

You can:
| Name | Cysteinyl leukotriene receptor 1 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | CYSLTR1 |
| Synonym | LTD4 receptor LTD4 leukotriene D4 receptor HMTMF81 HG55 [ Show all ] |
| Disease | N/A |
| Length | 337 |
| Amino acid sequence | MDETGNLTVSSATCHDTIDDFRNQVYSTLYSMISVVGFFGNGFVLYVLIKTYHKKSAFQVYMINLAVADLLCVCTLPLRVVYYVHKGIWLFGDFLCRLSTYALYVNLYCSIFFMTAMSFFRCIAIVFPVQNINLVTQKKARFVCVGIWIFVILTSSPFLMAKPQKDEKNNTKCFEPPQDNQTKNHVLVLHYVSLFVGFIIPFVIIIVCYTMIILTLLKKSMKKNLSSHKKAIGMIMVVTAAFLVSFMPYHIQRTIHLHFLHNETKPCDSVLRMQKSVVITLSLAASNCCFDPLLYFFSGGNFRKRLSTFRKHSLSSVTYVPRKKASLPEKGEEICKV |
| UniProt | Q9Y271 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | Q9Y271 |
| 3D structure model | This predicted structure model is from GPCR-EXP Q9Y271. |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL1798 |
| IUPHAR | 269 |
| DrugBank | BE0000705 |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 463265 | 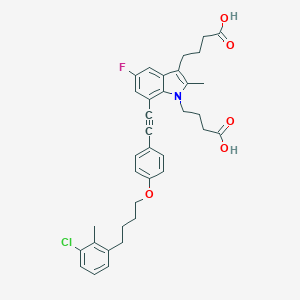 CHEMBL3597629 CHEMBL3597629 | C36H37ClFNO5 | 618.142 | 6 / 2 | 8.1 | No |
| 3333 | 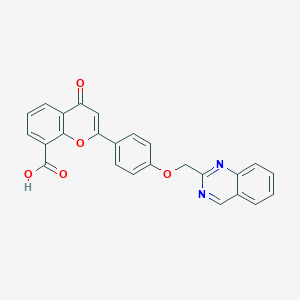 CHEMBL417135 CHEMBL417135 | C25H16N2O5 | 424.412 | 7 / 1 | 3.8 | Yes |
| 521595 | 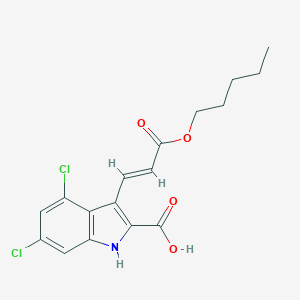 CHEMBL3809152 CHEMBL3809152 | C17H17Cl2NO4 | 370.226 | 4 / 2 | 5.2 | No |
| 463511 | 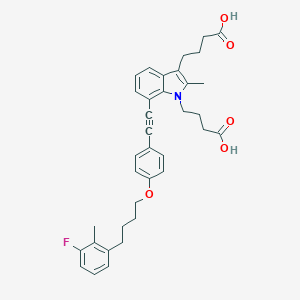 CHEMBL3597633 CHEMBL3597633 | C36H38FNO5 | 583.7 | 6 / 2 | 7.4 | No |
| 6792 | 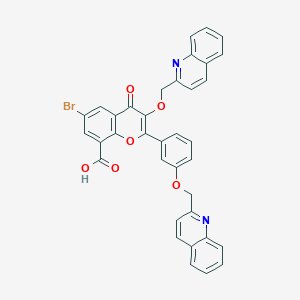 CHEMBL1213921 CHEMBL1213921 | C36H23BrN2O6 | 659.492 | 8 / 1 | 7.1 | No |
| 536127 | 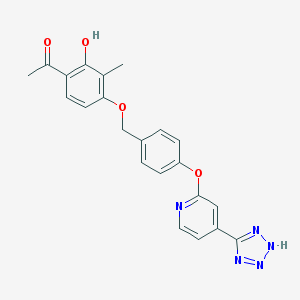 CHEMBL3899832 CHEMBL3899832 | C22H19N5O4 | 417.425 | 8 / 2 | 3.6 | Yes |
| 441994 | 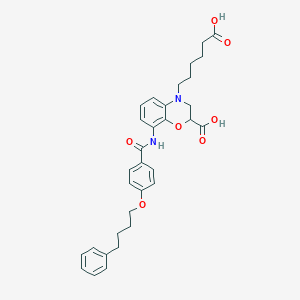 CHEMBL3342957 CHEMBL3342957 | C32H36N2O7 | 560.647 | 8 / 3 | 5.5 | No |
| 8136 | 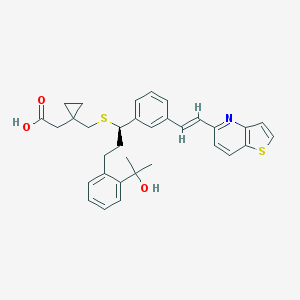 CHEMBL86264 CHEMBL86264 | C33H35NO3S2 | 557.767 | 6 / 2 | 6.9 | No |
| 442062 | 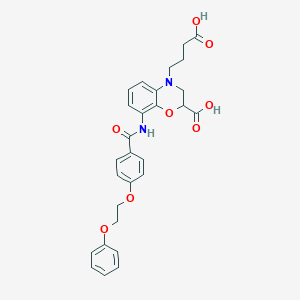 CHEMBL3342960 CHEMBL3342960 | C28H28N2O8 | 520.538 | 9 / 3 | 3.7 | No |
| 464042 | 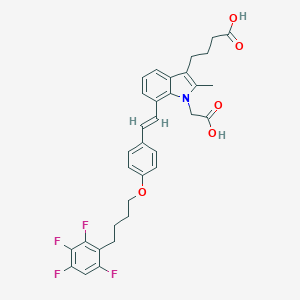 CHEMBL3597533 CHEMBL3597533 | C33H31F4NO5 | 597.607 | 9 / 2 | 7.5 | No |
| 11119 | 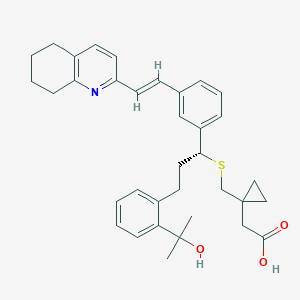 CHEMBL344180 CHEMBL344180 | C35H41NO3S | 555.777 | 5 / 2 | 7.0 | No |
| 442177 | 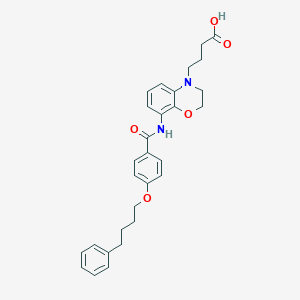 CHEMBL3401692 CHEMBL3401692 | C29H32N2O5 | 488.584 | 6 / 2 | 5.1 | No |
| 12928 | 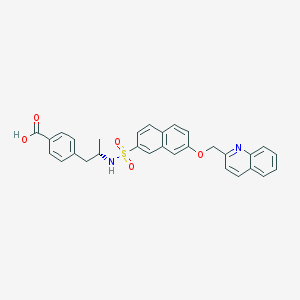 CHEMBL1214530 CHEMBL1214530 | C30H26N2O5S | 526.607 | 7 / 2 | 5.8 | No |
| 13188 | 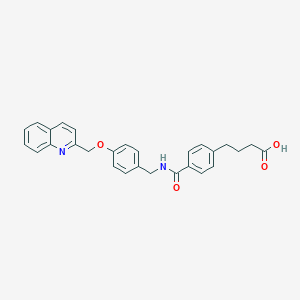 CHEMBL128151 CHEMBL128151 | C28H26N2O4 | 454.526 | 5 / 2 | 4.6 | Yes |
| 14202 | 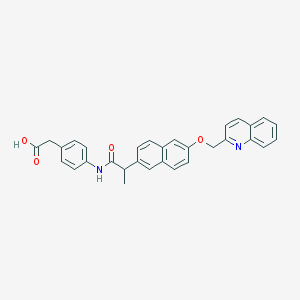 CHEMBL129866 CHEMBL129866 | C31H26N2O4 | 490.559 | 5 / 2 | 5.7 | No |
| 464917 | 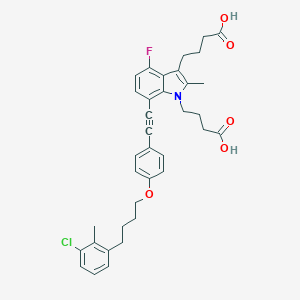 CHEMBL3597628 CHEMBL3597628 | C36H37ClFNO5 | 618.142 | 6 / 2 | 8.1 | No |
| 17092 | 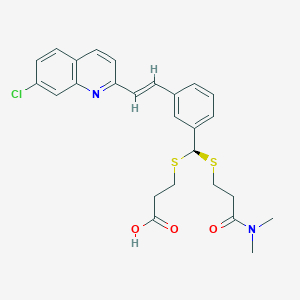 CHEMBL97257 CHEMBL97257 | C26H27ClN2O3S2 | 515.083 | 6 / 1 | 5.3 | No |
| 17093 | 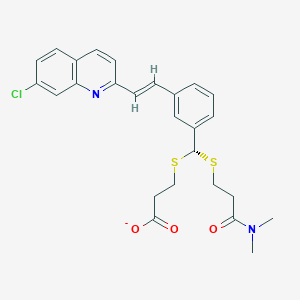 CHEMBL89340 CHEMBL89340 | C26H26ClN2O3S2- | 514.075 | 6 / 0 | 5.9 | No |
| 17096 | 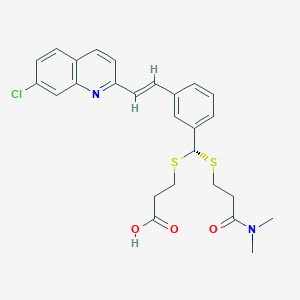 VERLUKAST VERLUKAST | C26H27ClN2O3S2 | 515.083 | 6 / 1 | 5.3 | No |
| 17098 | 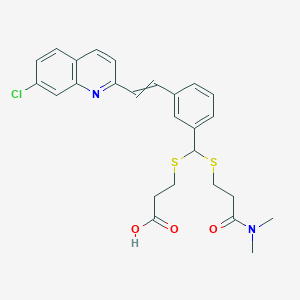 VERLUKAST VERLUKAST | C26H27ClN2O3S2 | 515.083 | 6 / 1 | 5.3 | No |
| 17101 | 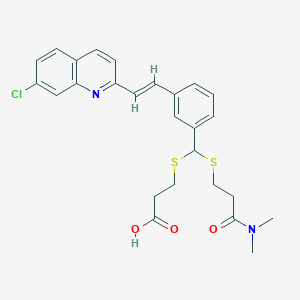 MK 571 MK 571 | C26H27ClN2O3S2 | 515.083 | 6 / 1 | 5.3 | No |
| 17126 | 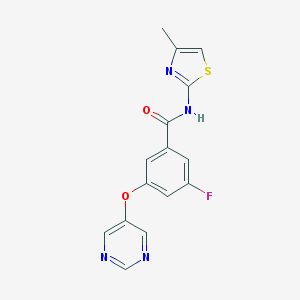 CHEMBL2440659 CHEMBL2440659 | C15H11FN4O2S | 330.337 | 7 / 1 | 2.4 | Yes |
| 522053 | 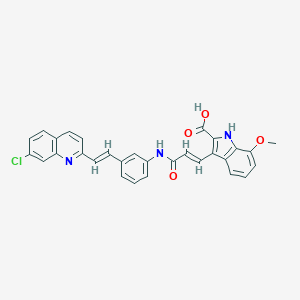 CHEMBL3810210 CHEMBL3810210 | C30H22ClN3O4 | 523.973 | 5 / 3 | 6.3 | No |
| 522078 | 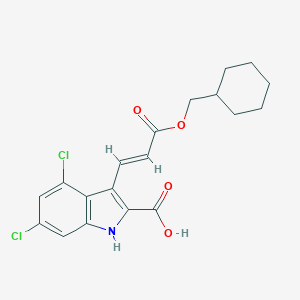 CHEMBL3809862 CHEMBL3809862 | C19H19Cl2NO4 | 396.264 | 4 / 2 | 5.8 | No |
| 465638 | 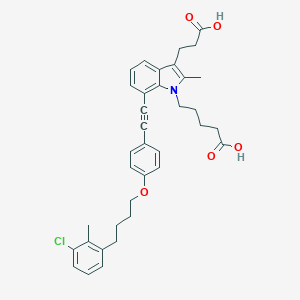 CHEMBL3597626 CHEMBL3597626 | C36H38ClNO5 | 600.152 | 5 / 2 | 8.0 | No |
| 522296 | 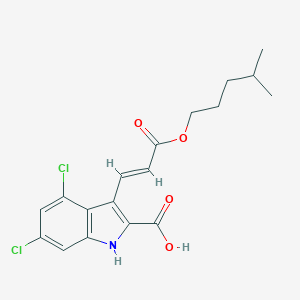 CHEMBL3809106 CHEMBL3809106 | C18H19Cl2NO4 | 384.253 | 4 / 2 | 5.5 | No |
| 24566 | 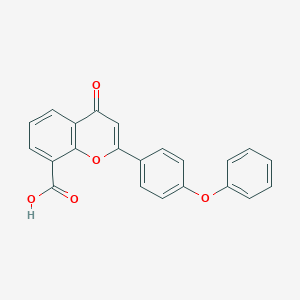 CHEMBL1213844 CHEMBL1213844 | C22H14O5 | 358.349 | 5 / 1 | 4.2 | Yes |
| 522399 | 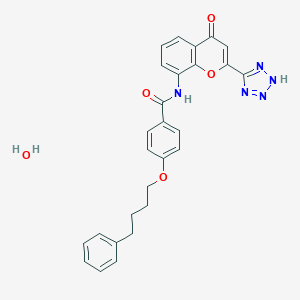 pranlukast hydrate pranlukast hydrate | C27H25N5O5 | 499.527 | 8 / 3 | N/A | N/A |
| 29861 | 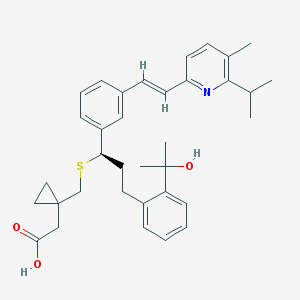 CHEMBL344169 CHEMBL344169 | C35H43NO3S | 557.793 | 5 / 2 | 7.3 | No |
| 466759 | 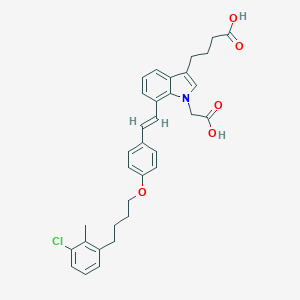 CHEMBL3597527 CHEMBL3597527 | C33H34ClNO5 | 560.087 | 5 / 2 | 7.7 | No |
| 31474 | 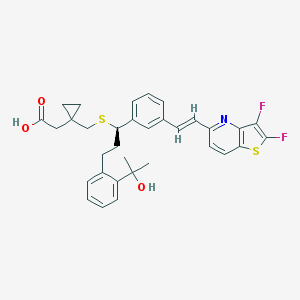 CHEMBL314237 CHEMBL314237 | C33H33F2NO3S2 | 593.748 | 8 / 2 | 7.4 | No |
| 31896 | 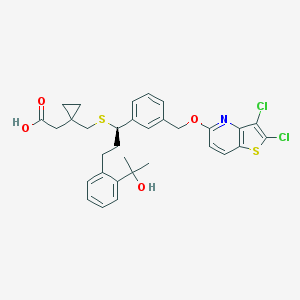 CHEMBL313544 CHEMBL313544 | C32H33Cl2NO4S2 | 630.639 | 7 / 2 | 7.9 | No |
| 35162 | 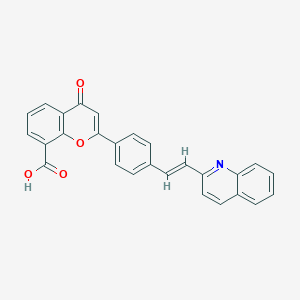 CHEMBL32764 CHEMBL32764 | C27H17NO4 | 419.436 | 5 / 1 | 5.3 | No |
| 443119 | 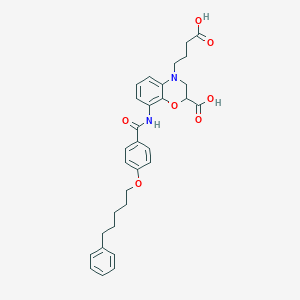 CHEMBL3342959 CHEMBL3342959 | C31H34N2O7 | 546.62 | 8 / 3 | 5.3 | No |
| 36595 | 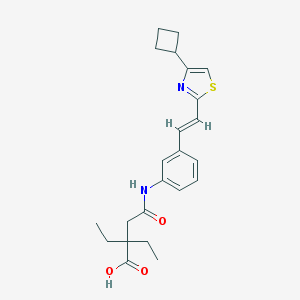 Cinalukast Cinalukast | C23H28N2O3S | 412.548 | 5 / 2 | 4.6 | Yes |
| 443213 | 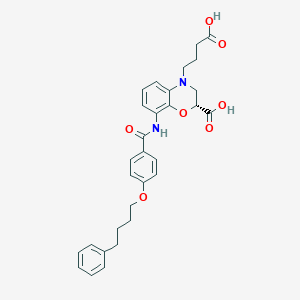 CHEMBL3342963 CHEMBL3342963 | C30H32N2O7 | 532.593 | 8 / 3 | 4.7 | No |
| 443215 | 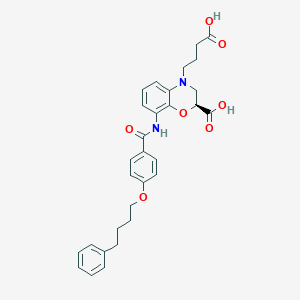 CHEMBL3342964 CHEMBL3342964 | C30H32N2O7 | 532.593 | 8 / 3 | 4.7 | No |
| 443216 | 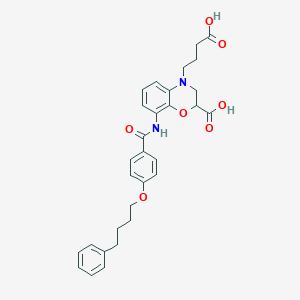 CHEMBL3342955 CHEMBL3342955 | C30H32N2O7 | 532.593 | 8 / 3 | 4.7 | No |
| 42145 | 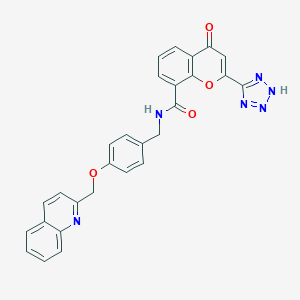 CHEMBL131287 CHEMBL131287 | C28H20N6O4 | 504.506 | 8 / 2 | 3.3 | No |
| 553456 | 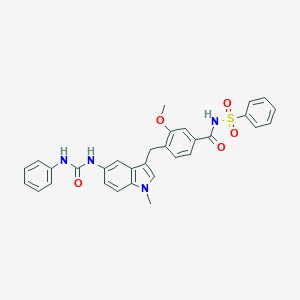 GTPL10002 GTPL10002 | C31H28N4O5S | 568.648 | 5 / 3 | 4.9 | No |
| 537112 | 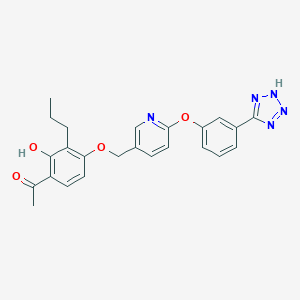 CHEMBL3920609 CHEMBL3920609 | C24H23N5O4 | 445.479 | 8 / 2 | 4.6 | Yes |
| 44234 | 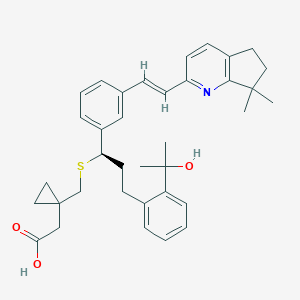 CHEMBL140381 CHEMBL140381 | C36H43NO3S | 569.804 | 5 / 2 | 7.3 | No |
| 537129 | 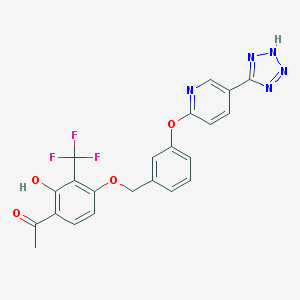 CHEMBL3973048 CHEMBL3973048 | C22H16F3N5O4 | 471.396 | 11 / 2 | 4.2 | No |
| 522857 | 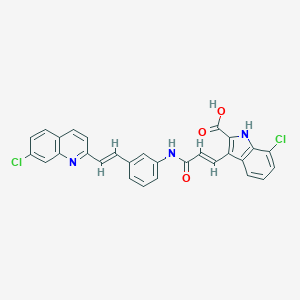 CHEMBL3809483 CHEMBL3809483 | C29H19Cl2N3O3 | 528.389 | 4 / 3 | 7.0 | No |
| 443534 | 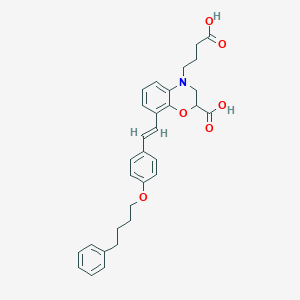 CHEMBL3401687 CHEMBL3401687 | C31H33NO6 | 515.606 | 7 / 2 | 6.2 | No |
| 47603 | 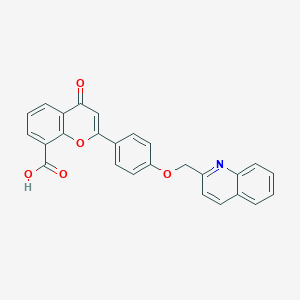 CHEMBL33294 CHEMBL33294 | C26H17NO5 | 423.424 | 6 / 1 | 4.5 | Yes |
| 469103 | 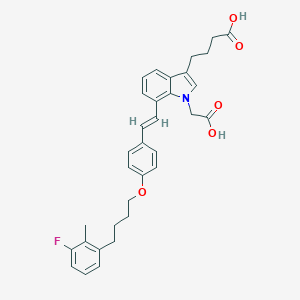 CHEMBL3597526 CHEMBL3597526 | C33H34FNO5 | 543.635 | 6 / 2 | 7.2 | No |
| 51383 | 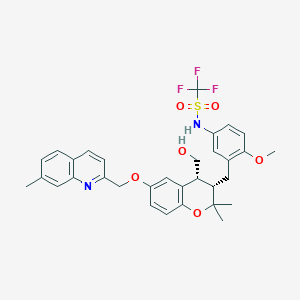 CHEMBL1214531 CHEMBL1214531 | C32H33F3N2O6S | 630.679 | 11 / 2 | 6.1 | No |
| 443867 | 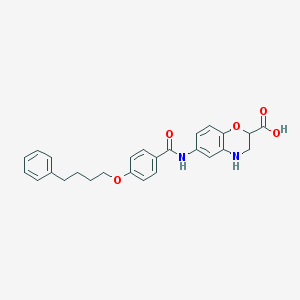 CHEMBL3342949 CHEMBL3342949 | C26H26N2O5 | 446.503 | 6 / 3 | 4.8 | Yes |
| 55972 | 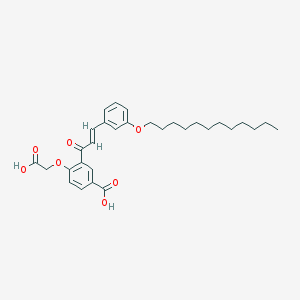 CHEMBL2413379 CHEMBL2413379 | C30H38O7 | 510.627 | 7 / 2 | 8.3 | No |
| 559013 | 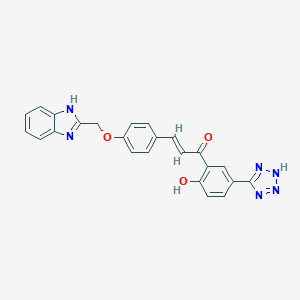 CHEMBL1214415 CHEMBL1214415 | C24H18N6O3 | 438.447 | 7 / 3 | 4.2 | Yes |
| 470596 | 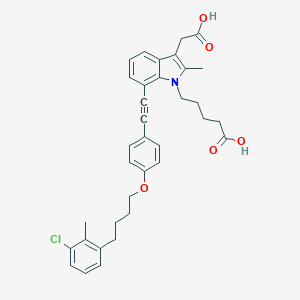 CHEMBL3597625 CHEMBL3597625 | C35H36ClNO5 | 586.125 | 5 / 2 | 7.7 | No |
| 63654 | 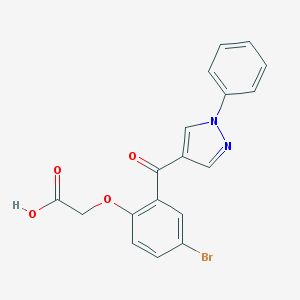 CHEMBL217053 CHEMBL217053 | C18H13BrN2O4 | 401.216 | 5 / 1 | 3.5 | Yes |
| 444269 | 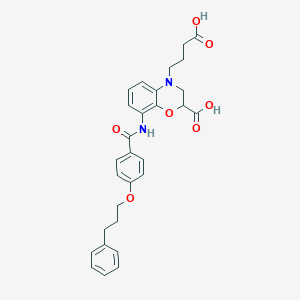 CHEMBL3342958 CHEMBL3342958 | C29H30N2O7 | 518.566 | 8 / 3 | 4.4 | No |
| 67193 | 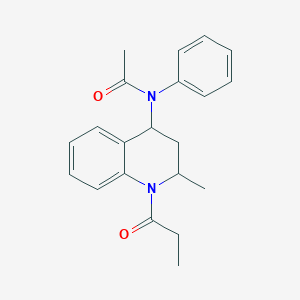 Probes1_000120 Probes1_000120 | C21H24N2O2 | 336.435 | 2 / 0 | 3.2 | Yes |
| 471384 | 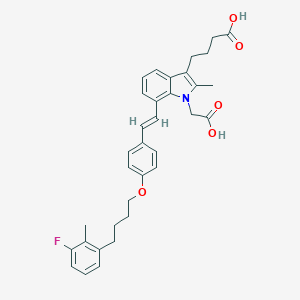 CHEMBL3597534 CHEMBL3597534 | C34H36FNO5 | 557.662 | 6 / 2 | 7.6 | No |
| 68893 | 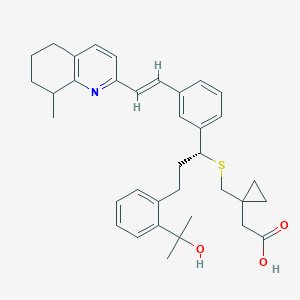 CHEMBL342705 CHEMBL342705 | C36H43NO3S | 569.804 | 5 / 2 | 7.3 | No |
| 471518 | 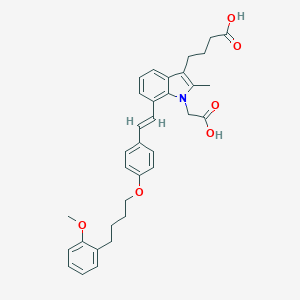 CHEMBL3597529 CHEMBL3597529 | C34H37NO6 | 555.671 | 6 / 2 | 7.1 | No |
| 70838 | 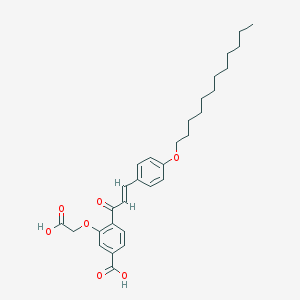 CHEMBL2413270 CHEMBL2413270 | C30H38O7 | 510.627 | 7 / 2 | 8.3 | No |
| 76198 | 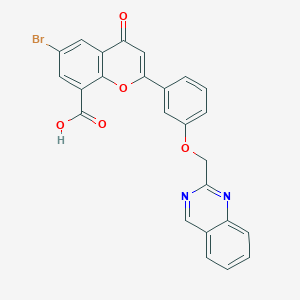 CHEMBL34740 CHEMBL34740 | C25H15BrN2O5 | 503.308 | 7 / 1 | 4.5 | No |
| 77765 | 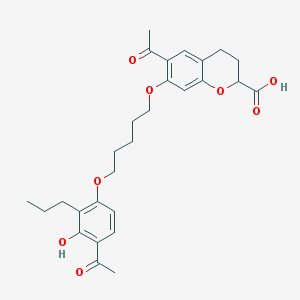 Ablukast Ablukast | C28H34O8 | 498.572 | 8 / 2 | 5.5 | No |
| 472559 | 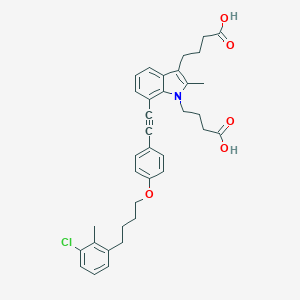 CHEMBL3597624 CHEMBL3597624 | C36H38ClNO5 | 600.152 | 5 / 2 | 8.0 | No |
| 79083 | 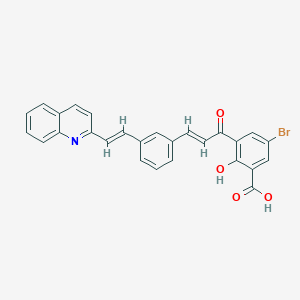 CHEMBL11349 CHEMBL11349 | C27H18BrNO4 | 500.348 | 5 / 2 | 6.7 | No |
| 523846 | 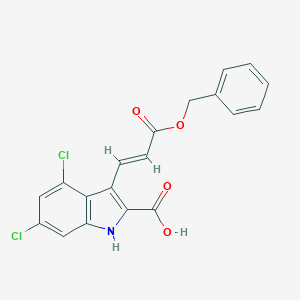 CHEMBL3810094 CHEMBL3810094 | C19H13Cl2NO4 | 390.216 | 4 / 2 | 4.9 | Yes |
| 444921 | 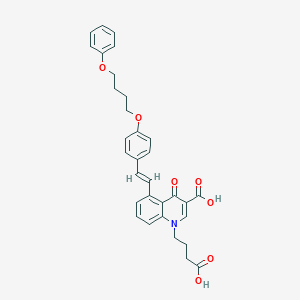 CHEMBL3403188 CHEMBL3403188 | C32H31NO7 | 541.6 | 8 / 2 | 6.5 | No |
| 81899 | 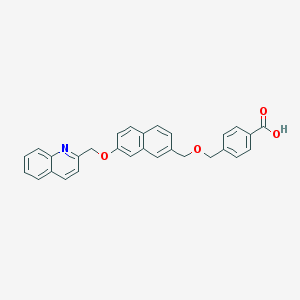 CHEMBL127842 CHEMBL127842 | C29H23NO4 | 449.506 | 5 / 1 | 5.6 | No |
| 82442 | 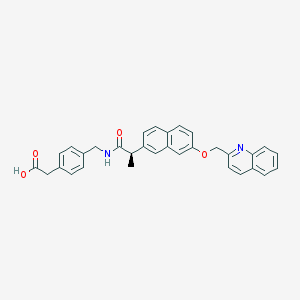 CHEMBL1214642 CHEMBL1214642 | C32H28N2O4 | 504.586 | 5 / 2 | 5.7 | No |
| 83203 | 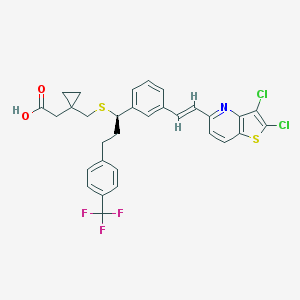 CHEMBL84841 CHEMBL84841 | C31H26Cl2F3NO2S2 | 636.569 | 8 / 1 | 9.6 | No |
| 553690 | 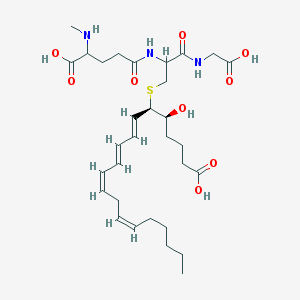 N-Methyl ltc4 N-Methyl ltc4 | C31H49N3O9S | 639.805 | 11 / 7 | 1.2 | No |
| 86631 | 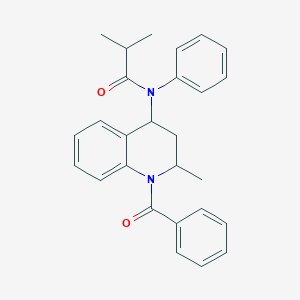 2-methyl-N-[2-methyl-1-(phenylcarbonyl)-1,2,3,4-tetrahydroquinolin-4-yl]-N-phenylpropanamide 2-methyl-N-[2-methyl-1-(phenylcarbonyl)-1,2,3,4-tetrahydroquinolin-4-yl]-N-phenylpropanamide | C27H28N2O2 | 412.533 | 2 / 0 | 5.4 | No |
| 473962 | 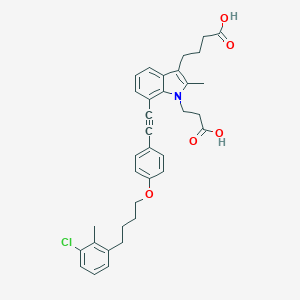 CHEMBL3597621 CHEMBL3597621 | C35H36ClNO5 | 586.125 | 5 / 2 | 7.6 | No |
| 538251 | 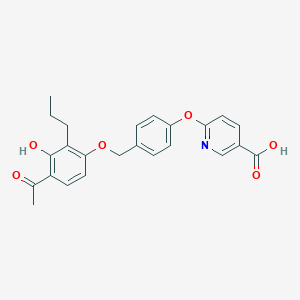 CHEMBL3935529 CHEMBL3935529 | C24H23NO6 | 421.449 | 7 / 2 | 4.9 | Yes |
| 445441 | 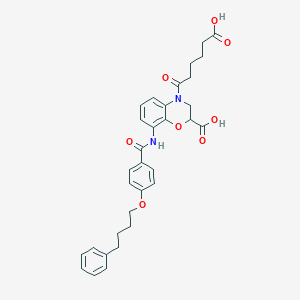 CHEMBL3342954 CHEMBL3342954 | C32H34N2O8 | 574.63 | 8 / 3 | 4.3 | No |
| 98870 | 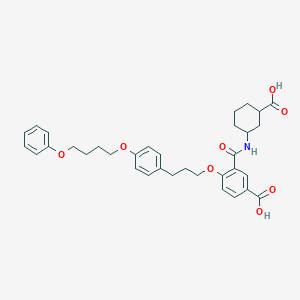 CysLT2cpd CysLT2cpd | C34H39NO8 | 589.685 | 8 / 3 | 6.2 | No |
| 538510 | 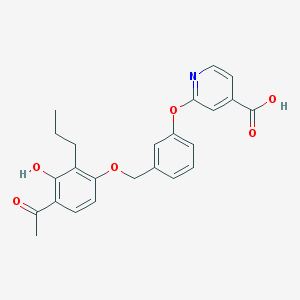 CHEMBL3890848 CHEMBL3890848 | C24H23NO6 | 421.449 | 7 / 2 | 4.9 | Yes |
| 103875 | 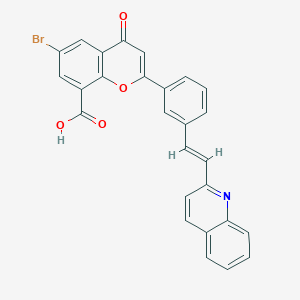 CHEMBL32136 CHEMBL32136 | C27H16BrNO4 | 498.332 | 5 / 1 | 6.0 | No |
| 106285 | 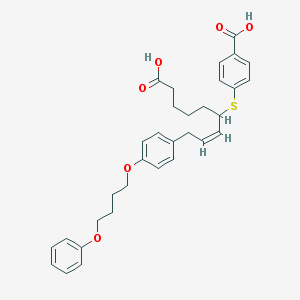 CHEMBL36607 CHEMBL36607 | C32H36O6S | 548.694 | 7 / 2 | 7.3 | No |
| 107097 | 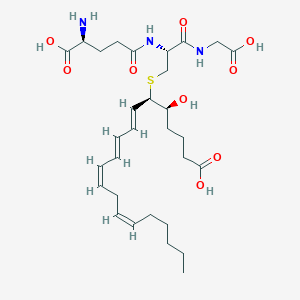 leukotriene C4 leukotriene C4 | C30H47N3O9S | 625.778 | 11 / 7 | 0.7 | No |
| 524660 | 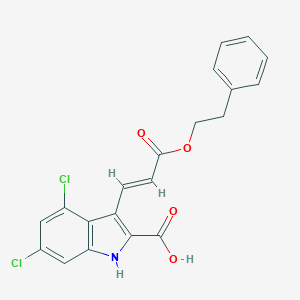 CHEMBL3810389 CHEMBL3810389 | C20H15Cl2NO4 | 404.243 | 4 / 2 | 5.4 | No |
| 524802 | 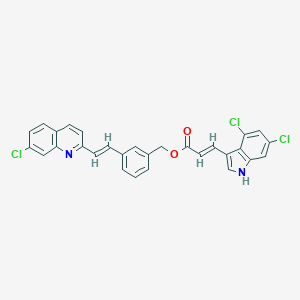 CHEMBL3810142 CHEMBL3810142 | C29H19Cl3N2O2 | 533.833 | 3 / 1 | 8.3 | No |
| 477548 | 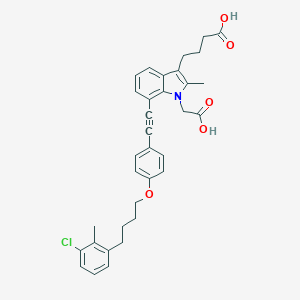 CHEMBL3597618 CHEMBL3597618 | C34H34ClNO5 | 572.098 | 5 / 2 | 7.9 | No |
| 477749 | 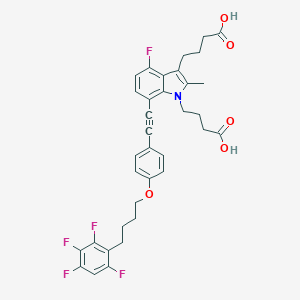 CHEMBL3597632 CHEMBL3597632 | C35H32F5NO5 | 641.635 | 10 / 2 | 7.5 | No |
| 446494 | 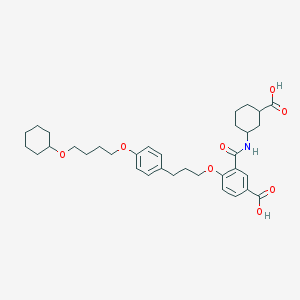 HAMI3379 HAMI3379 | C34H45NO8 | 595.733 | 8 / 3 | 6.2 | No |
| 446561 | 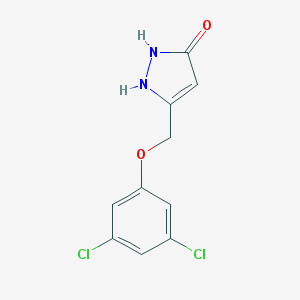 CHEMBL1939222 CHEMBL1939222 | C10H8Cl2N2O2 | 259.086 | 3 / 2 | 2.5 | Yes |
| 478242 | 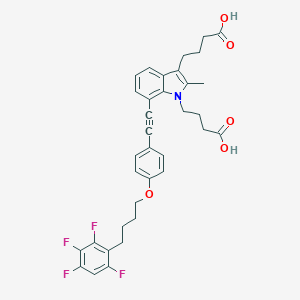 CHEMBL3597631 CHEMBL3597631 | C35H33F4NO5 | 623.645 | 9 / 2 | 7.4 | No |
| 125606 | 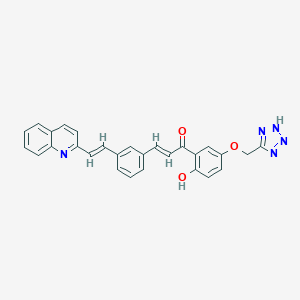 CHEMBL11870 CHEMBL11870 | C28H21N5O3 | 475.508 | 7 / 2 | 5.5 | No |
| 125607 | 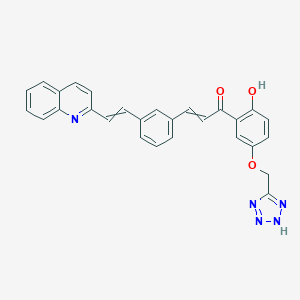 CID 57035651 CID 57035651 | C28H21N5O3 | 475.508 | 7 / 2 | 5.5 | No |
| 128982 | 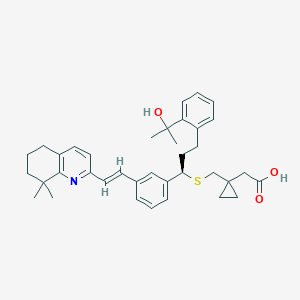 CHEMBL348229 CHEMBL348229 | C37H45NO3S | 583.831 | 5 / 2 | 7.9 | No |
| 539275 | 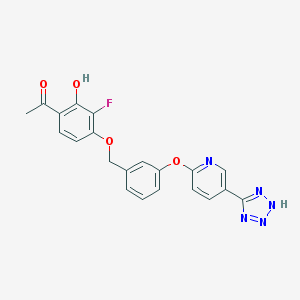 CHEMBL3943547 CHEMBL3943547 | C21H16FN5O4 | 421.388 | 9 / 2 | 3.4 | Yes |
| 525294 | 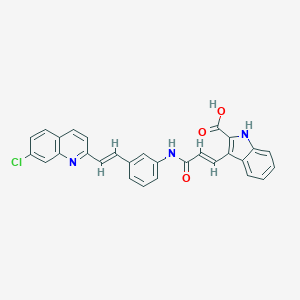 CHEMBL3809230 CHEMBL3809230 | C29H20ClN3O3 | 493.947 | 4 / 3 | 6.4 | No |
| 132042 | 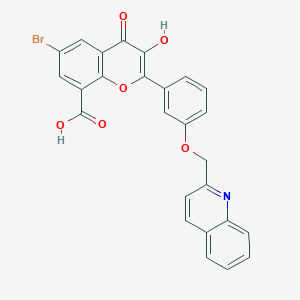 CHEMBL1213919 CHEMBL1213919 | C26H16BrNO6 | 518.319 | 7 / 2 | 5.0 | No |
| 132893 | 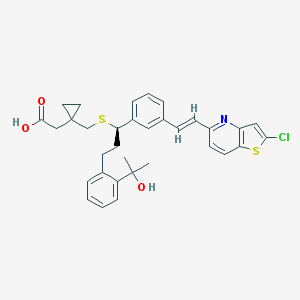 CHEMBL314340 CHEMBL314340 | C33H34ClNO3S2 | 592.209 | 6 / 2 | 7.8 | No |
| 134190 | 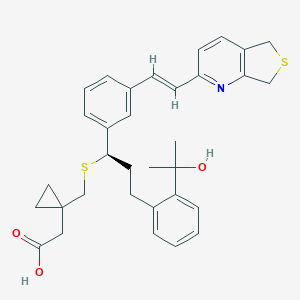 CHEMBL139757 CHEMBL139757 | C33H37NO3S2 | 559.783 | 6 / 2 | 5.8 | No |
| 447074 | 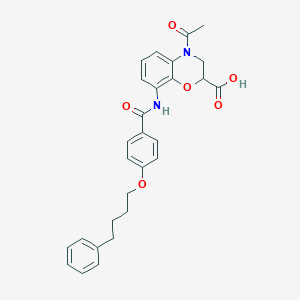 CHEMBL3342950 CHEMBL3342950 | C28H28N2O6 | 488.54 | 6 / 2 | 4.1 | Yes |
| 137051 | 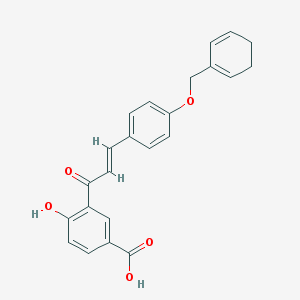 CHEMBL1213782 CHEMBL1213782 | C23H20O5 | 376.408 | 5 / 2 | 4.7 | Yes |
| 447216 | 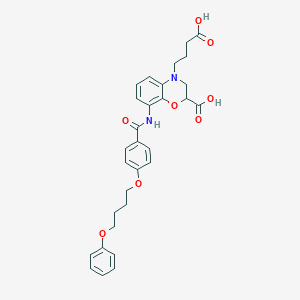 CHEMBL3342962 CHEMBL3342962 | C30H32N2O8 | 548.592 | 9 / 3 | 4.5 | No |
| 141612 | 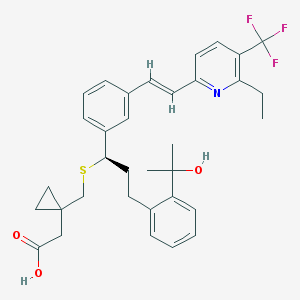 CHEMBL143115 CHEMBL143115 | C34H38F3NO3S | 597.737 | 8 / 2 | 7.5 | No |
| 553999 | 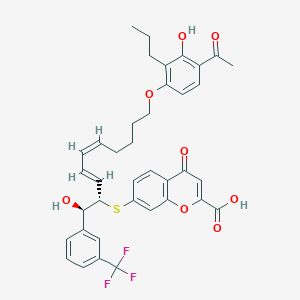 Iralukast Iralukast | C38H37F3O8S | 710.761 | 12 / 3 | 8.6 | No |
| 480808 | 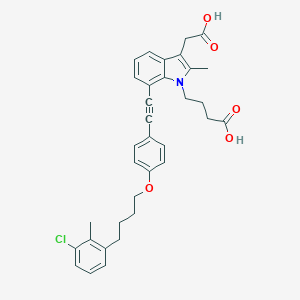 CHEMBL3597622 CHEMBL3597622 | C34H34ClNO5 | 572.098 | 5 / 2 | 7.3 | No |
| 148532 | 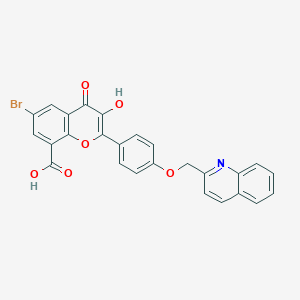 CHEMBL1213720 CHEMBL1213720 | C26H16BrNO6 | 518.319 | 7 / 2 | 5.0 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218