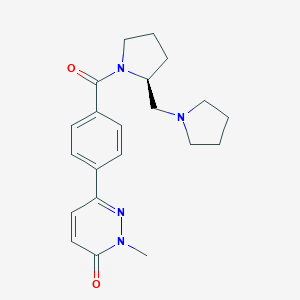 CHEMBL1823410 CHEMBL1823410 | C21H26N4O2 | 366.465 | 4 / 0 | 2.0 | Yes |
| 20775 | 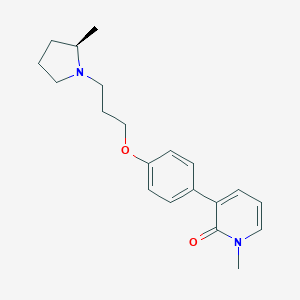 CHEMBL1923732 CHEMBL1923732 | C20H26N2O2 | 326.44 | 3 / 0 | 3.3 | Yes |
| 442510 | 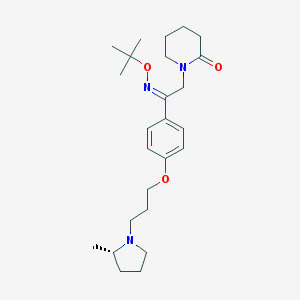 CHEMBL3331158 CHEMBL3331158 | C25H39N3O3 | 429.605 | 5 / 0 | 4.4 | Yes |
| 21570 | 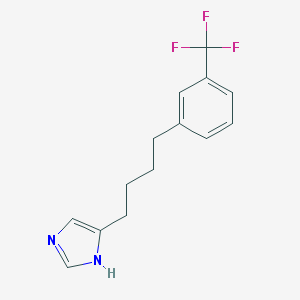 CHEMBL109498 CHEMBL109498 | C14H15F3N2 | 268.283 | 4 / 1 | 4.1 | Yes |
| 22405 | 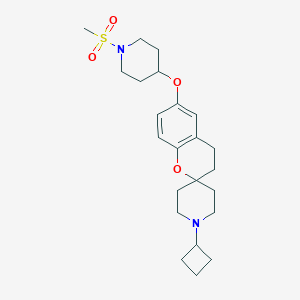 CHEMBL2013045 CHEMBL2013045 | C23H34N2O4S | 434.595 | 6 / 0 | 3.4 | Yes |
| 22438 | 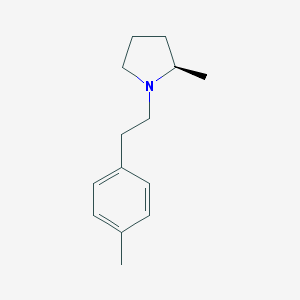 CHEMBL403052 CHEMBL403052 | C14H21N | 203.329 | 1 / 0 | 3.5 | Yes |
| 22440 | 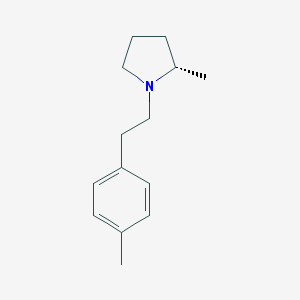 CHEMBL257911 CHEMBL257911 | C14H21N | 203.329 | 1 / 0 | 3.5 | Yes |
| 23010 | 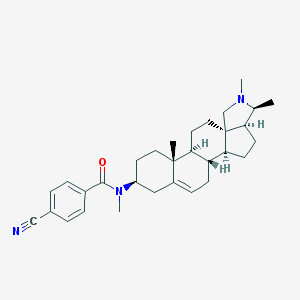 Conessine analogue, 12d Conessine analogue, 12d | C31H41N3O | 471.689 | 3 / 0 | 5.7 | No |
| 23391 | 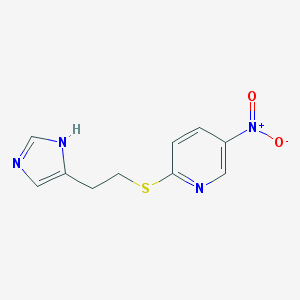 CHEMBL112504 CHEMBL112504 | C10H10N4O2S | 250.276 | 5 / 1 | 1.7 | Yes |
| 23646 | 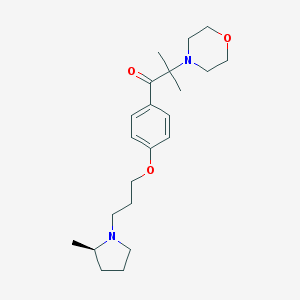 CHEMBL1951061 CHEMBL1951061 | C22H34N2O3 | 374.525 | 5 / 0 | 3.2 | Yes |
| 24129 | 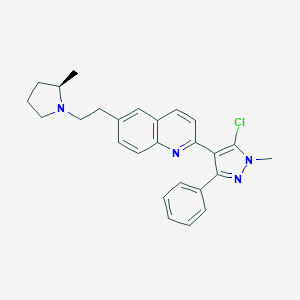 CHEMBL1085397 CHEMBL1085397 | C26H27ClN4 | 430.98 | 3 / 0 | 6.0 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218