

You can:
| Name | Muscarinic acetylcholine receptor M5 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | CHRM5 |
| Synonym | M5R M5 receptor cholinergic receptor cholinergic receptor, muscarinic 5 |
| Disease | Urinary incontinence Colitis Dysmenorrhea Irritable bowel syndrome Myasthenia gravis [ Show all ] |
| Length | 532 |
| Amino acid sequence | MEGDSYHNATTVNGTPVNHQPLERHRLWEVITIAAVTAVVSLITIVGNVLVMISFKVNSQLKTVNNYYLLSLACADLIIGIFSMNLYTTYILMGRWALGSLACDLWLALDYVASNASVMNLLVISFDRYFSITRPLTYRAKRTPKRAGIMIGLAWLISFILWAPAILCWQYLVGKRTVPLDECQIQFLSEPTITFGTAIAAFYIPVSVMTILYCRIYRETEKRTKDLADLQGSDSVTKAEKRKPAHRALFRSCLRCPRPTLAQRERNQASWSSSRRSTSTTGKPSQATGPSANWAKAEQLTTCSSYPSSEDEDKPATDPVLQVVYKSQGKESPGEEFSAEETEETFVKAETEKSDYDTPNYLLSPAAAHRPKSQKCVAYKFRLVVKADGNQETNNGCHKVKIMPCPFPVAKEPSTKGLNPNPSHQMTKRKRVVLVKERKAAQTLSAILLAFIITWTPYNIMVLVSTFCDKCVPVTLWHLGYWLCYVNSTVNPICYALCNRTFRKTFKMLLLCRWKKKKVEEKLYWQGNSKLP |
| UniProt | P08912 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | T79961 |
| ChEMBL | CHEMBL2035 |
| IUPHAR | 17 |
| DrugBank | BE0000247, BE0004890 |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 3 | 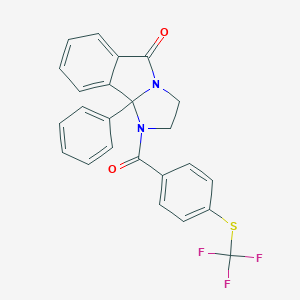 CHEMBL3105203 CHEMBL3105203 | C24H17F3N2O2S | 454.467 | 6 / 0 | 5.2 | No |
| 441679 | 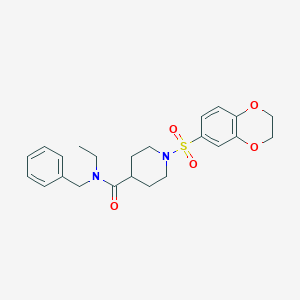 CHEMBL1602658 CHEMBL1602658 | C23H28N2O5S | 444.546 | 6 / 0 | 2.6 | Yes |
| 522 | 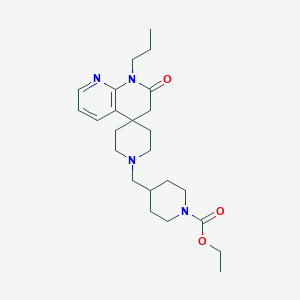 CHEMBL2407175 CHEMBL2407175 | C24H36N4O3 | 428.577 | 5 / 0 | 2.7 | Yes |
| 1023 | 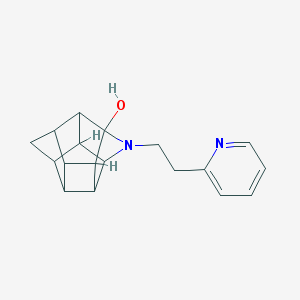 CHEMBL2205816 CHEMBL2205816 | C18H20N2O | 280.371 | 3 / 1 | 1.8 | Yes |
| 2259 | 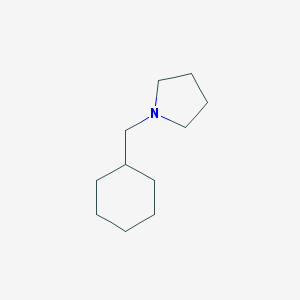 1-(cyclohexylmethyl)pyrrolidine 1-(cyclohexylmethyl)pyrrolidine | C11H21N | 167.296 | 1 / 0 | 3.1 | Yes |
| 3128 | 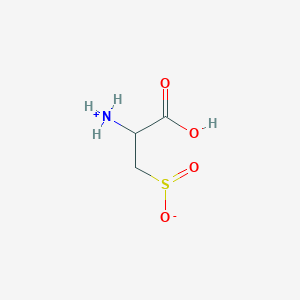 AC1N0YGA AC1N0YGA | C3H7NO4S | 153.152 | 5 / 2 | -4.7 | Yes |
| 3550 | 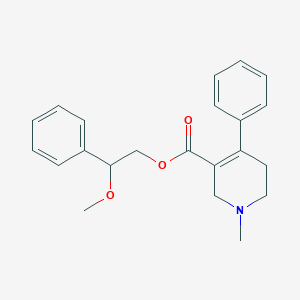 CHEMBL2312348 CHEMBL2312348 | C22H25NO3 | 351.446 | 4 / 0 | 3.1 | Yes |
| 4008 | 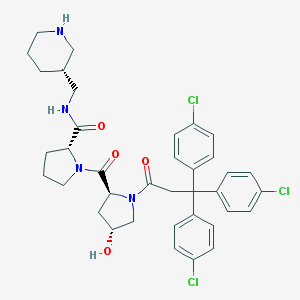 CHEMBL215894 CHEMBL215894 | C37H41Cl3N4O4 | 712.109 | 5 / 3 | 5.9 | No |
| 4318 | 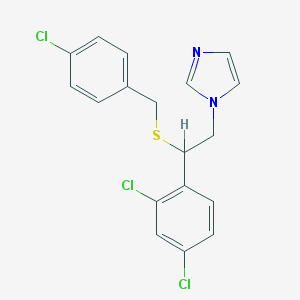 sulconazole sulconazole | C18H15Cl3N2S | 397.742 | 2 / 0 | 6.1 | No |
| 4918 | 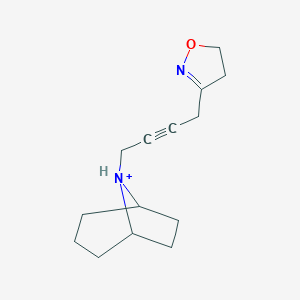 CHEMBL239657 CHEMBL239657 | C14H21N2O+ | 233.335 | 2 / 1 | 1.8 | Yes |
| 4986 | 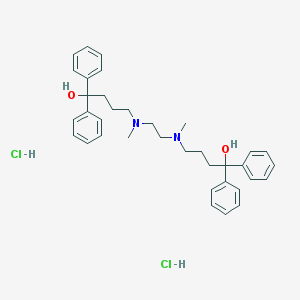 CHEMBL255692 CHEMBL255692 | C36H46Cl2N2O2 | 609.676 | 4 / 4 | N/A | No |
| 5666 | 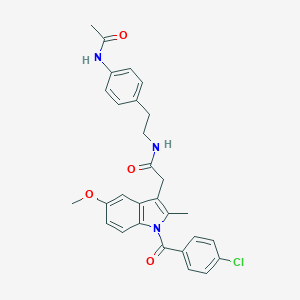 AC1L1HUK AC1L1HUK | C29H28ClN3O4 | 518.01 | 4 / 2 | 5.2 | No |
| 5747 | 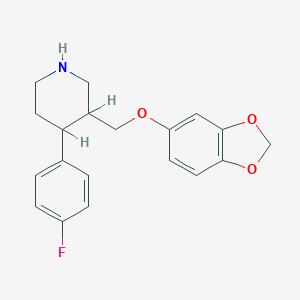 AHOUBRCZNHFOSL-UHFFFAOYSA-N AHOUBRCZNHFOSL-UHFFFAOYSA-N | C19H20FNO3 | 329.371 | 5 / 1 | 3.5 | Yes |
| 5759 | 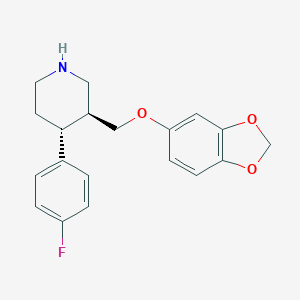 paroxetine paroxetine | C19H20FNO3 | 329.371 | 5 / 1 | 3.5 | Yes |
| 459279 | 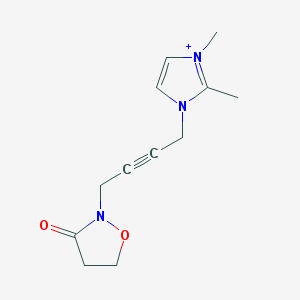 BDBM86295 BDBM86295 | C12H16N3O2+ | 234.279 | 2 / 0 | 0.0 | Yes |
| 6612 | 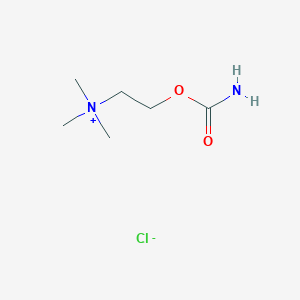 carbachol carbachol | C6H15ClN2O2 | 182.648 | 3 / 1 | N/A | N/A |
| 6685 | 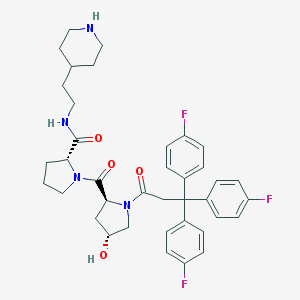 CHEMBL214029 CHEMBL214029 | C38H43F3N4O4 | 676.781 | 8 / 3 | 4.7 | No |
| 6836 | 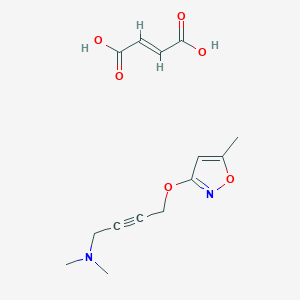 CID 49797317 CID 49797317 | C14H18N2O6 | 310.306 | 8 / 2 | N/A | N/A |
| 463725 | 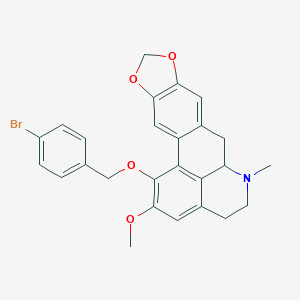 CHEMBL3633757 CHEMBL3633757 | C26H24BrNO4 | 494.385 | 5 / 0 | 5.4 | No |
| 7190 | 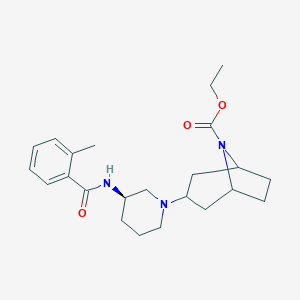 CHEMBL2024331 CHEMBL2024331 | C23H33N3O3 | 399.535 | 4 / 1 | 3.6 | Yes |
| 7781 | 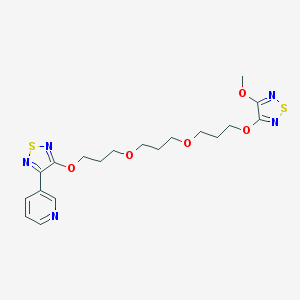 CHEMBL376295 CHEMBL376295 | C19H25N5O5S2 | 467.559 | 12 / 0 | 2.8 | No |
| 9533 | 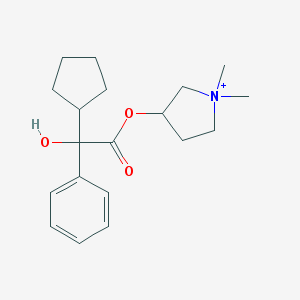 Glycopyrronium Glycopyrronium | C19H28NO3+ | 318.437 | 3 / 1 | 3.1 | Yes |
| 9854 | 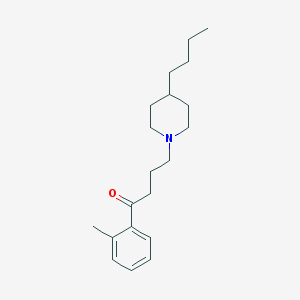 AC-42 AC-42 | C20H31NO | 301.474 | 2 / 0 | 5.2 | No |
| 10179 | 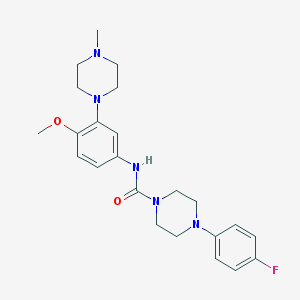 CHEMBL194206 CHEMBL194206 | C23H30FN5O2 | 427.524 | 6 / 1 | 2.7 | Yes |
| 442116 | 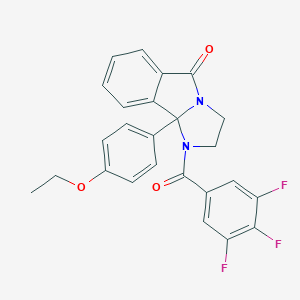 CHEMBL3353272 CHEMBL3353272 | C25H19F3N2O3 | 452.433 | 6 / 0 | 4.1 | Yes |
| 10390 | 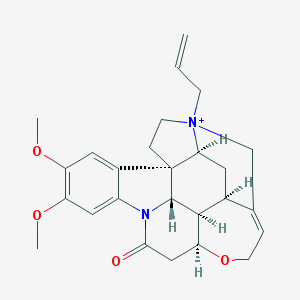 N-Allyl Brucine Iodide N-Allyl Brucine Iodide | C26H31N2O4+ | 435.544 | 4 / 0 | 1.6 | Yes |
| 10705 | 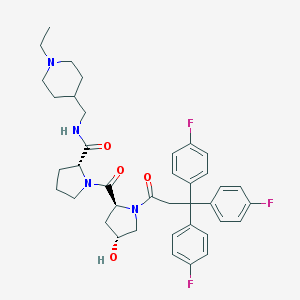 CHEMBL213709 CHEMBL213709 | C39H45F3N4O4 | 690.808 | 8 / 2 | 5.1 | No |
| 10749 | 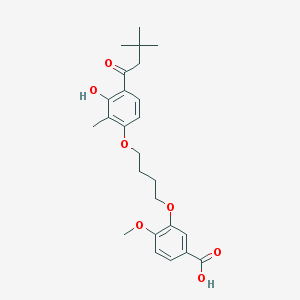 CHEMBL3287723 CHEMBL3287723 | C25H32O7 | 444.524 | 7 / 2 | 5.7 | No |
| 10910 | 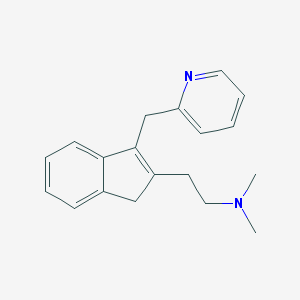 CHEMBL169494 CHEMBL169494 | C19H22N2 | 278.399 | 2 / 0 | 2.4 | Yes |
| 442147 | 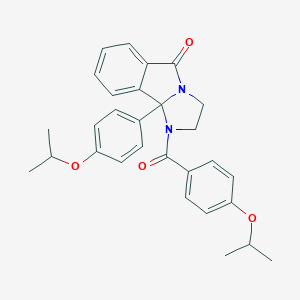 CHEMBL3353282 CHEMBL3353282 | C29H30N2O4 | 470.569 | 4 / 0 | 5.0 | Yes |
| 11495 | 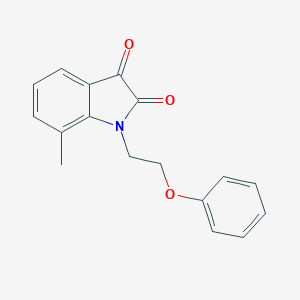 MLS000624518 MLS000624518 | C17H15NO3 | 281.311 | 3 / 0 | 2.9 | Yes |
| 11914 | 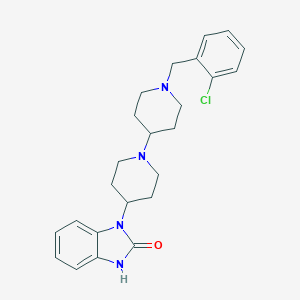 CHEMBL477821 CHEMBL477821 | C24H29ClN4O | 424.973 | 3 / 1 | 4.1 | Yes |
| 12545 | 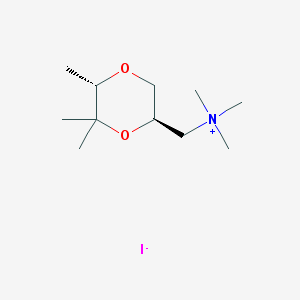 CHEMBL593620 CHEMBL593620 | C11H24INO2 | 329.222 | 3 / 0 | N/A | N/A |
| 12550 | 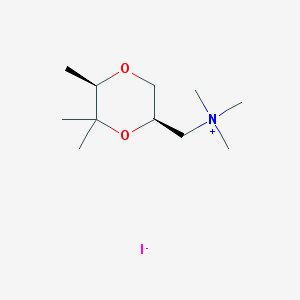 CHEMBL593685 CHEMBL593685 | C11H24INO2 | 329.222 | 3 / 0 | N/A | N/A |
| 12704 | 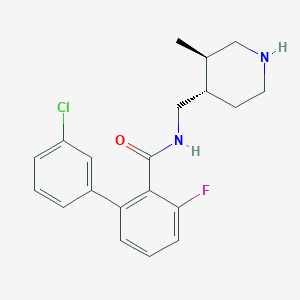 CHEMBL1083925 CHEMBL1083925 | C20H22ClFN2O | 360.857 | 3 / 2 | 4.3 | Yes |
| 13059 | 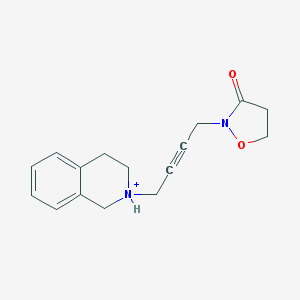 CHEMBL397538 CHEMBL397538 | C16H19N2O2+ | 271.34 | 2 / 1 | 1.1 | Yes |
| 13306 | 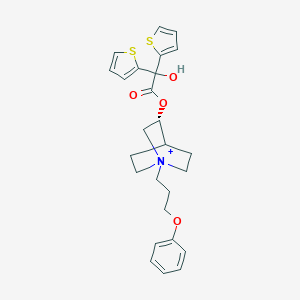 Aclidinium Aclidinium | C26H30NO4S2+ | 484.649 | 6 / 1 | 4.7 | Yes |
| 13775 | 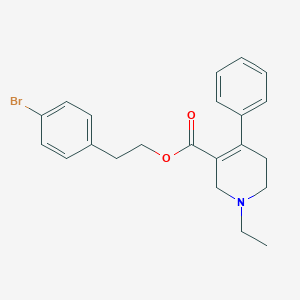 CHEMBL2312373 CHEMBL2312373 | C22H24BrNO2 | 414.343 | 3 / 0 | 4.7 | Yes |
| 13785 | 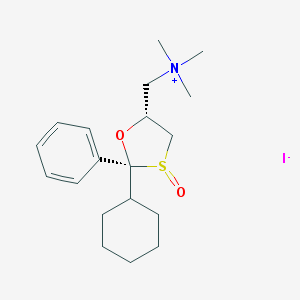 CHEMBL557808 CHEMBL557808 | C19H30INO2S | 463.418 | 4 / 0 | N/A | N/A |
| 14122 | 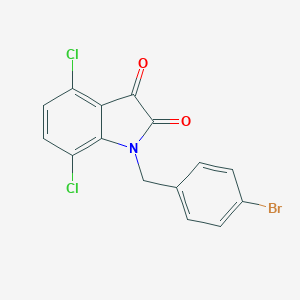 CHEMBL466851 CHEMBL466851 | C15H8BrCl2NO2 | 385.038 | 2 / 0 | 4.3 | Yes |
| 14780 | 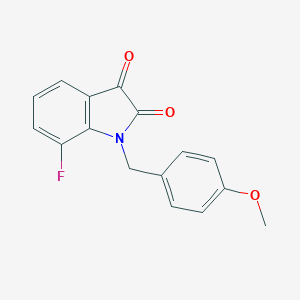 CHEMBL443334 CHEMBL443334 | C16H12FNO3 | 285.274 | 4 / 0 | 2.5 | Yes |
| 15154 | 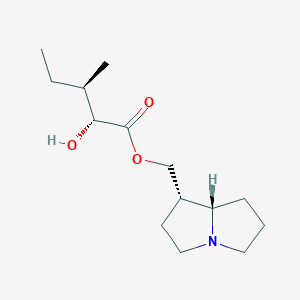 cremastrine cremastrine | C14H25NO3 | 255.358 | 4 / 1 | 2.2 | Yes |
| 15431 | 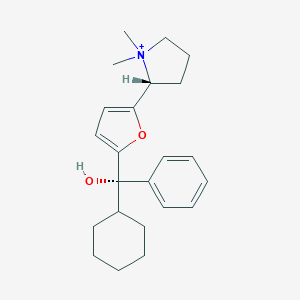 CHEMBL568775 CHEMBL568775 | C23H32NO2+ | 354.514 | 2 / 1 | 4.4 | Yes |
| 15432 | 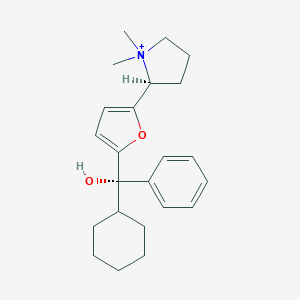 CHEMBL569089 CHEMBL569089 | C23H32NO2+ | 354.514 | 2 / 1 | 4.4 | Yes |
| 15439 | 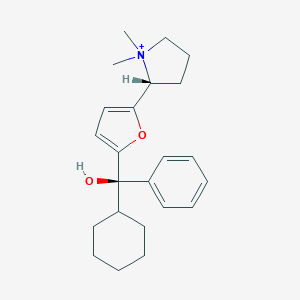 CHEMBL569307 CHEMBL569307 | C23H32NO2+ | 354.514 | 2 / 1 | 4.4 | Yes |
| 15444 | 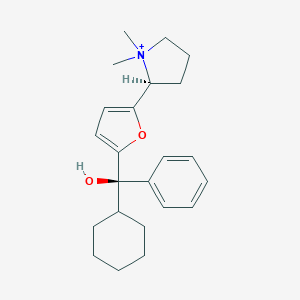 CHEMBL568870 CHEMBL568870 | C23H32NO2+ | 354.514 | 2 / 1 | 4.4 | Yes |
| 15449 | 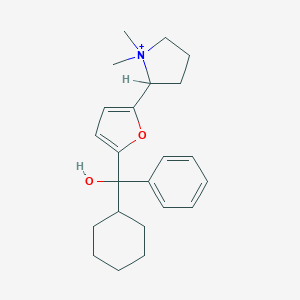 CHEMBL569760 CHEMBL569760 | C23H32NO2+ | 354.514 | 2 / 1 | 4.4 | Yes |
| 15472 | 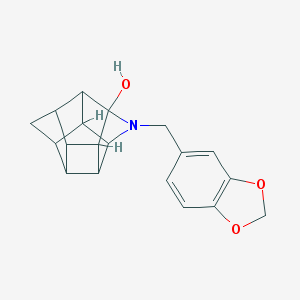 CHEMBL2205829 CHEMBL2205829 | C19H19NO3 | 309.365 | 4 / 1 | 2.2 | Yes |
| 15786 | 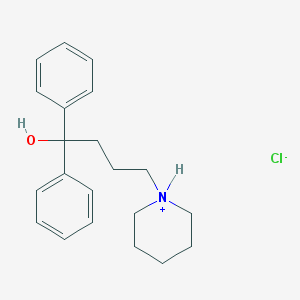 CID 44448670 CID 44448670 | C21H28ClNO | 345.911 | 2 / 2 | N/A | N/A |
| 442266 | 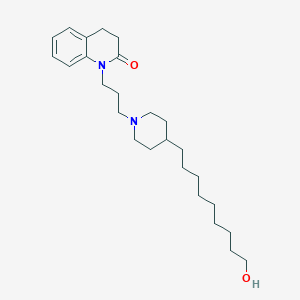 CHEMBL3354075 CHEMBL3354075 | C26H42N2O2 | 414.634 | 3 / 1 | 5.7 | No |
| 16049 | 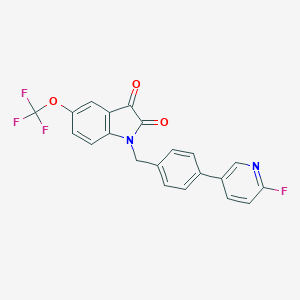 CHEMBL596509 CHEMBL596509 | C21H12F4N2O3 | 416.332 | 8 / 0 | 4.6 | Yes |
| 442305 | 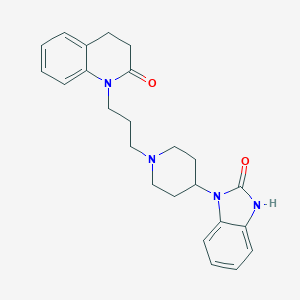 CHEMBL2059306 CHEMBL2059306 | C24H28N4O2 | 404.514 | 3 / 1 | 3.8 | Yes |
| 16957 | 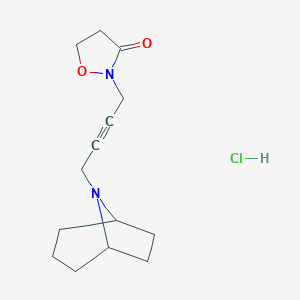 CHEMBL239865 CHEMBL239865 | C14H21ClN2O2 | 284.784 | 3 / 1 | N/A | N/A |
| 442338 | 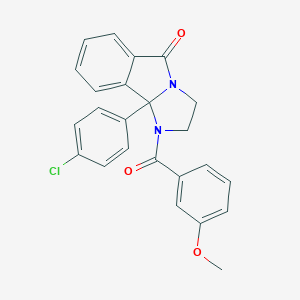 CHEMBL3361316 CHEMBL3361316 | C24H19ClN2O3 | 418.877 | 3 / 0 | 4.1 | Yes |
| 17717 | 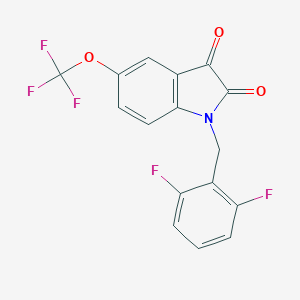 CHEMBL504572 CHEMBL504572 | C16H8F5NO3 | 357.236 | 8 / 0 | 3.8 | Yes |
| 19085 | 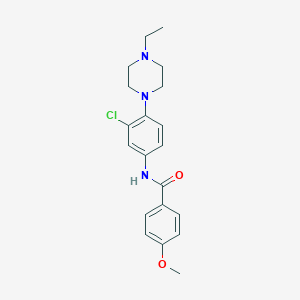 BAS 07465501 BAS 07465501 | C20H24ClN3O2 | 373.881 | 4 / 1 | 3.7 | Yes |
| 442434 | 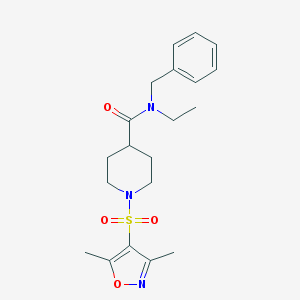 CHEMBL3329723 CHEMBL3329723 | C20H27N3O4S | 405.513 | 6 / 0 | 2.1 | Yes |
| 20186 | 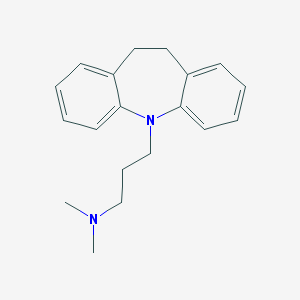 imipramine imipramine | C19H24N2 | 280.415 | 2 / 0 | 4.8 | Yes |
| 442463 | 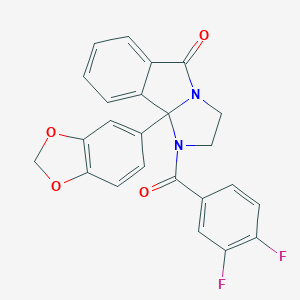 CHEMBL3353265 CHEMBL3353265 | C24H16F2N2O4 | 434.399 | 6 / 0 | 3.5 | Yes |
| 20542 | 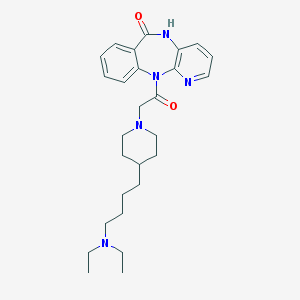 AQ-RA 741 AQ-RA 741 | C27H37N5O2 | 463.626 | 5 / 1 | 3.1 | Yes |
| 20974 | 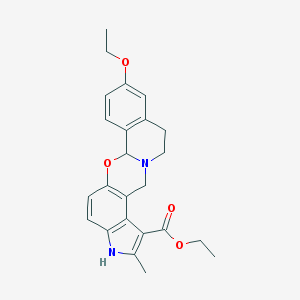 CHEMBL62247 CHEMBL62247 | C24H26N2O4 | 406.482 | 5 / 1 | 4.2 | Yes |
| 21068 | 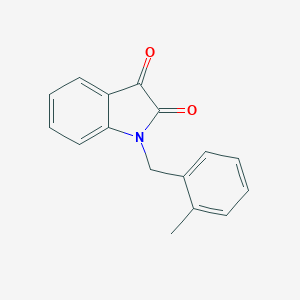 1-(2-methylbenzyl)-1H-indole-2,3-dione 1-(2-methylbenzyl)-1H-indole-2,3-dione | C16H13NO2 | 251.285 | 2 / 0 | 2.4 | Yes |
| 442508 | 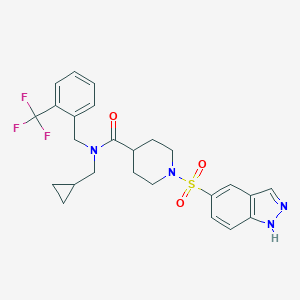 CHEMBL3329765 CHEMBL3329765 | C25H27F3N4O3S | 520.571 | 8 / 1 | 3.8 | No |
| 22067 | 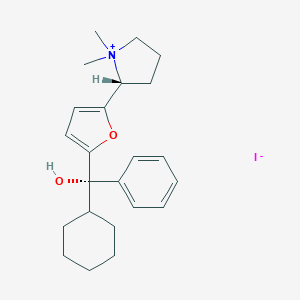 CHEMBL568775 CHEMBL568775 | C23H32INO2 | 481.418 | 3 / 1 | N/A | N/A |
| 22072 | 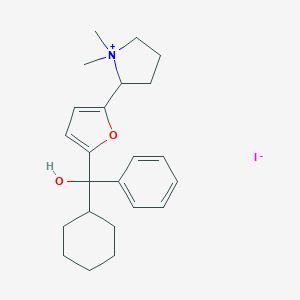 CHEMBL569760 CHEMBL569760 | C23H32INO2 | 481.418 | 3 / 1 | N/A | N/A |
| 22079 | 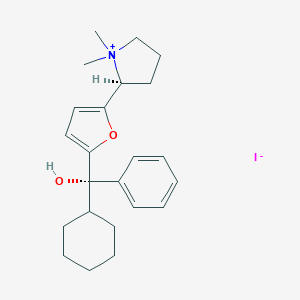 CHEMBL569089 CHEMBL569089 | C23H32INO2 | 481.418 | 3 / 1 | N/A | N/A |
| 22082 | 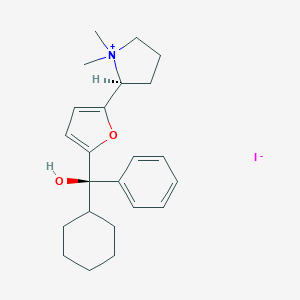 CHEMBL568870 CHEMBL568870 | C23H32INO2 | 481.418 | 3 / 1 | N/A | N/A |
| 22086 | 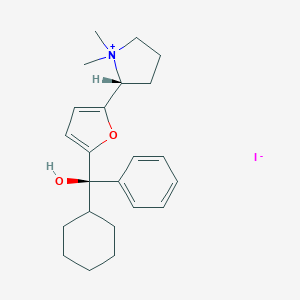 CHEMBL569307 CHEMBL569307 | C23H32INO2 | 481.418 | 3 / 1 | N/A | N/A |
| 522142 | 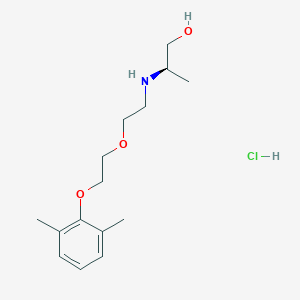 CHEMBL3775436 CHEMBL3775436 | C15H26ClNO3 | 303.827 | 4 / 3 | N/A | N/A |
| 522197 | 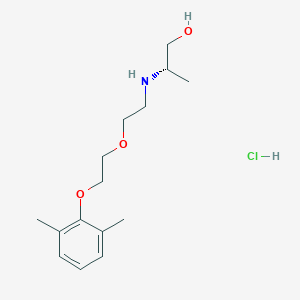 CHEMBL3775769 CHEMBL3775769 | C15H26ClNO3 | 303.827 | 4 / 3 | N/A | N/A |
| 22175 | 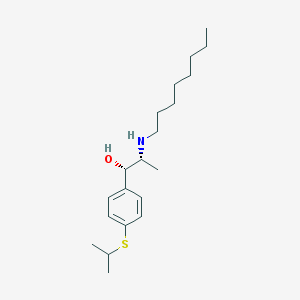 suloctidil suloctidil | C20H35NOS | 337.566 | 3 / 2 | 5.8 | No |
| 22244 | 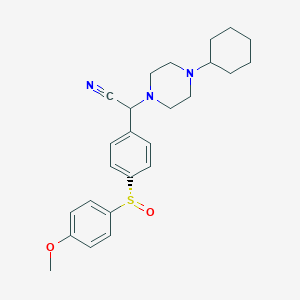 CHEMBL2111540 CHEMBL2111540 | C25H31N3O2S | 437.602 | 6 / 0 | 4.1 | Yes |
| 22297 | 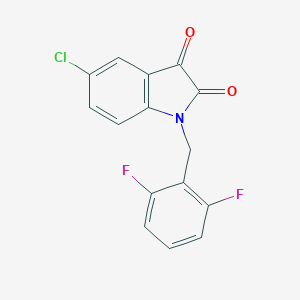 CHEMBL512236 CHEMBL512236 | C15H8ClF2NO2 | 307.681 | 4 / 0 | 3.2 | Yes |
| 22556 | 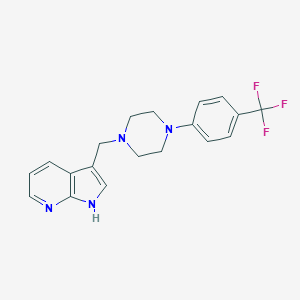 CHEMBL283036 CHEMBL283036 | C19H19F3N4 | 360.384 | 6 / 1 | 3.6 | Yes |
| 22872 | 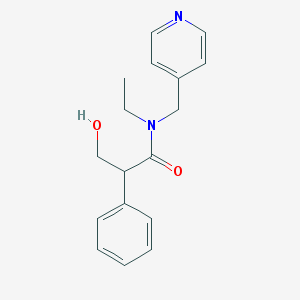 tropicamide tropicamide | C17H20N2O2 | 284.359 | 3 / 1 | 1.5 | Yes |
| 22907 | 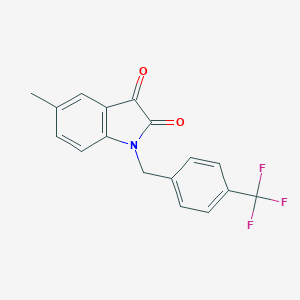 MLS002473340 MLS002473340 | C17H12F3NO2 | 319.283 | 5 / 0 | 3.6 | Yes |
| 23582 | 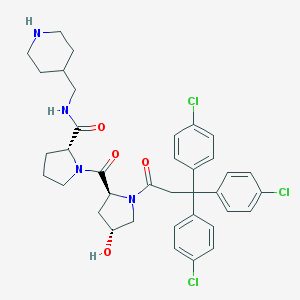 CHEMBL378872 CHEMBL378872 | C37H41Cl3N4O4 | 712.109 | 5 / 3 | 5.9 | No |
| 24596 | 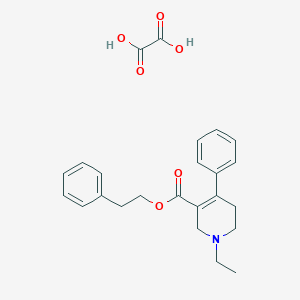 CHEMBL262198 CHEMBL262198 | C24H27NO6 | 425.481 | 7 / 2 | N/A | N/A |
| 24634 | 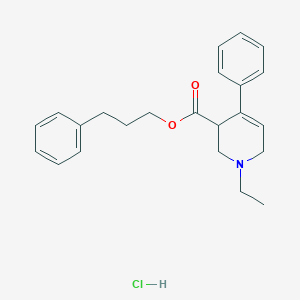 CHEMBL540316 CHEMBL540316 | C23H28ClNO2 | 385.932 | 3 / 1 | N/A | N/A |
| 25797 | 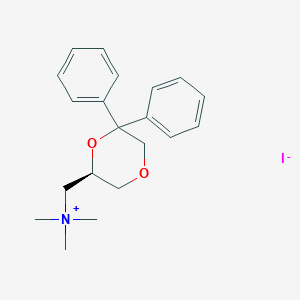 CHEMBL2042406 CHEMBL2042406 | C20H26INO2 | 439.337 | 3 / 0 | N/A | N/A |
| 25803 | 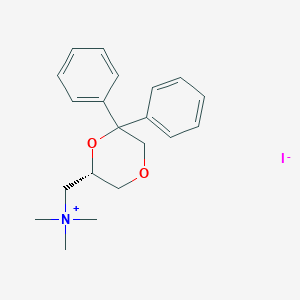 CHEMBL2042405 CHEMBL2042405 | C20H26INO2 | 439.337 | 3 / 0 | N/A | N/A |
| 25811 | 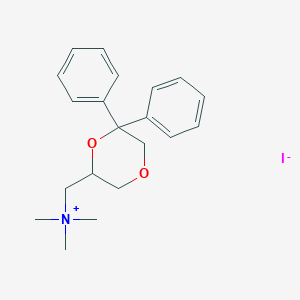 CHEMBL2042404 CHEMBL2042404 | C20H26INO2 | 439.337 | 3 / 0 | N/A | N/A |
| 25886 | 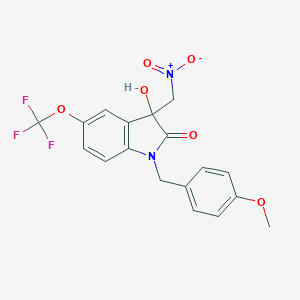 MLS004576082 MLS004576082 | C18H15F3N2O6 | 412.321 | 9 / 1 | 3.2 | Yes |
| 26381 | 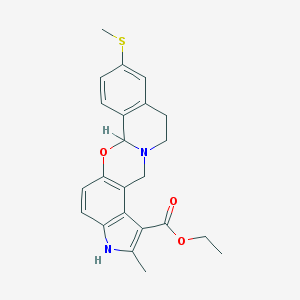 CHEMBL105122 CHEMBL105122 | C23H24N2O3S | 408.516 | 5 / 1 | 4.4 | Yes |
| 26440 | 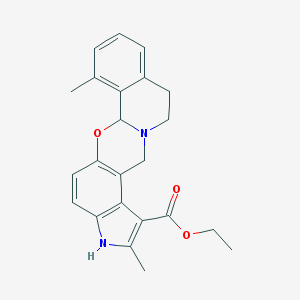 CHEMBL304036 CHEMBL304036 | C23H24N2O3 | 376.456 | 4 / 1 | 4.3 | Yes |
| 26491 | 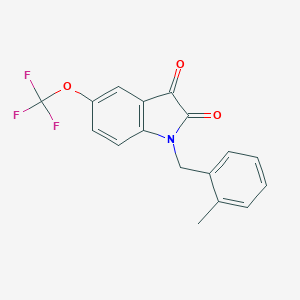 CHEMBL506043 CHEMBL506043 | C17H12F3NO3 | 335.282 | 6 / 0 | 3.9 | Yes |
| 26524 | 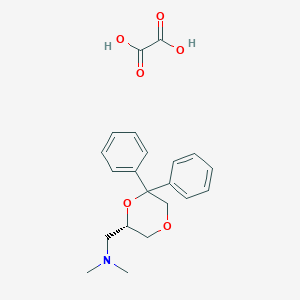 CHEMBL2042553 CHEMBL2042553 | C21H25NO6 | 387.432 | 7 / 2 | N/A | N/A |
| 26529 | 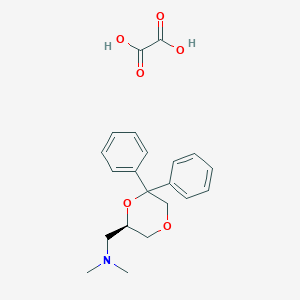 CHEMBL2042554 CHEMBL2042554 | C21H25NO6 | 387.432 | 7 / 2 | N/A | N/A |
| 26533 | 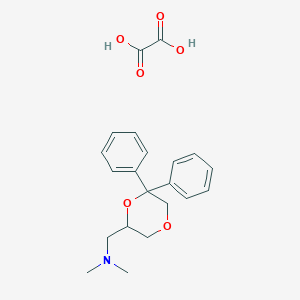 CHEMBL2042552 CHEMBL2042552 | C21H25NO6 | 387.432 | 7 / 2 | N/A | N/A |
| 26934 | 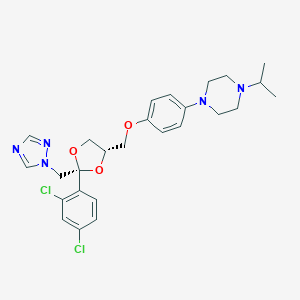 terconazole terconazole | C26H31Cl2N5O3 | 532.466 | 7 / 0 | 4.8 | No |
| 27218 | 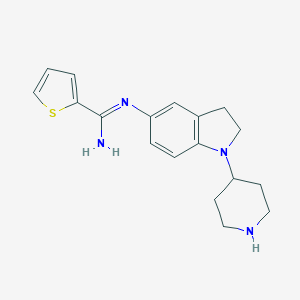 CHEMBL2203713 CHEMBL2203713 | C18H22N4S | 326.462 | 4 / 2 | 2.9 | Yes |
| 27490 | 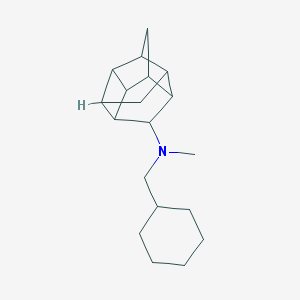 CHEMBL2432063 CHEMBL2432063 | C19H29N | 271.448 | 1 / 0 | 4.7 | Yes |
| 28359 | 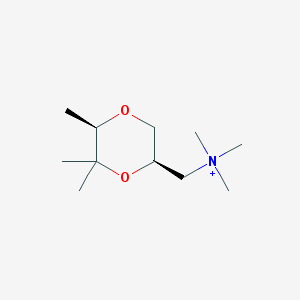 CHEMBL593685 CHEMBL593685 | C11H24NO2+ | 202.318 | 2 / 0 | 0.9 | Yes |
| 28361 | 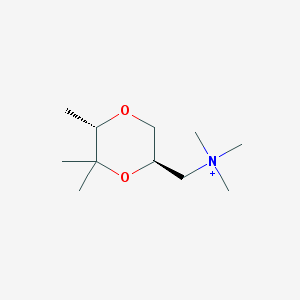 CHEMBL593620 CHEMBL593620 | C11H24NO2+ | 202.318 | 2 / 0 | 0.9 | Yes |
| 28541 | 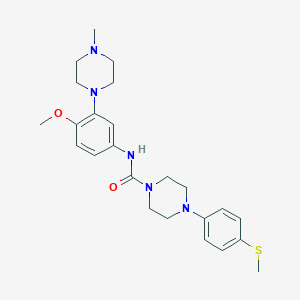 CHEMBL197796 CHEMBL197796 | C24H33N5O2S | 455.621 | 6 / 1 | 3.1 | Yes |
| 28619 | 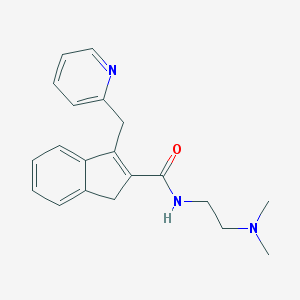 CHEMBL352779 CHEMBL352779 | C20H23N3O | 321.424 | 3 / 1 | 2.2 | Yes |
| 28943 | 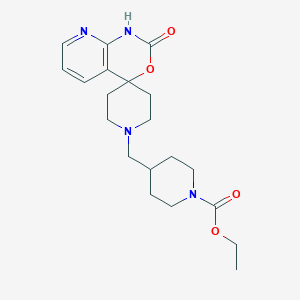 CHEMBL2407171 CHEMBL2407171 | C20H28N4O4 | 388.468 | 6 / 1 | 1.7 | Yes |
| 29118 | 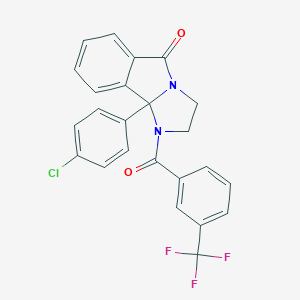 CHEMBL3105211 CHEMBL3105211 | C24H16ClF3N2O2 | 456.849 | 5 / 0 | 5.0 | Yes |
| 29604 | 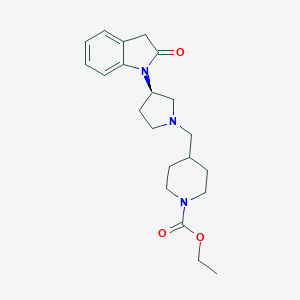 CHEMBL2336054 CHEMBL2336054 | C21H29N3O3 | 371.481 | 4 / 0 | 2.2 | Yes |
| 29610 | 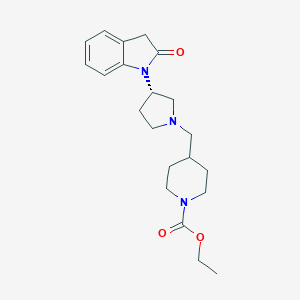 CHEMBL2336055 CHEMBL2336055 | C21H29N3O3 | 371.481 | 4 / 0 | 2.2 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218