

You can:
| Name | Melanocortin receptor 4 |
|---|---|
| Species | Mus musculus (Mouse) |
| Gene | Mc4r |
| Synonym | MC4 receptor MC4-R |
| Disease | N/A for non-human GPCRs |
| Length | 332 |
| Amino acid sequence | MNSTHHHGMYTSLHLWNRSSYGLHGNASESLGKGHPDGGCYEQLFVSPEVFVTLGVISLLENILVIVAIAKNKNLHSPMYFFICSLAVADMLVSVSNGSETIVITLLNSTDTDAQSFTVNIDNVIDSVICSSLLASICSLLSIAVDRYFTIFYALQYHNIMTVRRVGIIISCIWAACTVSGVLFIIYSDSSAVIICLISMFFTMLVLMASLYVHMFLMARLHIKRIAVLPGTGTIRQGTNMKGAITLTILIGVFVVCWAPFFLHLLFYISCPQNPYCVCFMSHFNLYLILIMCNAVIDPLIYALRSQELRKTFKEIICFYPLGGICELSSRY |
| UniProt | P56450 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL3719 |
| IUPHAR | N/A |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 463042 | 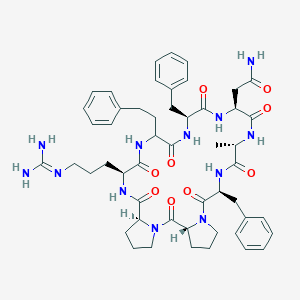 CHEMBL3577985 CHEMBL3577985 | C51H66N12O9 | 991.164 | 10 / 9 | 1.0 | No |
| 5972 | 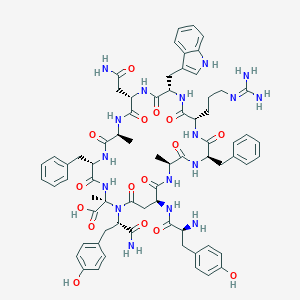 CHEMBL2369967 CHEMBL2369967 | C70H85N17O16 | 1420.55 | 18 / 18 | -2.5 | No |
| 5976 | 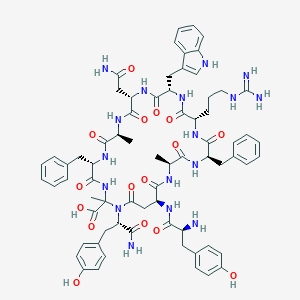 CHEMBL2096721 CHEMBL2096721 | C70H85N17O16 | 1420.55 | 18 / 19 | -2.2 | No |
| 8133 | 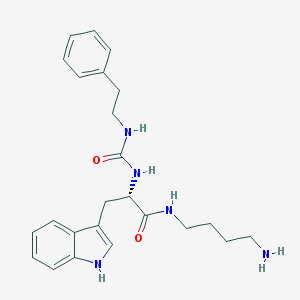 CHEMBL431523 CHEMBL431523 | C24H31N5O2 | 421.545 | 3 / 5 | 1.7 | Yes |
| 10924 | 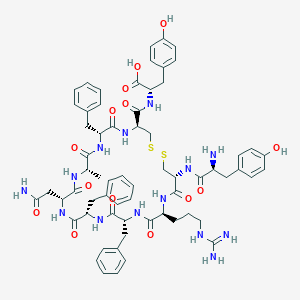 BDBM50106532 BDBM50106532 | C64H78N14O14S2 | 1331.53 | 18 / 17 | -0.7 | No |
| 11090 | 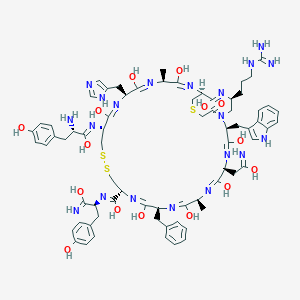 CID 102096776 CID 102096776 | C71H90N20O15S3 | 1559.81 | 33 / 22 | 4.0 | No |
| 521795 | 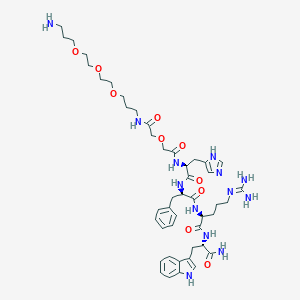 CHEMBL3799563 CHEMBL3799563 | C46H67N13O10 | 962.123 | 13 / 11 | -1.4 | No |
| 12238 | 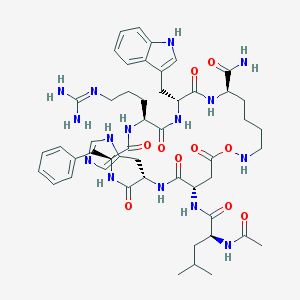 CHEMBL216295 CHEMBL216295 | C50H69N15O10 | 1040.2 | 13 / 13 | 0.3 | No |
| 12396 | 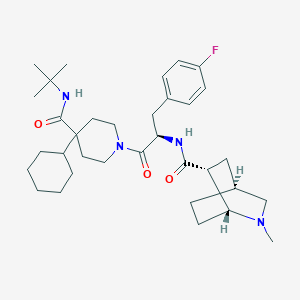 RY-764 RY-764 | C34H51FN4O3 | 582.805 | 5 / 2 | 5.4 | No |
| 15600 | 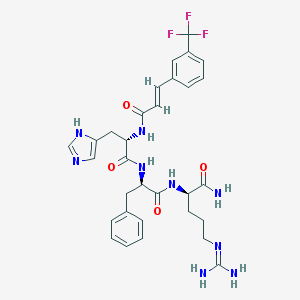 CHEMBL569696 CHEMBL569696 | C31H36F3N9O4 | 655.683 | 9 / 7 | 1.5 | No |
| 16256 | 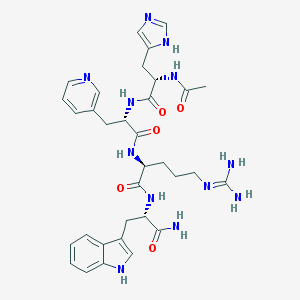 CHEMBL104052 CHEMBL104052 | C33H42N12O5 | 686.778 | 8 / 9 | -1.1 | No |
| 18262 | 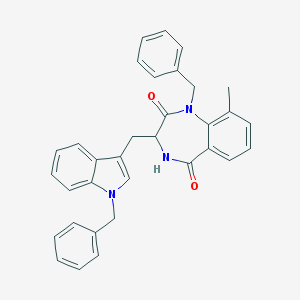 CHEMBL410435 CHEMBL410435 | C33H29N3O2 | 499.614 | 2 / 1 | 5.9 | No |
| 522049 | 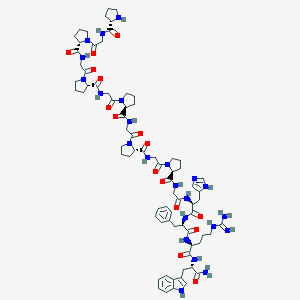 CHEMBL3799094 CHEMBL3799094 | C74H101N23O16 | 1568.77 | 19 / 17 | -3.2 | No |
| 20120 | 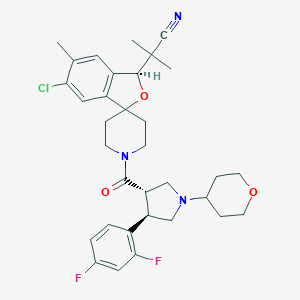 CHEMBL1209788 CHEMBL1209788 | C33H38ClF2N3O3 | 598.132 | 7 / 0 | 4.8 | No |
| 23001 | 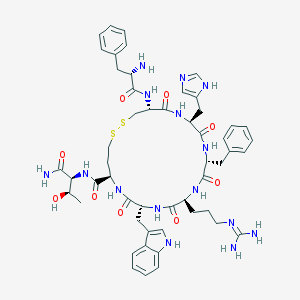 CHEMBL406891 CHEMBL406891 | C52H67N15O9S2 | 1110.32 | 14 / 14 | -0.2 | No |
| 26906 | 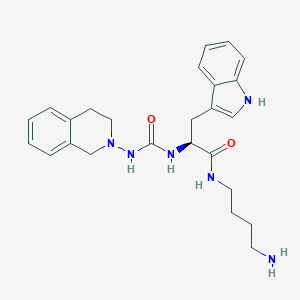 CHEMBL62228 CHEMBL62228 | C25H32N6O2 | 448.571 | 4 / 5 | 2.4 | Yes |
| 28883 | 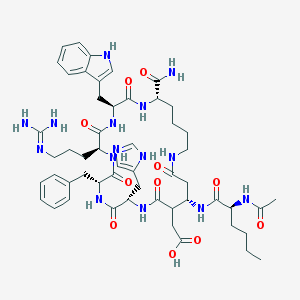 CHEMBL224277 CHEMBL224277 | C53H73N15O11 | 1096.26 | 13 / 14 | -0.5 | No |
| 31251 | 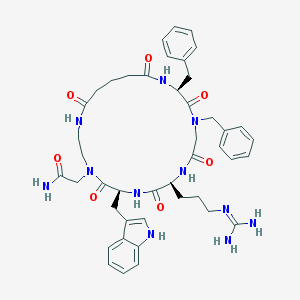 CHEMBL602654 CHEMBL602654 | C44H55N11O7 | 849.994 | 8 / 8 | 0.4 | No |
| 467202 | 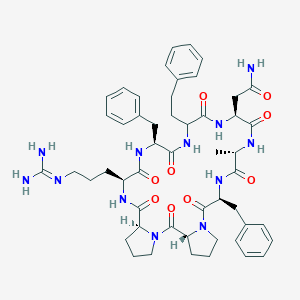 CHEMBL3577991 CHEMBL3577991 | C51H66N12O9 | 991.164 | 10 / 9 | 1.0 | No |
| 35309 | 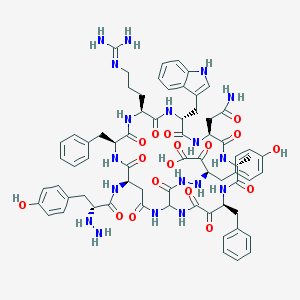 CID 44310242 CID 44310242 | C68H80N18O17 | 1421.5 | 21 / 20 | -2.4 | No |
| 35313 | 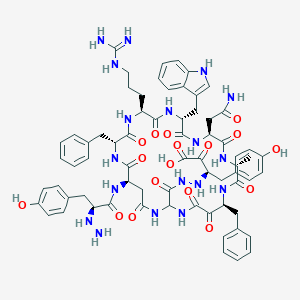 BDBM50144874 BDBM50144874 | C68H80N18O17 | 1421.5 | 21 / 21 | -2.0 | No |
| 35367 | 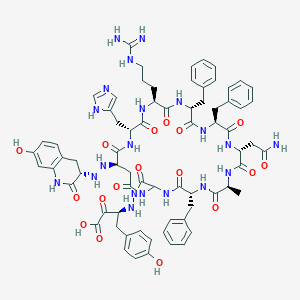 CHEMBL414900 CHEMBL414900 | C71H84N20O17 | 1489.58 | 22 / 22 | -2.9 | No |
| 35370 | 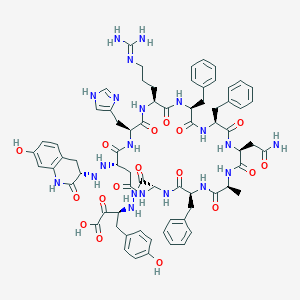 CHEMBL2371699 CHEMBL2371699 | C71H84N20O17 | 1489.58 | 22 / 21 | -3.3 | No |
| 36345 | 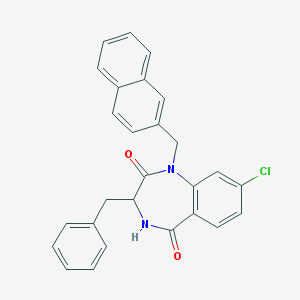 CHEMBL270691 CHEMBL270691 | C27H21ClN2O2 | 440.927 | 2 / 1 | 5.7 | No |
| 467502 | 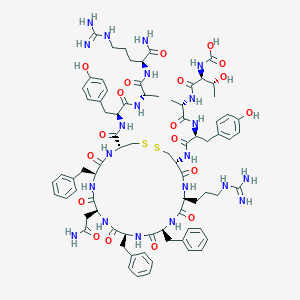 CHEMBL3578001 CHEMBL3578001 | C78H103N21O19S2 | 1702.93 | 23 / 25 | -0.9 | No |
| 522718 | 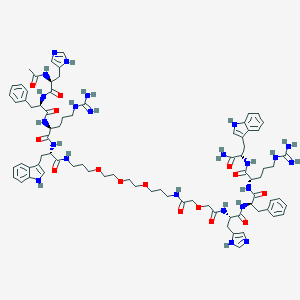 CHEMBL3798421 CHEMBL3798421 | C80H107N23O15 | 1630.88 | 19 / 21 | 0.4 | No |
| 39994 | 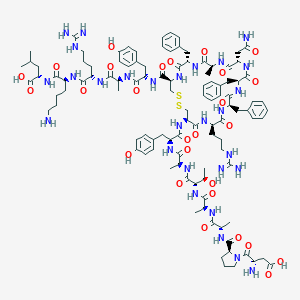 CHEMBL2304248 CHEMBL2304248 | C107H152N28O27S2 | 2326.68 | 33 / 32 | -6.4 | No |
| 40000 | 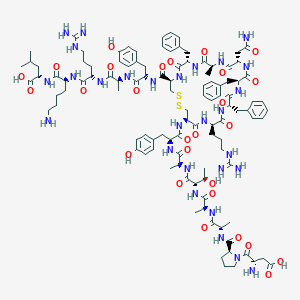 CHEMBL2304250 CHEMBL2304250 | C107H152N28O27S2 | 2326.68 | 33 / 32 | -6.4 | No |
| 40004 | 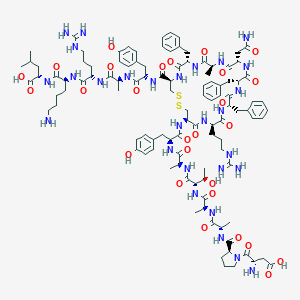 CHEMBL2304246 CHEMBL2304246 | C107H152N28O27S2 | 2326.68 | 33 / 32 | -6.4 | No |
| 40008 | 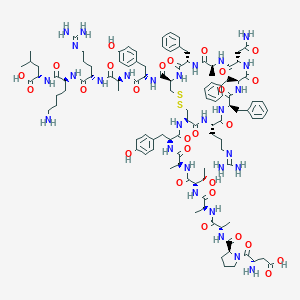 BDBM50422954 BDBM50422954 | C107H152N28O27S2 | 2326.68 | 33 / 30 | -7.1 | No |
| 40013 | 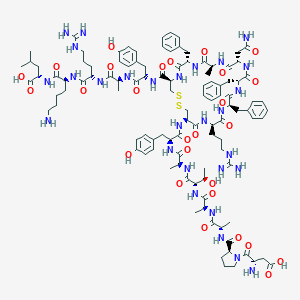 CHEMBL2369131 CHEMBL2369131 | C107H152N28O27S2 | 2326.68 | 33 / 32 | -6.4 | No |
| 40014 | 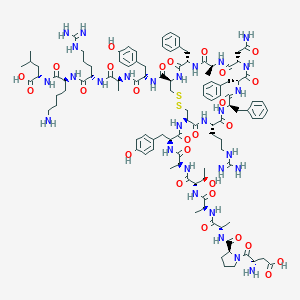 CHEMBL2304249 CHEMBL2304249 | C107H152N28O27S2 | 2326.68 | 33 / 32 | -6.4 | No |
| 443273 | 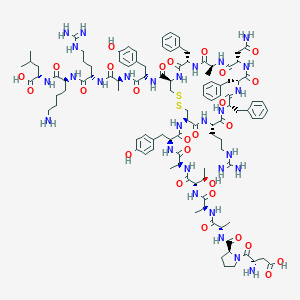 CHEMBL3350327 CHEMBL3350327 | C107H152N28O27S2 | 2326.68 | 33 / 32 | -6.4 | No |
| 443290 | 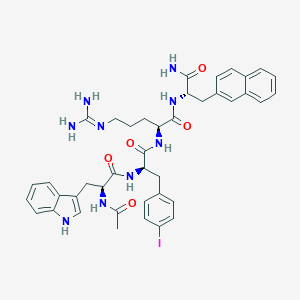 CHEMBL3422432 CHEMBL3422432 | C41H46IN9O5 | 871.781 | 6 / 8 | 3.7 | No |
| 443293 | 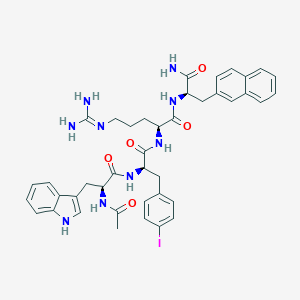 CHEMBL3422433 CHEMBL3422433 | C41H46IN9O5 | 871.781 | 6 / 8 | 3.7 | No |
| 41145 | 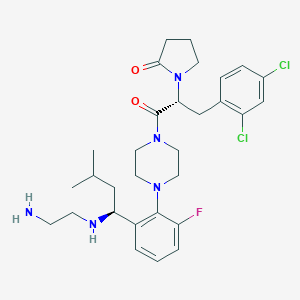 CHEMBL393576 CHEMBL393576 | C30H40Cl2FN5O2 | 592.581 | 6 / 2 | 4.4 | No |
| 44413 | 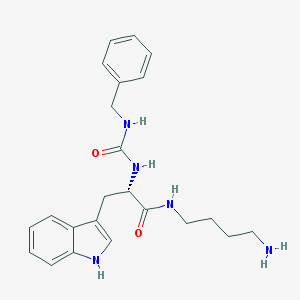 CHEMBL305559 CHEMBL305559 | C23H29N5O2 | 407.518 | 3 / 5 | 1.9 | Yes |
| 44417 | 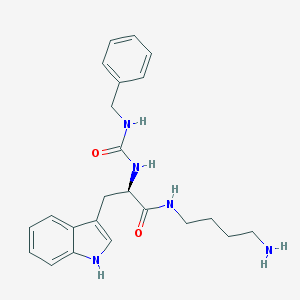 CHEMBL64954 CHEMBL64954 | C23H29N5O2 | 407.518 | 3 / 5 | 1.9 | Yes |
| 468542 | 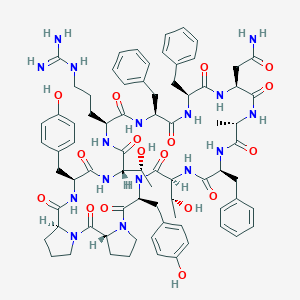 CHEMBL3577980 CHEMBL3577980 | C76H96N16O17 | 1505.7 | 18 / 18 | 2.3 | No |
| 46073 | 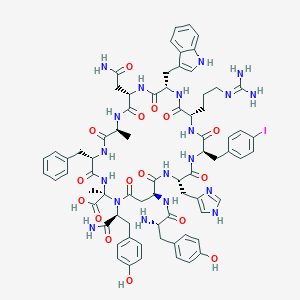 CHEMBL2369973 CHEMBL2369973 | C73H86IN19O16 | 1612.51 | 19 / 19 | -2.1 | No |
| 46895 | 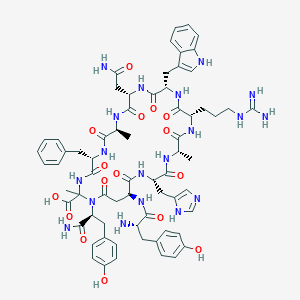 CHEMBL2096722 CHEMBL2096722 | C67H83N19O16 | 1410.52 | 19 / 20 | -4.0 | No |
| 46898 | 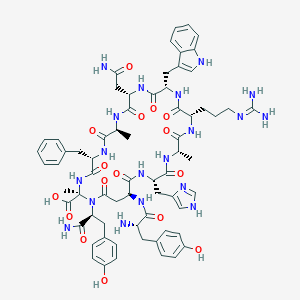 CHEMBL2369968 CHEMBL2369968 | C67H83N19O16 | 1410.52 | 19 / 19 | -4.4 | No |
| 49620 | 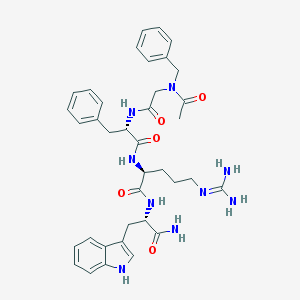 CHEMBL138771 CHEMBL138771 | C37H45N9O5 | 695.825 | 6 / 7 | 1.2 | No |
| 469160 | 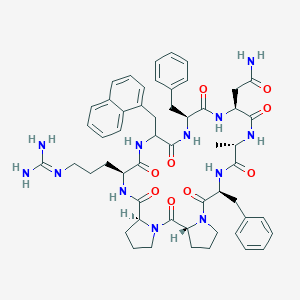 CHEMBL3577986 CHEMBL3577986 | C54H66N12O9 | 1027.2 | 10 / 9 | 1.9 | No |
| 523031 | 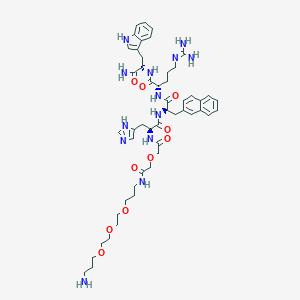 CHEMBL3798912 CHEMBL3798912 | C50H69N13O10 | 1012.18 | 13 / 11 | -0.2 | No |
| 51829 | 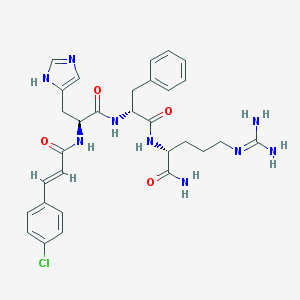 CHEMBL568577 CHEMBL568577 | C30H36ClN9O4 | 622.127 | 6 / 7 | 1.2 | No |
| 52138 | 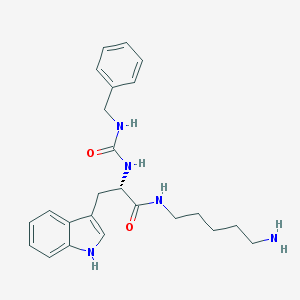 CHEMBL305391 CHEMBL305391 | C24H31N5O2 | 421.545 | 3 / 5 | 2.3 | Yes |
| 52480 | 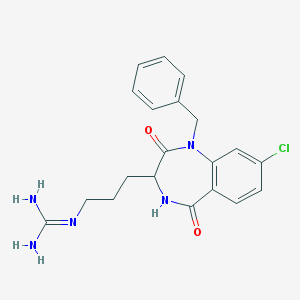 CHEMBL272439 CHEMBL272439 | C20H22ClN5O2 | 399.879 | 3 / 3 | 1.7 | Yes |
| 53549 | 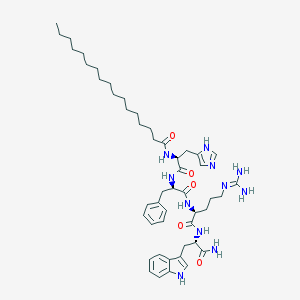 CHEMBL191304 CHEMBL191304 | C49H73N11O5 | 896.195 | 7 / 9 | 7.9 | No |
| 469597 | 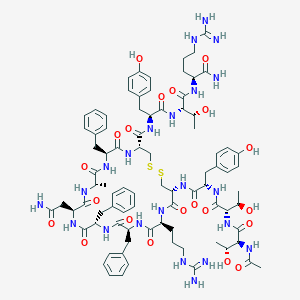 CHEMBL3577977 CHEMBL3577977 | C84H114N22O21S2 | 1832.09 | 25 / 27 | -2.5 | No |
| 54367 | 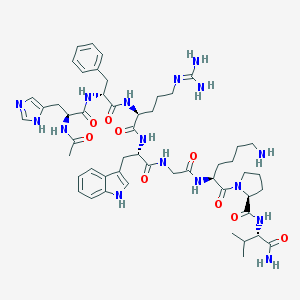 CHEMBL405335 CHEMBL405335 | C52H74N16O9 | 1067.27 | 12 / 13 | -0.4 | No |
| 56084 | 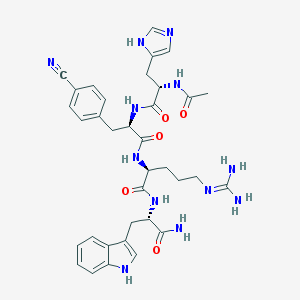 CHEMBL448536 CHEMBL448536 | C35H42N12O5 | 710.8 | 8 / 9 | -0.3 | No |
| 58385 | 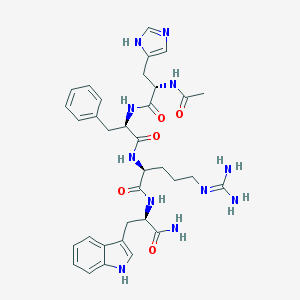 CHEMBL433296 CHEMBL433296 | C34H43N11O5 | 685.79 | 7 / 9 | -0.3 | No |
| 58387 | 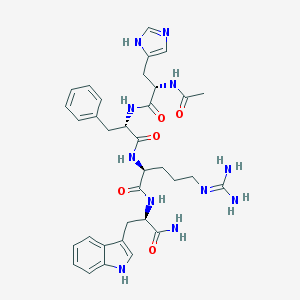 Ac-His-Phe-Arg-DTrp-NH2 Ac-His-Phe-Arg-DTrp-NH2 | C34H43N11O5 | 685.79 | 7 / 9 | -0.3 | No |
| 58399 | 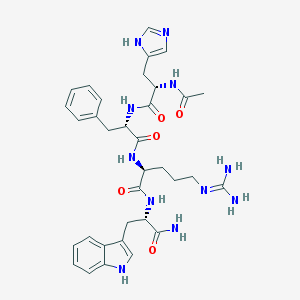 Ac-His-Phe-Arg-Trp-NH2 Ac-His-Phe-Arg-Trp-NH2 | C34H43N11O5 | 685.79 | 7 / 9 | -0.3 | No |
| 58404 | 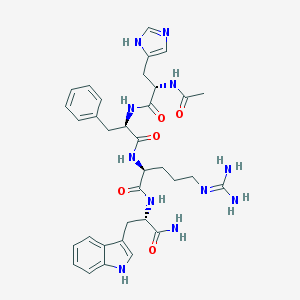 CHEMBL264190 CHEMBL264190 | C34H43N11O5 | 685.79 | 7 / 9 | -0.3 | No |
| 58411 | 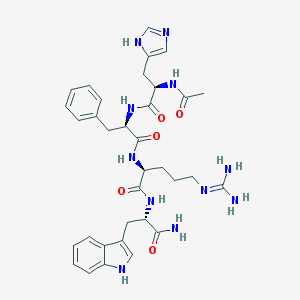 CHEMBL94391 CHEMBL94391 | C34H43N11O5 | 685.79 | 7 / 9 | -0.3 | No |
| 58416 | 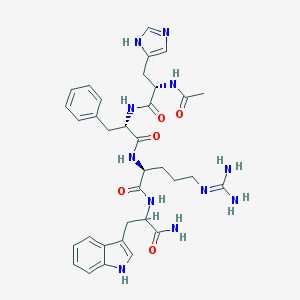 CHEMBL152612 CHEMBL152612 | C34H43N11O5 | 685.79 | 7 / 9 | -0.3 | No |
| 58849 | 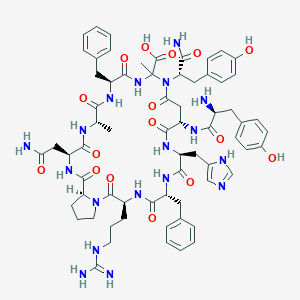 CHEMBL2096716 CHEMBL2096716 | C67H84N18O16 | 1397.52 | 19 / 18 | -3.8 | No |
| 58852 | 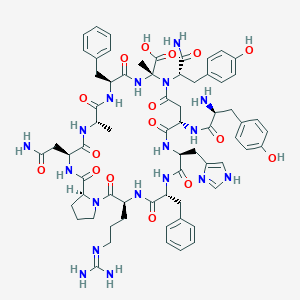 CHEMBL2369962 CHEMBL2369962 | C67H84N18O16 | 1397.52 | 19 / 17 | -4.2 | No |
| 444090 | 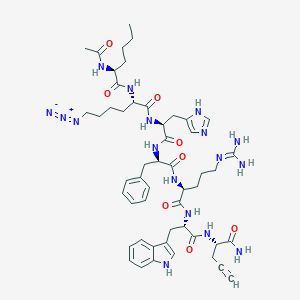 CHEMBL3358537 CHEMBL3358537 | C51H69N17O8 | 1048.22 | 12 / 12 | 2.5 | No |
| 64601 | 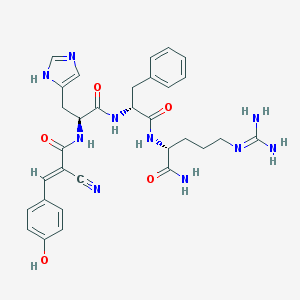 CHEMBL565900 CHEMBL565900 | C31H36N10O5 | 628.694 | 8 / 8 | 0.2 | No |
| 64655 | 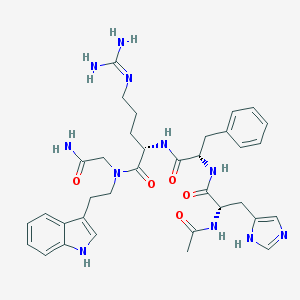 CHEMBL138212 CHEMBL138212 | C35H45N11O5 | 699.817 | 7 / 8 | 0.1 | No |
| 470883 | 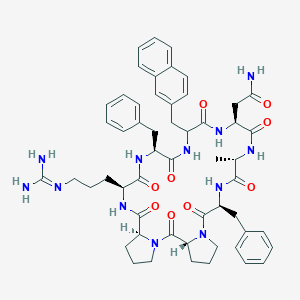 CHEMBL3577994 CHEMBL3577994 | C54H66N12O9 | 1027.2 | 10 / 9 | 1.9 | No |
| 470927 | 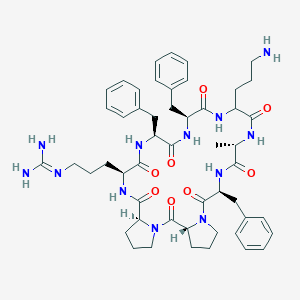 CHEMBL3577998 CHEMBL3577998 | C51H68N12O8 | 977.181 | 10 / 9 | 1.6 | No |
| 65152 | 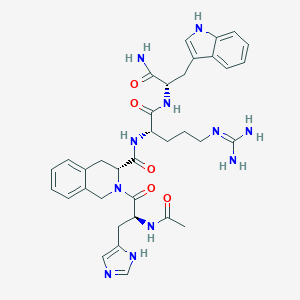 CHEMBL607103 CHEMBL607103 | C35H43N11O5 | 697.801 | 7 / 8 | -0.2 | No |
| 65157 | 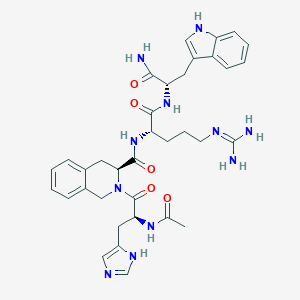 CHEMBL607104 CHEMBL607104 | C35H43N11O5 | 697.801 | 7 / 8 | -0.2 | No |
| 65162 | 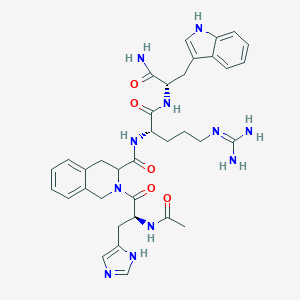 BDBM50115367 BDBM50115367 | C35H43N11O5 | 697.801 | 7 / 8 | -0.2 | No |
| 65902 | 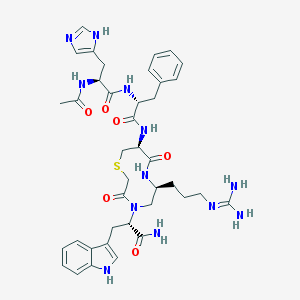 CHEMBL1688108 CHEMBL1688108 | C39H50N12O6S | 814.967 | 9 / 9 | 0.0 | No |
| 66079 | 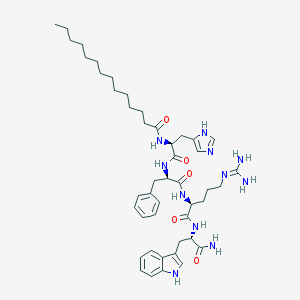 CHEMBL370261 CHEMBL370261 | C46H67N11O5 | 854.114 | 7 / 9 | 6.2 | No |
| 66255 | 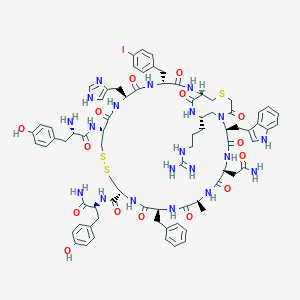 CID 71606706 CID 71606706 | C77H93IN20O15S3 | 1761.8 | 21 / 20 | 1.3 | No |
| 70082 | 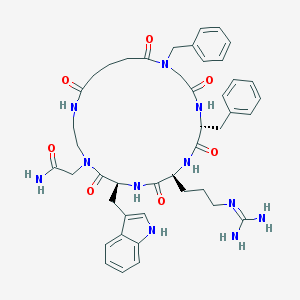 CHEMBL589468 CHEMBL589468 | C44H55N11O7 | 849.994 | 8 / 8 | 0.4 | No |
| 70474 | 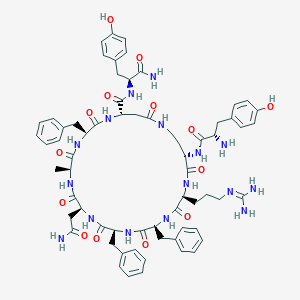 CHEMBL2370696 CHEMBL2370696 | C66H82N16O14 | 1323.48 | 16 / 17 | -0.5 | No |
| 70830 | 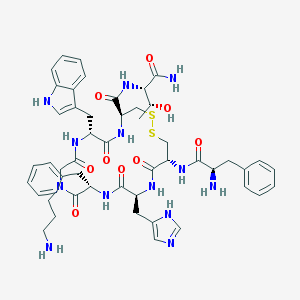 CHEMBL415200 CHEMBL415200 | C50H63N13O9S2 | 1054.26 | 14 / 12 | -0.2 | No |
| 444618 | 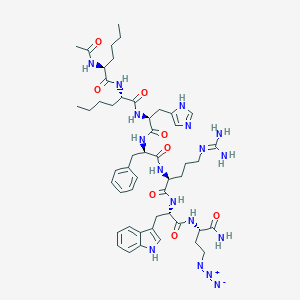 CHEMBL3352878 CHEMBL3352878 | C50H71N17O8 | 1038.23 | 12 / 12 | 3.0 | No |
| 77644 | 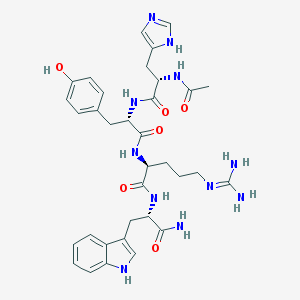 CHEMBL104880 CHEMBL104880 | C34H43N11O6 | 701.789 | 8 / 10 | -0.4 | No |
| 77647 | 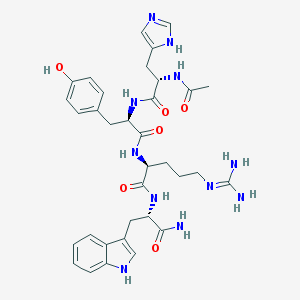 CHEMBL105113 CHEMBL105113 | C34H43N11O6 | 701.789 | 8 / 10 | -0.4 | No |
| 472624 | 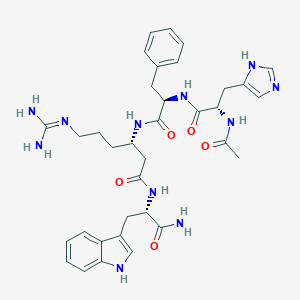 CHEMBL3582445 CHEMBL3582445 | C35H45N11O5 | 699.817 | 7 / 9 | -0.4 | No |
| 80091 | 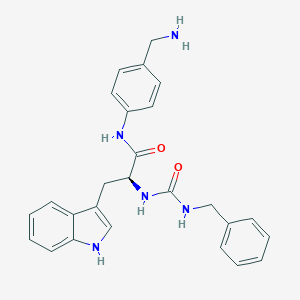 CHEMBL2112213 CHEMBL2112213 | C26H27N5O2 | 441.535 | 3 / 5 | 2.6 | Yes |
| 472831 | 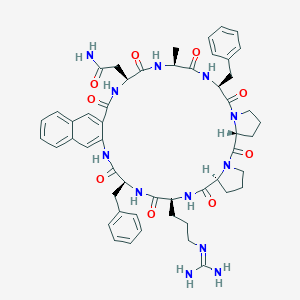 CHEMBL3577992 CHEMBL3577992 | C52H62N12O9 | 999.143 | 10 / 9 | 1.4 | No |
| 80548 | 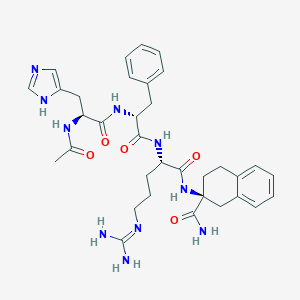 CHEMBL2371147 CHEMBL2371147 | C34H44N10O5 | 672.791 | 7 / 8 | 0.1 | No |
| 80552 | 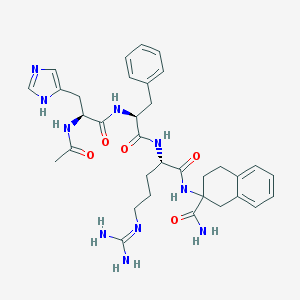 Ac-His-D-Phe-Arg-Atc-NH2 Ac-His-D-Phe-Arg-Atc-NH2 | C34H44N10O5 | 672.791 | 7 / 8 | 0.1 | No |
| 81409 | 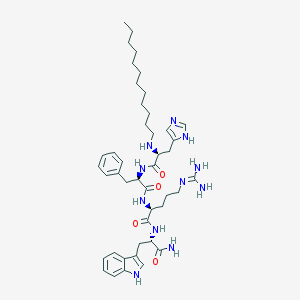 CHEMBL195468 CHEMBL195468 | C44H65N11O4 | 812.077 | 7 / 9 | 5.8 | No |
| 82015 | 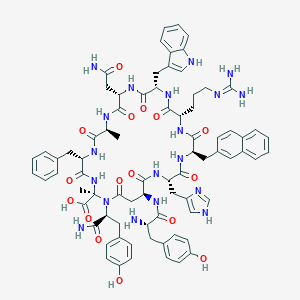 CHEMBL2369974 CHEMBL2369974 | C77H89N19O16 | 1536.68 | 19 / 19 | -1.5 | No |
| 473108 | 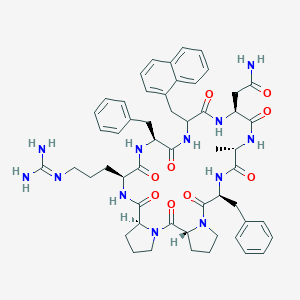 CHEMBL3577993 CHEMBL3577993 | C54H66N12O9 | 1027.2 | 10 / 9 | 1.9 | No |
| 85416 | 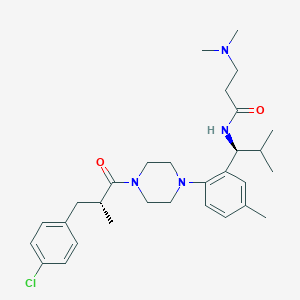 CHEMBL427860 CHEMBL427860 | C30H43ClN4O2 | 527.15 | 4 / 1 | 5.2 | No |
| 86754 | 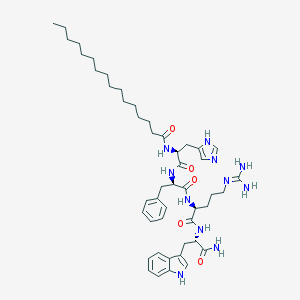 CHEMBL265236 CHEMBL265236 | C48H71N11O5 | 882.168 | 7 / 9 | 7.3 | No |
| 86759 | 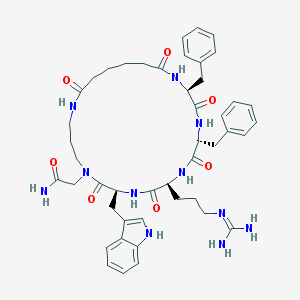 CHEMBL271586 CHEMBL271586 | C46H59N11O7 | 878.048 | 8 / 9 | 1.4 | No |
| 92926 | 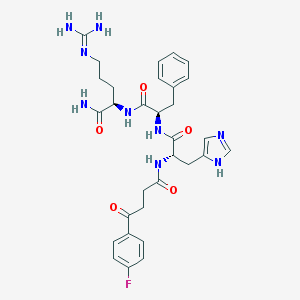 CHEMBL568836 CHEMBL568836 | C31H38FN9O5 | 635.701 | 8 / 7 | -0.1 | No |
| 93856 | 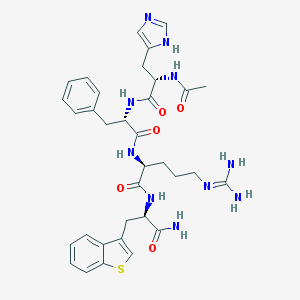 CHEMBL151906 CHEMBL151906 | C34H42N10O5S | 702.835 | 8 / 8 | 0.9 | No |
| 474518 | 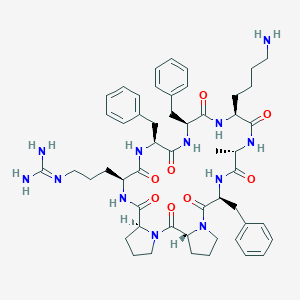 CHEMBL3577999 CHEMBL3577999 | C52H70N12O8 | 991.208 | 10 / 9 | 1.9 | No |
| 96205 | 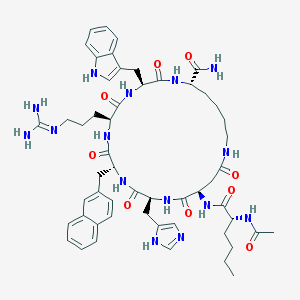 CHEMBL2096742 CHEMBL2096742 | C54H71N15O9 | 1074.26 | 11 / 13 | 1.3 | No |
| 445498 | 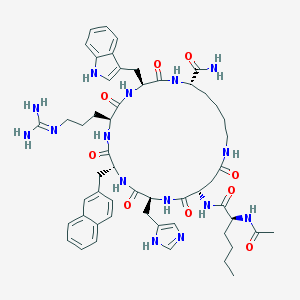 CHEMBL3422426 CHEMBL3422426 | C54H71N15O9 | 1074.26 | 11 / 13 | 1.3 | No |
| 524344 | 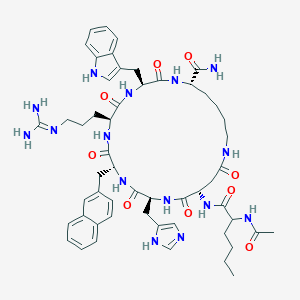 CHEMBL3735607 CHEMBL3735607 | C54H71N15O9 | 1074.26 | 11 / 13 | 1.3 | No |
| 445640 | 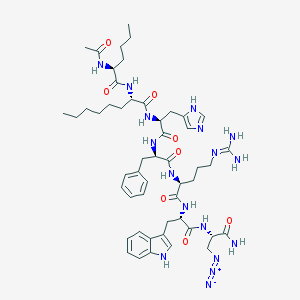 CHEMBL3358536 CHEMBL3358536 | C51H73N17O8 | 1052.26 | 12 / 12 | 3.7 | No |
| 101877 | 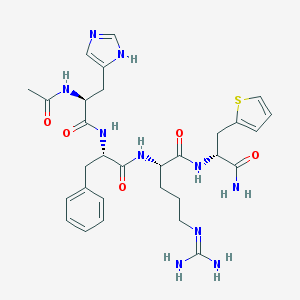 CHEMBL349795 CHEMBL349795 | C30H40N10O5S | 652.775 | 8 / 8 | -0.4 | No |
| 106375 | 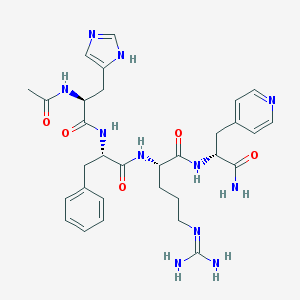 CHEMBL433522 CHEMBL433522 | C31H41N11O5 | 647.741 | 8 / 8 | -1.2 | No |
| 106752 | 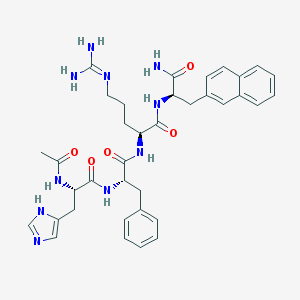 CHEMBL357558 CHEMBL357558 | C36H44N10O5 | 696.813 | 7 / 8 | 1.1 | No |
| 106758 | 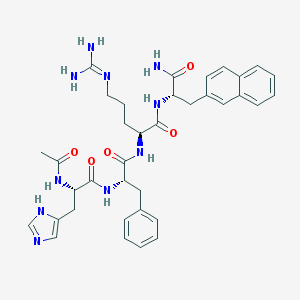 CHEMBL347216 CHEMBL347216 | C36H44N10O5 | 696.813 | 7 / 8 | 1.1 | No |
| 107139 | 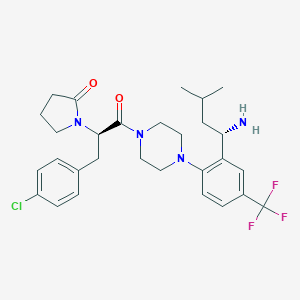 CHEMBL439560 CHEMBL439560 | C29H36ClF3N4O2 | 565.078 | 7 / 1 | 5.0 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218