

You can:
| Name | Neuropeptide S receptor |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | NPSR1 |
| Synonym | G-protein coupled receptor for asthma susceptibility vasopressin receptor-related receptor 1 G protein-coupled receptor 154 PGR14 NPS receptor [ Show all ] |
| Disease | Neurological disease |
| Length | 371 |
| Amino acid sequence | MPANFTEGSFDSSGTGQTLDSSPVACTETVTFTEVVEGKEWGSFYYSFKTEQLITLWVLFVFTIVGNSVVLFSTWRRKKKSRMTFFVTQLAITDSFTGLVNILTDINWRFTGDFTAPDLVCRVVRYLQVVLLYASTYVLVSLSIDRYHAIVYPMKFLQGEKQARVLIVIAWSLSFLFSIPTLIIFGKRTLSNGEVQCWALWPDDSYWTPYMTIVAFLVYFIPLTIISIMYGIVIRTIWIKSKTYETVISNCSDGKLCSSYNRGLISKAKIKAIKYSIIIILAFICCWSPYFLFDILDNFNLLPDTQERFYASVIIQNLPALNSAINPLIYCVFSSSISFPCREQRSQDSRMTFRERTERHEMQILSKPEFI |
| UniProt | Q6W5P4 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | Q6W5P4 |
| 3D structure model | This predicted structure model is from GPCR-EXP Q6W5P4. |
| BioLiP | N/A |
| Therapeutic Target Database | T20958 |
| ChEMBL | CHEMBL5162 |
| IUPHAR | 302 |
| DrugBank | BE0000948 |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 11 | 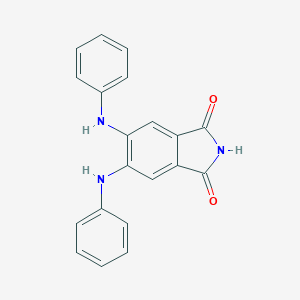 DAPH DAPH | C20H15N3O2 | 329.359 | 4 / 3 | 3.8 | Yes |
| 19 | 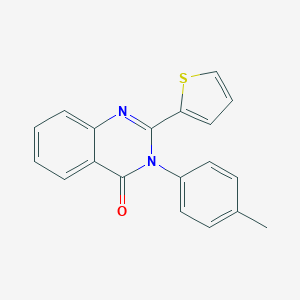 BAS 00216743 BAS 00216743 | C19H14N2OS | 318.394 | 3 / 0 | 4.5 | Yes |
| 36 | 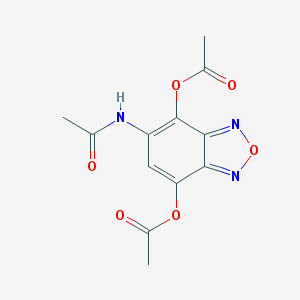 MLS000077136 MLS000077136 | C12H11N3O6 | 293.235 | 8 / 1 | -0.1 | Yes |
| 114 | 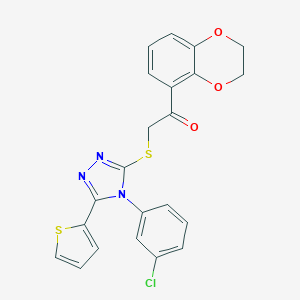 SMR000155400 SMR000155400 | C22H16ClN3O3S2 | 469.958 | 7 / 0 | 5.1 | No |
| 209 | 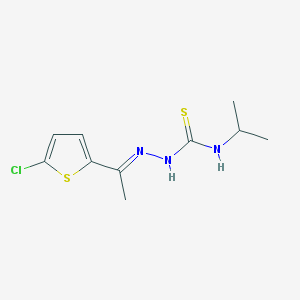 MLS000392662 MLS000392662 | C10H14ClN3S2 | 275.813 | 3 / 2 | 3.6 | Yes |
| 216 | 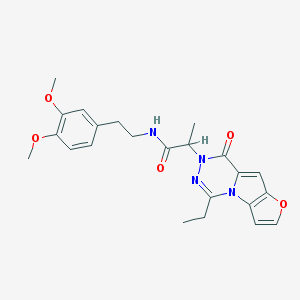 MLS000093986 MLS000093986 | C23H26N4O5 | 438.484 | 6 / 1 | 3.0 | Yes |
| 280 | 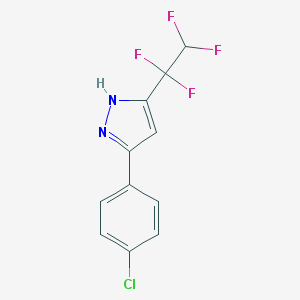 3-(4-chlorophenyl)-5-(1,1,2,2-tetrafluoroethyl)-1H-pyrazole 3-(4-chlorophenyl)-5-(1,1,2,2-tetrafluoroethyl)-1H-pyrazole | C11H7ClF4N2 | 278.635 | 5 / 1 | 3.8 | Yes |
| 299 | 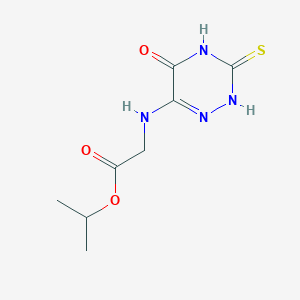 BAS 00662566 BAS 00662566 | C8H12N4O3S | 244.269 | 5 / 3 | 0.2 | Yes |
| 316 | 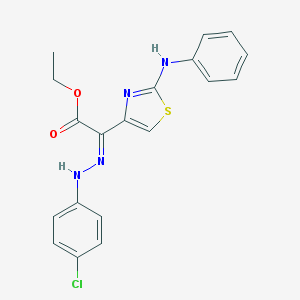 SMR000155256 SMR000155256 | C19H17ClN4O2S | 400.881 | 7 / 2 | 6.6 | No |
| 351 | 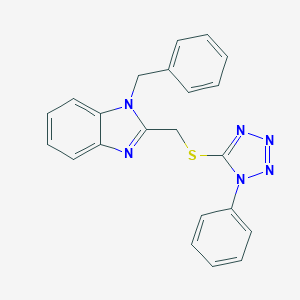 1-Benzyl-2-(1-phenyl-1H-tetrazol-5-ylsulfanylmethyl)-1H-benzoimidazole 1-Benzyl-2-(1-phenyl-1H-tetrazol-5-ylsulfanylmethyl)-1H-benzoimidazole | C22H18N6S | 398.488 | 5 / 0 | 4.7 | Yes |
| 377 | 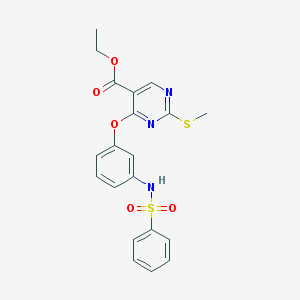 ethyl 2-(methylsulfanyl)-4-{3-[(phenylsulfonyl)amino]phenoxy}-5-pyrimidinecarboxylate ethyl 2-(methylsulfanyl)-4-{3-[(phenylsulfonyl)amino]phenoxy}-5-pyrimidinecarboxylate | C20H19N3O5S2 | 445.508 | 9 / 1 | 3.7 | Yes |
| 380 | 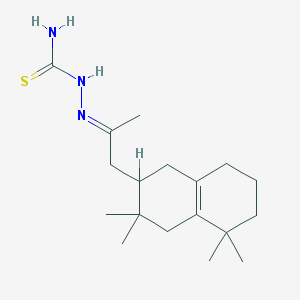 MLS000526436 MLS000526436 | C18H31N3S | 321.527 | 2 / 2 | 3.7 | Yes |
| 419 | 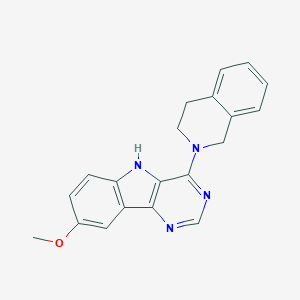 4-(3,4-Dihydro-1H-isoquinolin-2-yl)-8-methoxy-5H-pyrimido[5,4-b]indole 4-(3,4-Dihydro-1H-isoquinolin-2-yl)-8-methoxy-5H-pyrimido[5,4-b]indole | C20H18N4O | 330.391 | 4 / 1 | 3.7 | Yes |
| 427 | 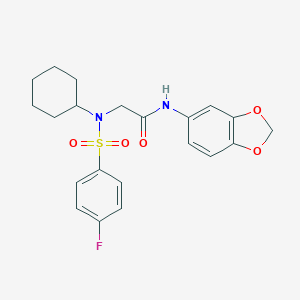 SMR000123112 SMR000123112 | C21H23FN2O5S | 434.482 | 7 / 1 | 3.7 | Yes |
| 459 | 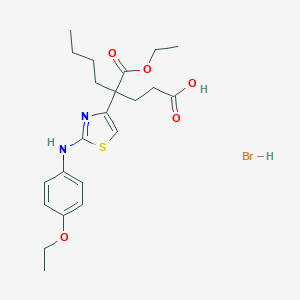 C22H31BrN2O5S C22H31BrN2O5S | C22H31BrN2O5S | 515.463 | 8 / 3 | N/A | No |
| 484 | 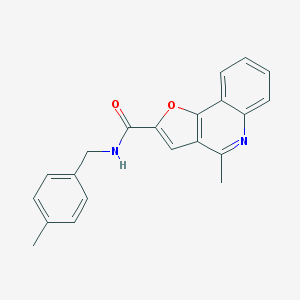 SMR000127857 SMR000127857 | C21H18N2O2 | 330.387 | 3 / 1 | 4.5 | Yes |
| 501 | 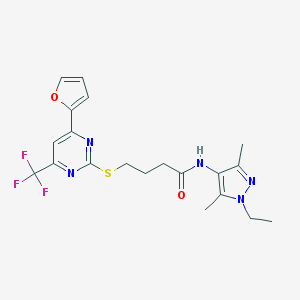 SMR000012485 SMR000012485 | C20H22F3N5O2S | 453.484 | 9 / 1 | 3.3 | Yes |
| 511 | 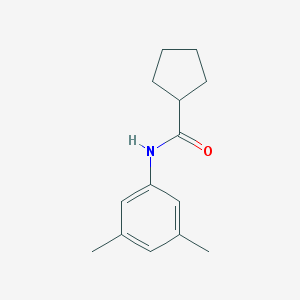 N-(3,5-dimethylphenyl)cyclopentanecarboxamide N-(3,5-dimethylphenyl)cyclopentanecarboxamide | C14H19NO | 217.312 | 1 / 1 | 3.4 | Yes |
| 557321 | 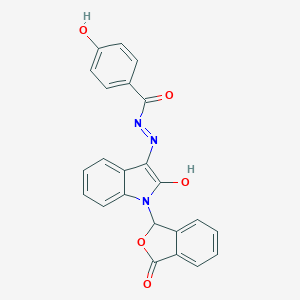 SMR000145689 SMR000145689 | C23H15N3O5 | 413.389 | 6 / 2 | 4.6 | Yes |
| 539 | 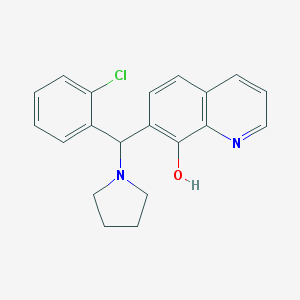 MLS000761483 MLS000761483 | C20H19ClN2O | 338.835 | 3 / 1 | 4.5 | Yes |
| 579 | 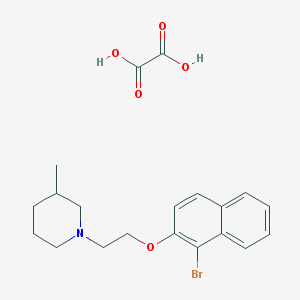 AC1MGH8D AC1MGH8D | C20H24BrNO5 | 438.318 | 6 / 2 | N/A | N/A |
| 646 | 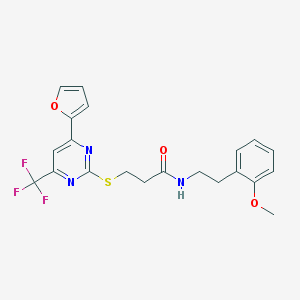 BAS 03609370 BAS 03609370 | C21H20F3N3O3S | 451.464 | 9 / 1 | 3.9 | Yes |
| 687 | 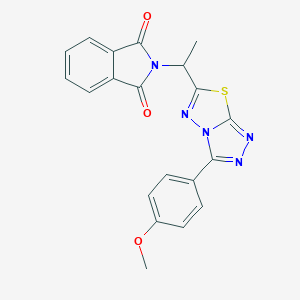 SMR000091760 SMR000091760 | C20H15N5O3S | 405.432 | 7 / 0 | 2.8 | Yes |
| 747 | 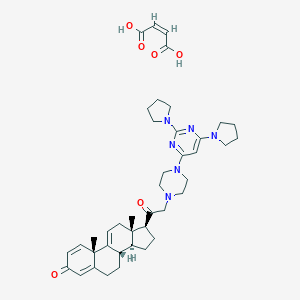 u-74389g u-74389g | C41H54N6O6 | 726.919 | 12 / 2 | N/A | No |
| 751 | 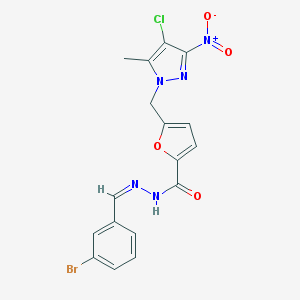 MLS000325386 MLS000325386 | C17H13BrClN5O4 | 466.676 | 6 / 1 | 4.1 | Yes |
| 790 | 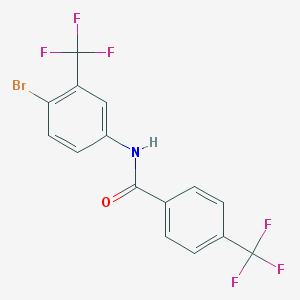 MLS000830224 MLS000830224 | C15H8BrF6NO | 412.129 | 7 / 1 | 5.2 | No |
| 839 | 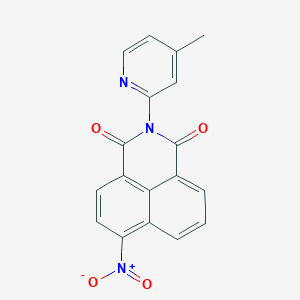 SMR000178156 SMR000178156 | C18H11N3O4 | 333.303 | 5 / 0 | 3.2 | Yes |
| 851 | 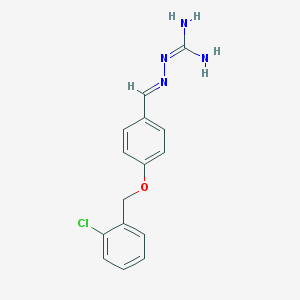 MLS000572570 MLS000572570 | C15H15ClN4O | 302.762 | 3 / 2 | 2.6 | Yes |
| 880 | 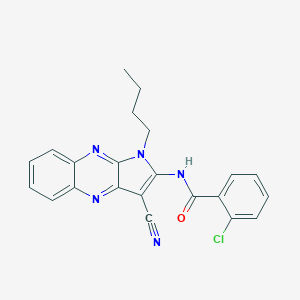 MLS000068755 MLS000068755 | C22H18ClN5O | 403.87 | 4 / 1 | 4.7 | Yes |
| 923 | 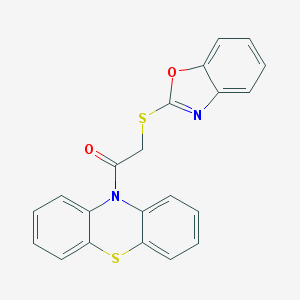 MLS000559294 MLS000559294 | C21H14N2O2S2 | 390.475 | 5 / 0 | 5.4 | No |
| 963 | 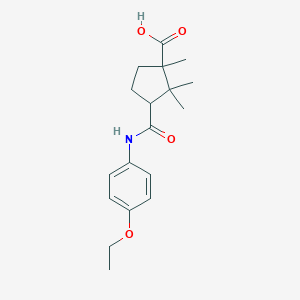 DEXTRO-CAMPHORIC P-PHENETIDIDE DEXTRO-CAMPHORIC P-PHENETIDIDE | C18H25NO4 | 319.401 | 4 / 2 | 3.0 | Yes |
| 965 | 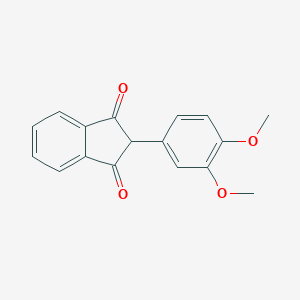 2-(3,4-Dimethoxyphenyl)-1H-indene-1,3(2H)-dione 2-(3,4-Dimethoxyphenyl)-1H-indene-1,3(2H)-dione | C17H14O4 | 282.295 | 4 / 0 | 2.9 | Yes |
| 995 | 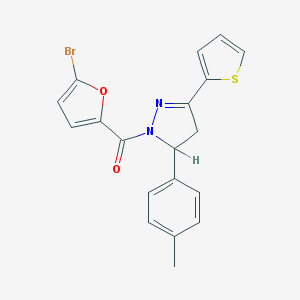 TCMDC-125610 TCMDC-125610 | C19H15BrN2O2S | 415.305 | 4 / 0 | 5.2 | No |
| 1003 | 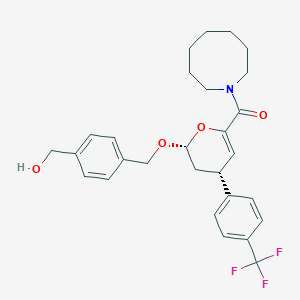 MLS000834357 MLS000834357 | C28H32F3NO4 | 503.562 | 7 / 1 | 5.5 | No |
| 1115 | 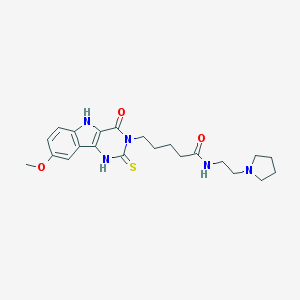 MLS000092917 MLS000092917 | C22H29N5O3S | 443.566 | 5 / 3 | 2.1 | Yes |
| 1157 | 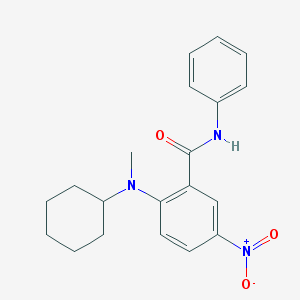 2-[cyclohexyl(methyl)amino]-5-nitro-N-phenylbenzamide 2-[cyclohexyl(methyl)amino]-5-nitro-N-phenylbenzamide | C20H23N3O3 | 353.422 | 4 / 1 | 4.5 | Yes |
| 1165 | 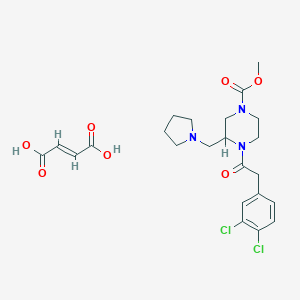 GR 89696 fumarate GR 89696 fumarate | C23H29Cl2N3O7 | 530.399 | 8 / 2 | N/A | No |
| 1197 | 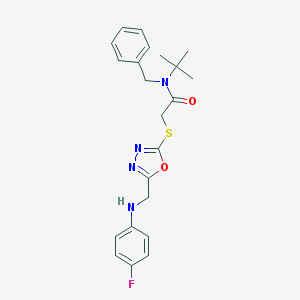 SMR000002891 SMR000002891 | C22H25FN4O2S | 428.526 | 7 / 1 | 4.2 | Yes |
| 1302 | 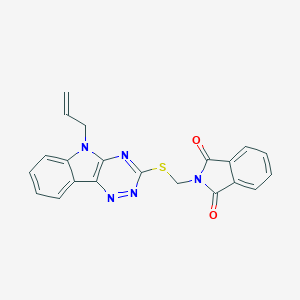 MLS000109191 MLS000109191 | C21H15N5O2S | 401.444 | 6 / 0 | 3.4 | Yes |
| 1304 | 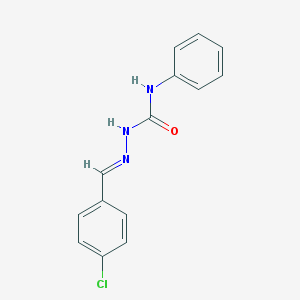 4-chlorobenzaldehyde N-phenylsemicarbazone 4-chlorobenzaldehyde N-phenylsemicarbazone | C14H12ClN3O | 273.72 | 2 / 2 | 3.2 | Yes |
| 1323 | 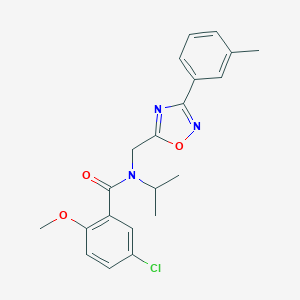 SMR000095165 SMR000095165 | C21H22ClN3O3 | 399.875 | 5 / 0 | 4.6 | Yes |
| 1336 | 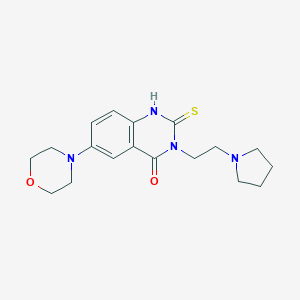 SMR000030980 SMR000030980 | C18H24N4O2S | 360.476 | 5 / 1 | 1.5 | Yes |
| 1389 | 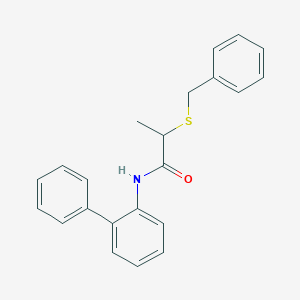 AC1MGWUV AC1MGWUV | C22H21NOS | 347.476 | 2 / 1 | 4.6 | Yes |
| 1392 | 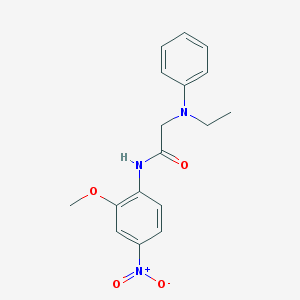 AC1MGB6Y AC1MGB6Y | C17H19N3O4 | 329.356 | 5 / 1 | 3.1 | Yes |
| 1402 | 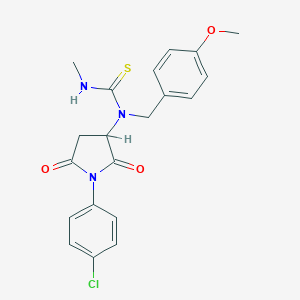 N-[1-(4-chlorophenyl)-2,5-dioxo-3-pyrrolidinyl]-N-(4-methoxybenzyl)-N'-methylthiourea N-[1-(4-chlorophenyl)-2,5-dioxo-3-pyrrolidinyl]-N-(4-methoxybenzyl)-N'-methylthiourea | C20H20ClN3O3S | 417.908 | 4 / 1 | 2.9 | Yes |
| 1429 | 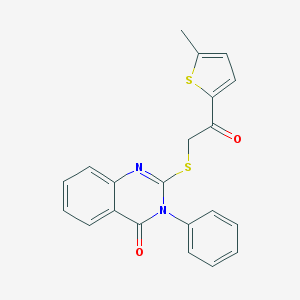 AC1LLNFY AC1LLNFY | C21H16N2O2S2 | 392.491 | 5 / 0 | 5.2 | No |
| 1431 | 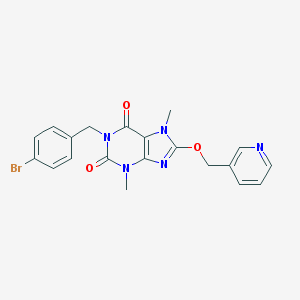 SMR000035518 SMR000035518 | C20H18BrN5O3 | 456.3 | 5 / 0 | 2.5 | Yes |
| 1484 | 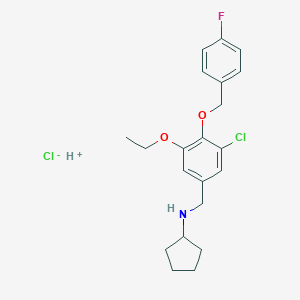 MLS-0106925.0002 MLS-0106925.0002 | C21H26Cl2FNO2 | 414.342 | 5 / 2 | N/A | N/A |
| 1499 | 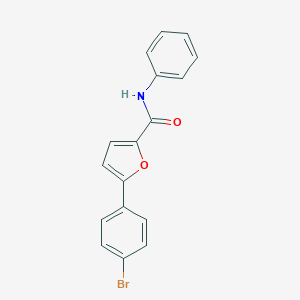 5-(4-bromophenyl)-N-phenyl-2-furamide 5-(4-bromophenyl)-N-phenyl-2-furamide | C17H12BrNO2 | 342.192 | 2 / 1 | 4.5 | Yes |
| 1579 | 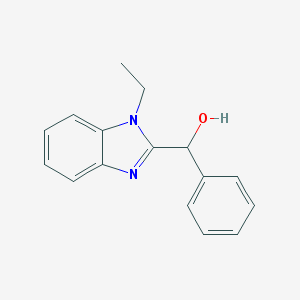 (1-Ethyl-1H-benzoimidazol-2-yl)-phenyl-methanol (1-Ethyl-1H-benzoimidazol-2-yl)-phenyl-methanol | C16H16N2O | 252.317 | 2 / 1 | 2.6 | Yes |
| 1615 | 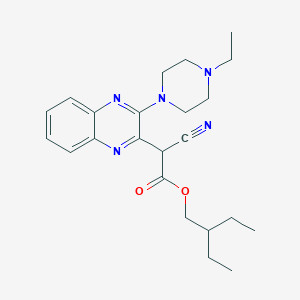 SMR000044012 SMR000044012 | C23H31N5O2 | 409.534 | 7 / 0 | 3.7 | Yes |
| 1619 | 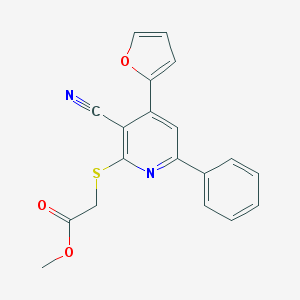 MLS000037786 MLS000037786 | C19H14N2O3S | 350.392 | 6 / 0 | 3.7 | Yes |
| 1627 | 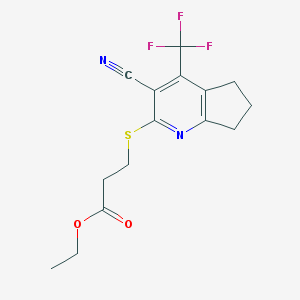 SMR000078129 SMR000078129 | C15H15F3N2O2S | 344.352 | 8 / 0 | 3.2 | Yes |
| 1647 | 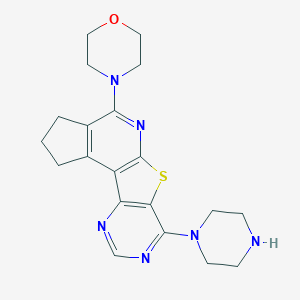 SMR000044353 SMR000044353 | C20H24N6OS | 396.513 | 8 / 1 | 2.6 | Yes |
| 1653 | 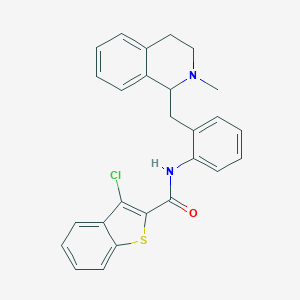 NSC309874 NSC309874 | C26H23ClN2OS | 446.993 | 3 / 1 | 6.6 | No |
| 557346 | 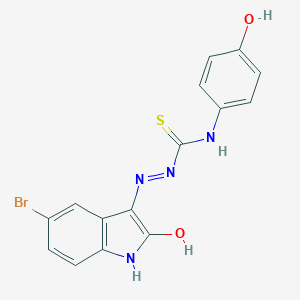 5-bromo-1H-indole-2,3-dione 3-[N-(4-hydroxyphenyl)thiosemicarbazone] 5-bromo-1H-indole-2,3-dione 3-[N-(4-hydroxyphenyl)thiosemicarbazone] | C15H11BrN4O2S | 391.243 | 4 / 4 | 4.3 | Yes |
| 1670 | 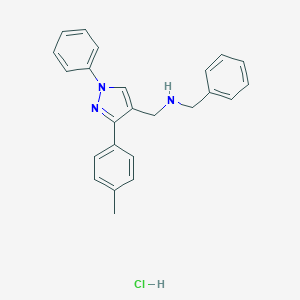 MLS000672046 MLS000672046 | C24H24ClN3 | 389.927 | 2 / 2 | N/A | N/A |
| 1719 | 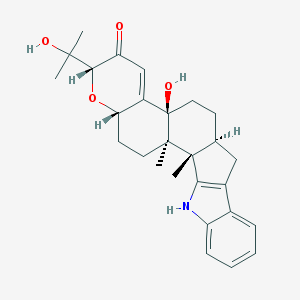 paxilline paxilline | C27H33NO4 | 435.564 | 4 / 3 | 3.5 | Yes |
| 1750 | 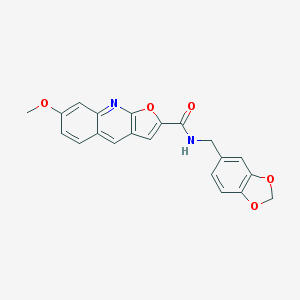 N-(1,3-benzodioxol-5-ylmethyl)-7-methoxyfuro[2,3-b]quinoline-2-carboxamide N-(1,3-benzodioxol-5-ylmethyl)-7-methoxyfuro[2,3-b]quinoline-2-carboxamide | C21H16N2O5 | 376.368 | 6 / 1 | 3.8 | Yes |
| 1762 | 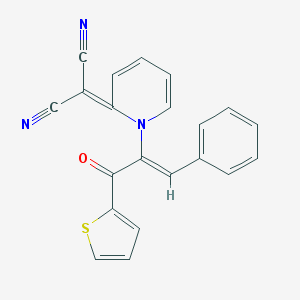 MLS000418503 MLS000418503 | C21H13N3OS | 355.415 | 5 / 0 | 4.8 | Yes |
| 1846 | 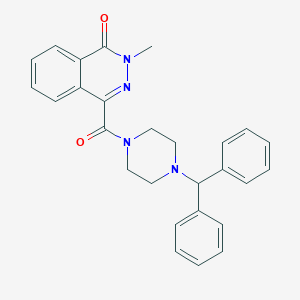 4-{[4-(diphenylmethyl)piperazin-1-yl]carbonyl}-2-methylphthalazin-1(2H)-one 4-{[4-(diphenylmethyl)piperazin-1-yl]carbonyl}-2-methylphthalazin-1(2H)-one | C27H26N4O2 | 438.531 | 4 / 0 | 3.8 | Yes |
| 1874 | 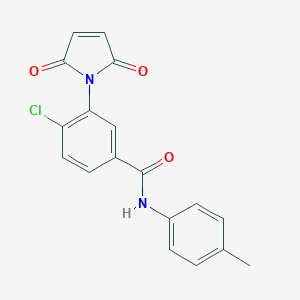 MLS000574219 MLS000574219 | C18H13ClN2O3 | 340.763 | 3 / 1 | 2.9 | Yes |
| 1881 | 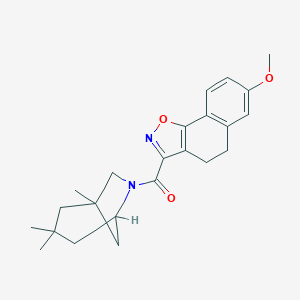 MLS000087448 MLS000087448 | C23H28N2O3 | 380.488 | 4 / 0 | 4.6 | Yes |
| 1892 | 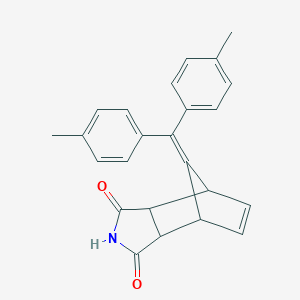 MLS000537663 MLS000537663 | C24H21NO2 | 355.437 | 2 / 1 | 4.1 | Yes |
| 1946 | 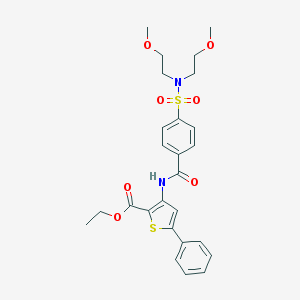 ethyl 3-(4-(N,N-bis(2-methoxyethyl)sulfamoyl)benzamido)-5-phenylthiophene-2-carboxylate ethyl 3-(4-(N,N-bis(2-methoxyethyl)sulfamoyl)benzamido)-5-phenylthiophene-2-carboxylate | C26H30N2O7S2 | 546.653 | 9 / 1 | 4.1 | No |
| 1957 | 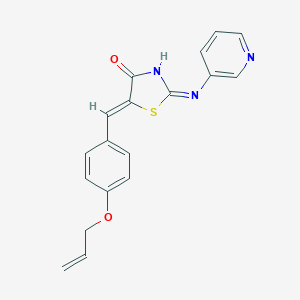 MLS000333603 MLS000333603 | C18H15N3O2S | 337.397 | 5 / 1 | 3.6 | Yes |
| 2007 | 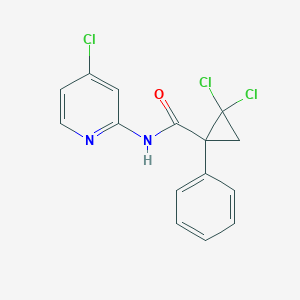 AC1NE6QE AC1NE6QE | C15H11Cl3N2O | 341.616 | 2 / 1 | 3.7 | Yes |
| 2020 | 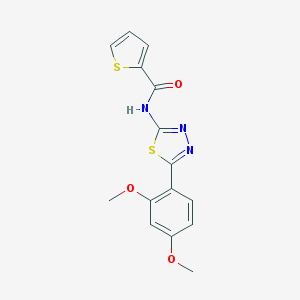 MLS001006960 MLS001006960 | C15H13N3O3S2 | 347.407 | 7 / 1 | 3.2 | Yes |
| 2067 | 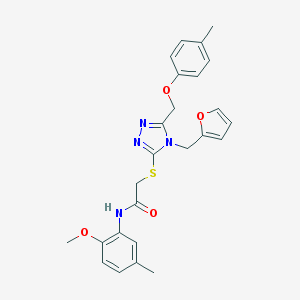 SMR000046468 SMR000046468 | C25H26N4O4S | 478.567 | 7 / 1 | 4.1 | Yes |
| 2078 | 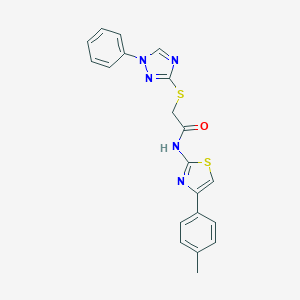 N-[4-(4-methylphenyl)-1,3-thiazol-2-yl]-2-[(1-phenyl-1H-1,2,4-triazol-3-yl)thio]acetamide N-[4-(4-methylphenyl)-1,3-thiazol-2-yl]-2-[(1-phenyl-1H-1,2,4-triazol-3-yl)thio]acetamide | C20H17N5OS2 | 407.51 | 6 / 1 | 4.8 | Yes |
| 2081 | 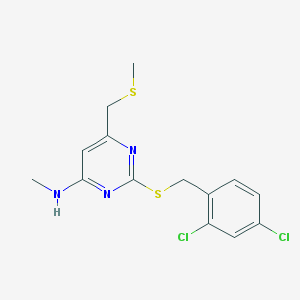 2-[(2,4-dichlorobenzyl)sulfanyl]-N-methyl-6-[(methylsulfanyl)methyl]-4-pyrimidinamine 2-[(2,4-dichlorobenzyl)sulfanyl]-N-methyl-6-[(methylsulfanyl)methyl]-4-pyrimidinamine | C14H15Cl2N3S2 | 360.315 | 5 / 1 | 4.6 | Yes |
| 2107 | 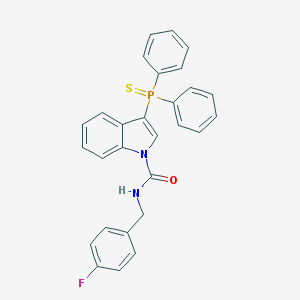 SCHEMBL2590676 SCHEMBL2590676 | C28H22FN2OPS | 484.529 | 3 / 1 | 7.2 | No |
| 2126 | 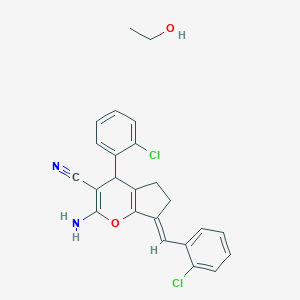 MLS000587548 MLS000587548 | C24H22Cl2N2O2 | 441.352 | 4 / 2 | N/A | N/A |
| 2156 | 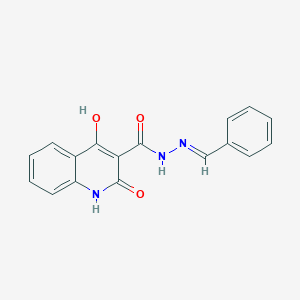 4-hydroxy-2-oxo-N'-[(E)-phenylmethylidene]-1,2-dihydroquinoline-3-carbohydrazide 4-hydroxy-2-oxo-N'-[(E)-phenylmethylidene]-1,2-dihydroquinoline-3-carbohydrazide | C17H13N3O3 | 307.309 | 4 / 3 | 2.9 | Yes |
| 2210 | 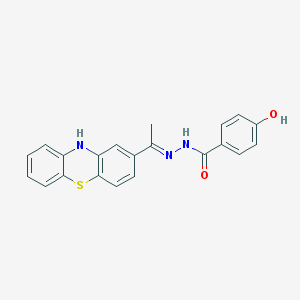 SR-01000697372 SR-01000697372 | C21H17N3O2S | 375.446 | 5 / 3 | 4.4 | Yes |
| 2226 | 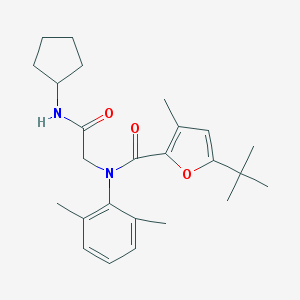 MLS000556811 MLS000556811 | C25H34N2O3 | 410.558 | 3 / 1 | 5.8 | No |
| 2294 | 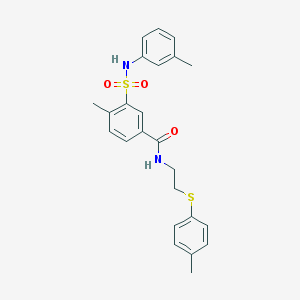 SMR000201550 SMR000201550 | C24H26N2O3S2 | 454.603 | 5 / 2 | 5.0 | Yes |
| 2303 | 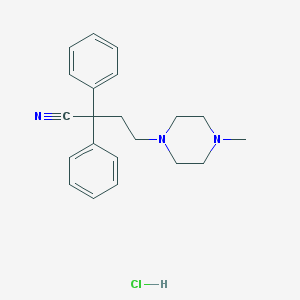 SMR000042447 SMR000042447 | C21H26ClN3 | 355.91 | 3 / 1 | N/A | N/A |
| 2316 | 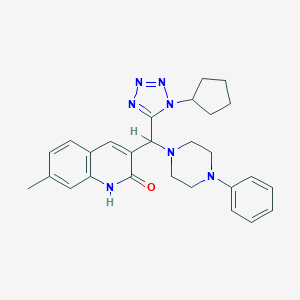 MLS000071942 MLS000071942 | C27H31N7O | 469.593 | 6 / 1 | 3.7 | Yes |
| 2400 | 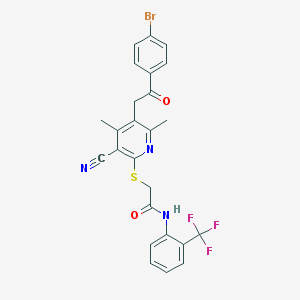 MLS000537437 MLS000537437 | C25H19BrF3N3O2S | 562.405 | 8 / 1 | 5.9 | No |
| 2928 | 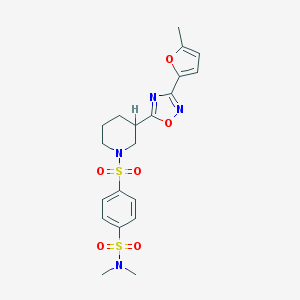 SMR000124404 SMR000124404 | C20H24N4O6S2 | 480.554 | 10 / 0 | 1.9 | Yes |
| 2940 | 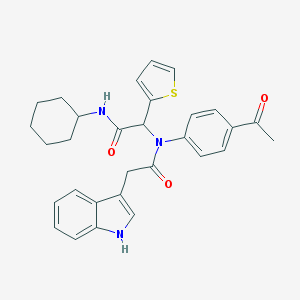 SMR000172878 SMR000172878 | C30H31N3O3S | 513.656 | 4 / 2 | 5.3 | No |
| 2976 | 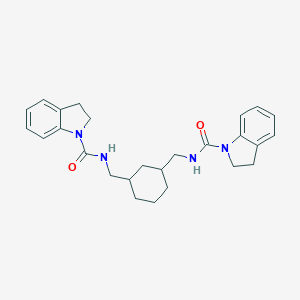 SMR000151229 SMR000151229 | C26H32N4O2 | 432.568 | 2 / 2 | 4.0 | Yes |
| 3036 | 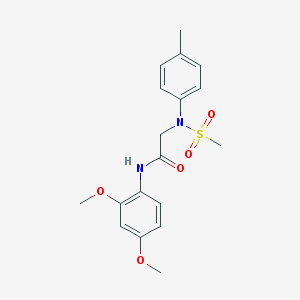 SMR000104366 SMR000104366 | C18H22N2O5S | 378.443 | 6 / 1 | 2.3 | Yes |
| 3070 | 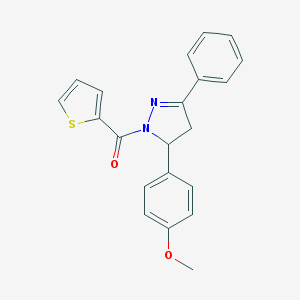 SMR000193353 SMR000193353 | C21H18N2O2S | 362.447 | 4 / 0 | 4.3 | Yes |
| 3075 | 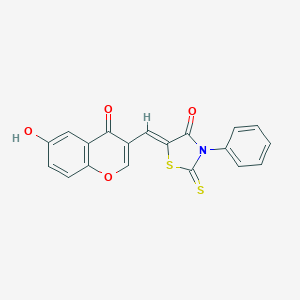 AC1LXTED AC1LXTED | C19H11NO4S2 | 381.42 | 6 / 1 | 3.8 | Yes |
| 3077 | 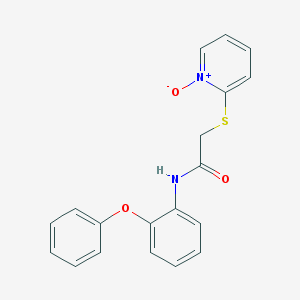 MLS001006036 MLS001006036 | C19H16N2O3S | 352.408 | 4 / 1 | 3.1 | Yes |
| 3194 | 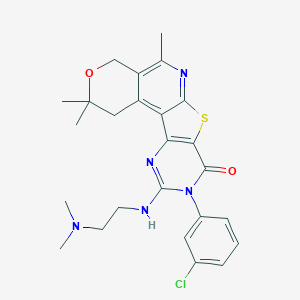 AC1NKK02 AC1NKK02 | C25H28ClN5O2S | 498.042 | 6 / 1 | 4.2 | Yes |
| 3196 | 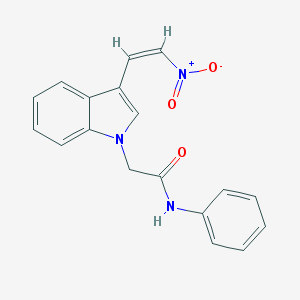 AC1NY53V AC1NY53V | C18H15N3O3 | 321.336 | 3 / 1 | 3.3 | Yes |
| 3203 | 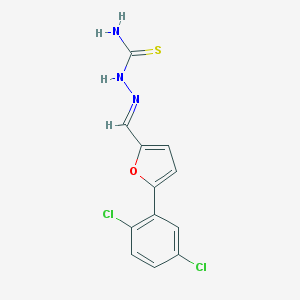 AC1OC89K AC1OC89K | C12H9Cl2N3OS | 314.184 | 3 / 2 | 3.5 | Yes |
| 3229 | 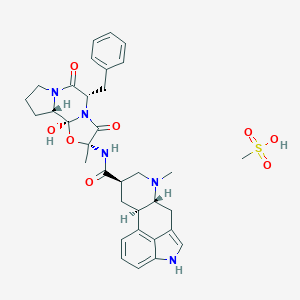 Dihydroergotamine mesylate Dihydroergotamine mesylate | C34H41N5O8S | 679.789 | 9 / 4 | N/A | No |
| 3242 | 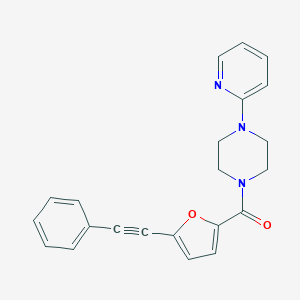 (5-Phenylethynyl-furan-2-yl)-(4-pyridin-2-yl-piperazin-1-yl)-methanone (5-Phenylethynyl-furan-2-yl)-(4-pyridin-2-yl-piperazin-1-yl)-methanone | C22H19N3O2 | 357.413 | 4 / 0 | 3.7 | Yes |
| 3248 | 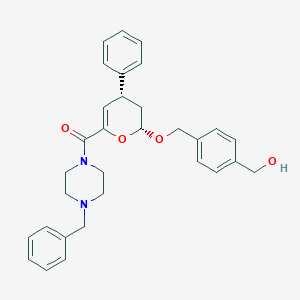 MLS000766439 MLS000766439 | C31H34N2O4 | 498.623 | 5 / 1 | 4.2 | Yes |
| 3256 | 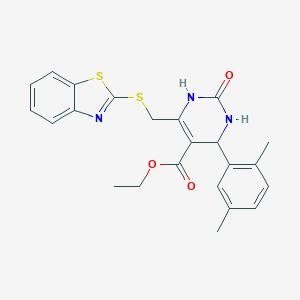 SMR000201450 SMR000201450 | C23H23N3O3S2 | 453.575 | 6 / 2 | 4.5 | Yes |
| 3259 | 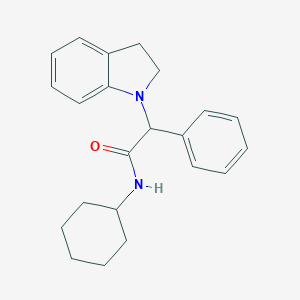 SMR000127622 SMR000127622 | C22H26N2O | 334.463 | 2 / 1 | 4.9 | Yes |
| 3279 | 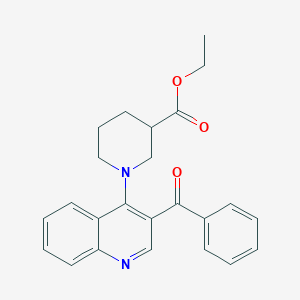 ethyl 1-(3-benzoylquinolin-4-yl)piperidine-3-carboxylate ethyl 1-(3-benzoylquinolin-4-yl)piperidine-3-carboxylate | C24H24N2O3 | 388.467 | 5 / 0 | 4.4 | Yes |
| 3280 | 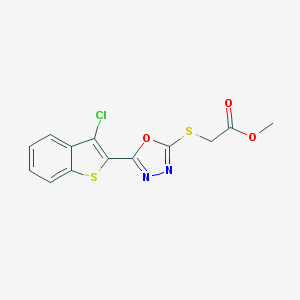 SMR000196863 SMR000196863 | C13H9ClN2O3S2 | 340.796 | 7 / 0 | 3.9 | Yes |
| 3295 | 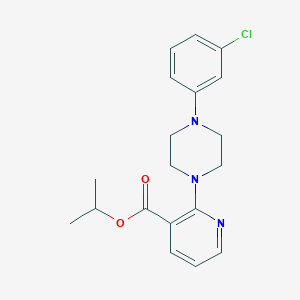 isopropyl 2-[4-(3-chlorophenyl)piperazino]nicotinate isopropyl 2-[4-(3-chlorophenyl)piperazino]nicotinate | C19H22ClN3O2 | 359.854 | 5 / 0 | 4.2 | Yes |
| 3339 | 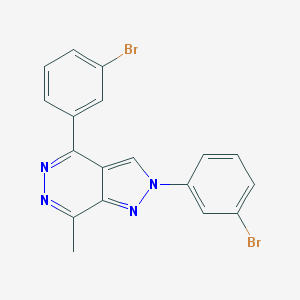 MLS000544661 MLS000544661 | C18H12Br2N4 | 444.13 | 3 / 0 | 4.6 | Yes |
| 3386 | 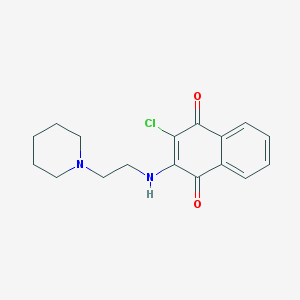 2-chloro-3-[(2-piperidinoethyl)amino]naphthoquinone 2-chloro-3-[(2-piperidinoethyl)amino]naphthoquinone | C17H19ClN2O2 | 318.801 | 4 / 1 | 3.4 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218