

You can:
| Name | Prostaglandin E2 receptor EP3 subtype |
|---|---|
| Species | Rattus norvegicus (Rat) |
| Gene | Ptger3 |
| Synonym | EP3 receptor PGE receptor EP3 subtype PGE2 receptor EP3 subtype prostaglandin E receptor 3 prostanoid EP3 receptor |
| Disease | N/A for non-human GPCRs |
| Length | 365 |
| Amino acid sequence | MAGVWAPEHSVEAHSNQSSAADGCGSVSVAFPITMMVTGFVGNALAMLLVVRSYRRRESKRKKSFLLCIGWLALTDLVGQLLTSPVVILVYLSQRRWEQLDPSGRLCTFFGLTMTVFGLSSLLVASAMAVERALAIRAPHWYASHMKTRATPVLLGVWLSVLAFALLPVLGVGRYSVQWPGTWCFISTGPAGNETDSAREPGSVAFASAFACLGLLALVVTFACNLATIKALVSRCRAKAAASQSSAQWGRITTETAIQLMGIMCVLSVCWSPLLIMMLKMIFNQMSVEQCKTQMGKEKECNSFLIAVRLASLNQILDPWVYLLLRKILLRKFCQIRDHTNYASSSTSLPCPGSSVLMWSDQLER |
| UniProt | P34980 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL5674 |
| IUPHAR | 342 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 553368 | 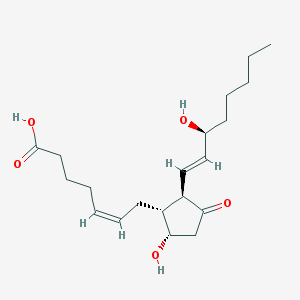 prostaglandin D2 prostaglandin D2 | C20H32O5 | 352.471 | 5 / 3 | 2.6 | Yes |
| 555634 | 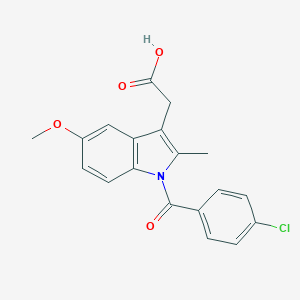 indomethacin indomethacin | C19H16ClNO4 | 357.79 | 4 / 1 | 4.3 | Yes |
| 553582 | 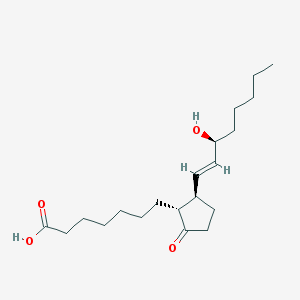 Doproston Doproston | C20H34O4 | 338.488 | 4 / 2 | 4.2 | Yes |
| 553680 | 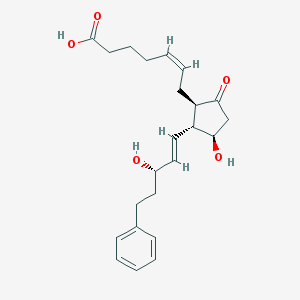 17-phenyl-trinor-PGE2 17-phenyl-trinor-PGE2 | C23H30O5 | 386.488 | 5 / 3 | 2.8 | Yes |
| 94048 | 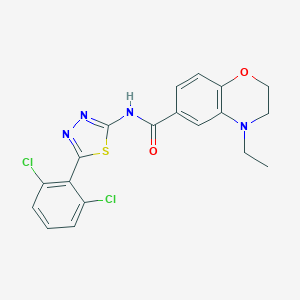 CHEMBL563480 CHEMBL563480 | C19H16Cl2N4O2S | 435.323 | 6 / 1 | 4.7 | Yes |
| 553747 | 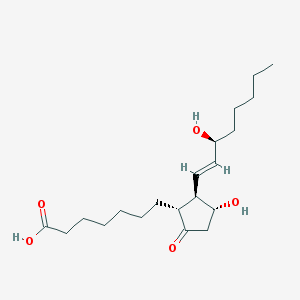 alprostadil alprostadil | C20H34O5 | 354.487 | 5 / 3 | 3.2 | Yes |
| 110550 | 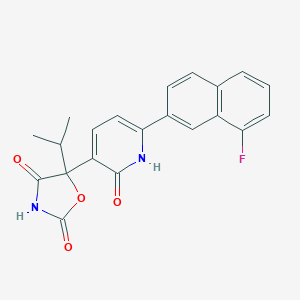 CHEMBL2035510 CHEMBL2035510 | C21H17FN2O4 | 380.375 | 5 / 2 | 3.7 | Yes |
| 553839 | 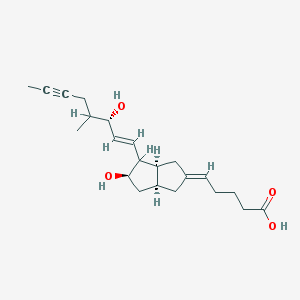 ILOPROST ILOPROST | C22H32O4 | 360.494 | 4 / 3 | 2.8 | Yes |
| 126731 | 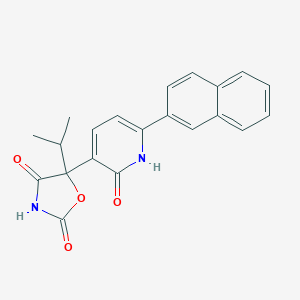 CHEMBL1770317 CHEMBL1770317 | C21H18N2O4 | 362.385 | 4 / 2 | 3.6 | Yes |
| 554185 | 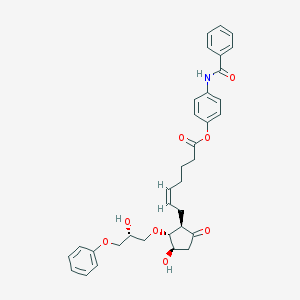 GR 63799 GR 63799 | C34H37NO8 | 587.669 | 8 / 3 | 3.9 | No |
| 186695 | 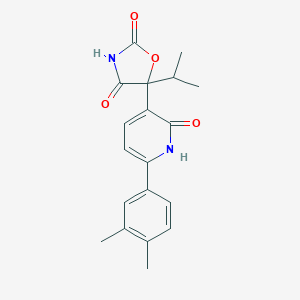 CHEMBL2035509 CHEMBL2035509 | C19H20N2O4 | 340.379 | 4 / 2 | 3.0 | Yes |
| 554240 | 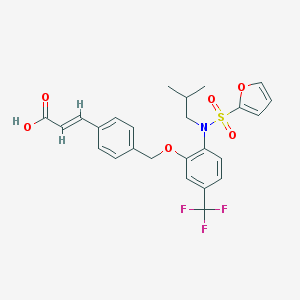 ONO-8713 ONO-8713 | C25H24F3NO6S | 523.523 | 10 / 1 | 5.7 | No |
| 554263 | 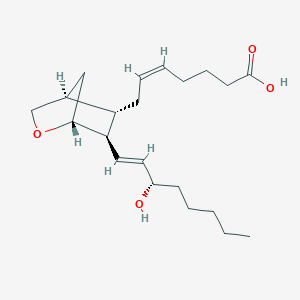 U 46619 U 46619 | C21H34O4 | 350.499 | 4 / 2 | 3.9 | Yes |
| 217452 | 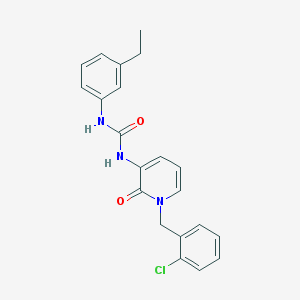 CHEMBL1271476 CHEMBL1271476 | C21H20ClN3O2 | 381.86 | 2 / 2 | 4.1 | Yes |
| 554443 | 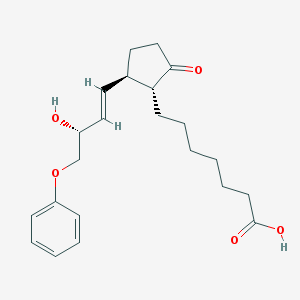 Dpt-prostaglandin E1 Dpt-prostaglandin E1 | C22H30O5 | 374.477 | 5 / 2 | 3.5 | Yes |
| 554540 | 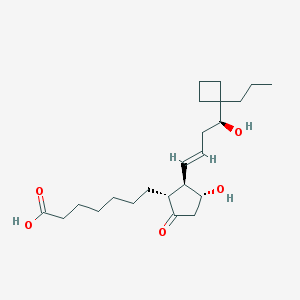 butaprost acid butaprost acid | C23H38O5 | 394.552 | 5 / 3 | 3.9 | Yes |
| 554600 | 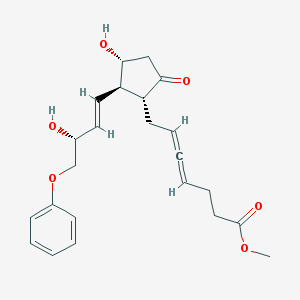 enprostil enprostil | C23H28O6 | 400.471 | 6 / 2 | 1.3 | Yes |
| 554624 | 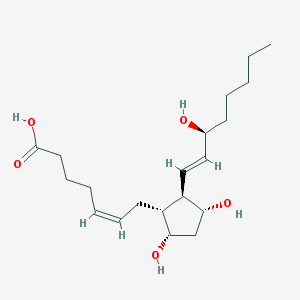 dinoprost dinoprost | C20H34O5 | 354.487 | 5 / 4 | 2.7 | Yes |
| 280145 | 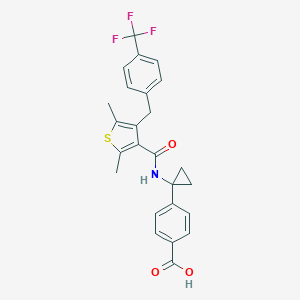 MK-2894 MK-2894 | C25H22F3NO3S | 473.51 | 7 / 2 | 5.9 | No |
| 556704 | 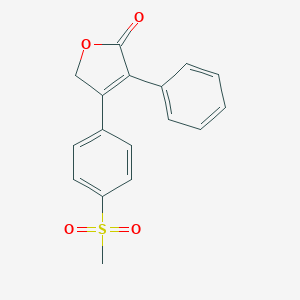 rofecoxib rofecoxib | C17H14O4S | 314.355 | 4 / 0 | 2.3 | Yes |
| 335482 | 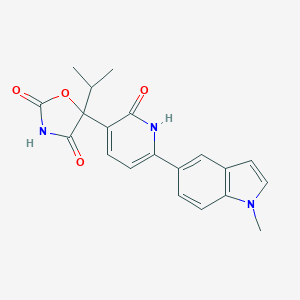 CHEMBL2035508 CHEMBL2035508 | C20H19N3O4 | 365.389 | 4 / 2 | 2.4 | Yes |
| 339599 | 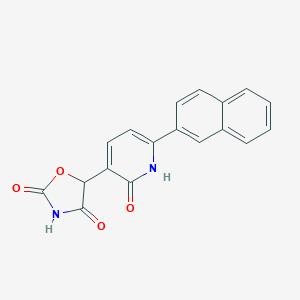 CID 70681702 CID 70681702 | C18H12N2O4 | 320.304 | 4 / 2 | 2.4 | Yes |
| 554986 | 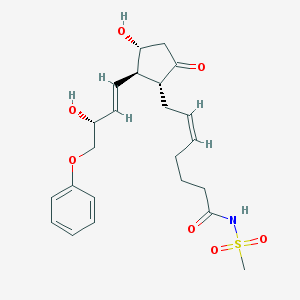 Sulprostone Sulprostone | C23H31NO7S | 465.561 | 7 / 3 | 1.5 | Yes |
| 555248 | 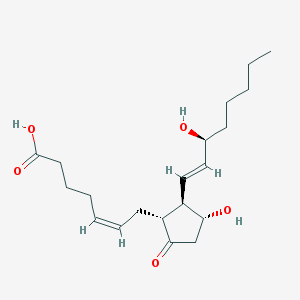 Prostaglandin E2 Prostaglandin E2 | C20H32O5 | 352.471 | 5 / 3 | 2.8 | Yes |
| 418834 | 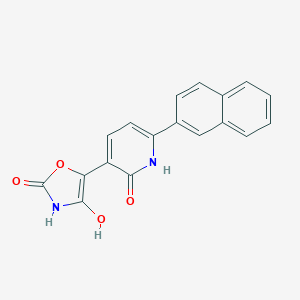 BDBM50384442 BDBM50384442 | C18H12N2O4 | 320.304 | 4 / 3 | 2.4 | Yes |
| 555475 | 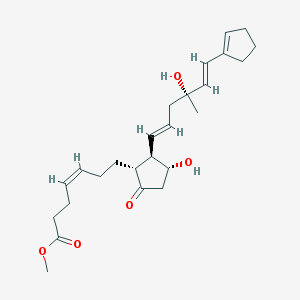 SC46275 SC46275 | C25H36O5 | 416.558 | 5 / 2 | 2.8 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218