

You can:
| Name | CHEMBL136684 |
|---|---|
| Molecular formula | C19H20N4O2 |
| IUPAC name | 2-(3,4-dihydro-1H-isoquinolin-2-yl)-6,7-dimethoxyquinazolin-4-amine |
| Molecular weight | 336.395 |
| Hydrogen bond acceptor | 6 |
| Hydrogen bond donor | 1 |
| XlogP | 3.1 |
| Synonyms | 2-(3,4-Dihydro-1H-isoquinolin-2-yl)-6,7-dimethoxy-quinazolin-4-ylamine BDBM50073575 SR-01000651481 2-[3,4-Dihydroisoquinoline-2(1H)-yl]-6,7-dimethoxyquinazoline-4-amine SR-01000651481-1 |
| Inchi Key | AMHBGRVKWJLMMP-UHFFFAOYSA-N |
| Inchi ID | InChI=1S/C19H20N4O2/c1-24-16-9-14-15(10-17(16)25-2)21-19(22-18(14)20)23-8-7-12-5-3-4-6-13(12)11-23/h3-6,9-10H,7-8,11H2,1-2H3,(H2,20,21,22) |
| PubChem CID | 10664606 |
| ChEMBL | CHEMBL136684 |
| IUPHAR | N/A |
| BindingDB | 50073575 |
| DrugBank | N/A |
Structure | 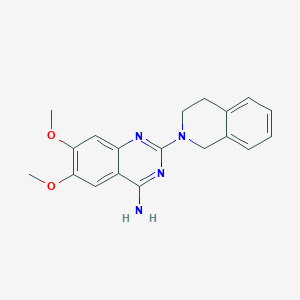 |
| Lipinski's druglikeness | This ligand satisfies Lipinski's rule of five. |
You can:
| GLASS ID | Name | UniProt | Gene | Species | Length |
|---|---|---|---|---|---|
| 8879 | 5-hydroxytryptamine receptor 1A | P19327 | Htr1a | Rattus norvegicus (Rat) | 422 |
| 8878 | 5-hydroxytryptamine receptor 2A | P14842 | Htr2a | Rattus norvegicus (Rat) | 471 |
| 8880 | D(2) dopamine receptor | P61169 | Drd2 | Rattus norvegicus (Rat) | 444 |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218