

You can:
| Name | Mu-type opioid receptor |
|---|---|
| Species | Rattus norvegicus (Rat) |
| Gene | Oprm1 |
| Synonym | Opioid receptor B opioid receptor OP3 mu receptor MOR-1 [ Show all ] |
| Disease | N/A for non-human GPCRs |
| Length | 398 |
| Amino acid sequence | MDSSTGPGNTSDCSDPLAQASCSPAPGSWLNLSHVDGNQSDPCGLNRTGLGGNDSLCPQTGSPSMVTAITIMALYSIVCVVGLFGNFLVMYVIVRYTKMKTATNIYIFNLALADALATSTLPFQSVNYLMGTWPFGTILCKIVISIDYYNMFTSIFTLCTMSVDRYIAVCHPVKALDFRTPRNAKIVNVCNWILSSAIGLPVMFMATTKYRQGSIDCTLTFSHPTWYWENLLKICVFIFAFIMPVLIITVCYGLMILRLKSVRMLSGSKEKDRNLRRITRMVLVVVAVFIVCWTPIHIYVIIKALITIPETTFQTVSWHFCIALGYTNSCLNPVLYAFLDENFKRCFREFCIPTSSTIEQQNSTRVRQNTREHPSTANTVDRTNHQLENLEAETAPLP |
| UniProt | P33535 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL270 |
| IUPHAR | 319 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 270 | 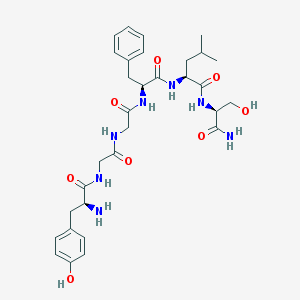 CHEMBL132776 CHEMBL132776 | C31H43N7O8 | 641.726 | 9 / 9 | -0.4 | No |
| 574 | 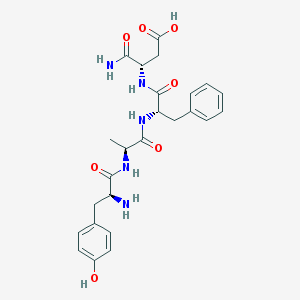 CHEMBL44172 CHEMBL44172 | C25H31N5O7 | 513.551 | 8 / 7 | -3.4 | No |
| 589 | 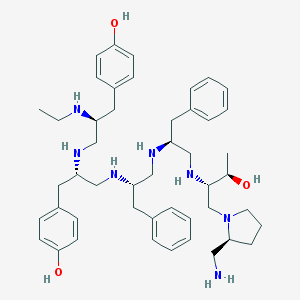 CHEMBL213366 CHEMBL213366 | C47H69N7O3 | 780.115 | 10 / 9 | 5.3 | No |
| 593 | 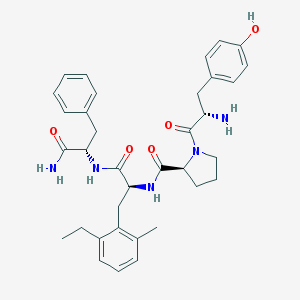 Tyr-Pro-Emp-Phe-NH2 Tyr-Pro-Emp-Phe-NH2 | C35H43N5O5 | 613.759 | 6 / 5 | 3.8 | No |
| 627 | 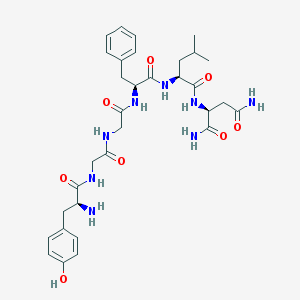 CHEMBL134023 CHEMBL134023 | C32H44N8O8 | 668.752 | 9 / 9 | -0.8 | No |
| 694 | 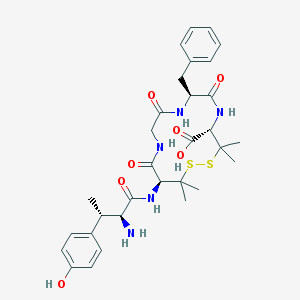 CHEMBL76086 CHEMBL76086 | C31H41N5O7S2 | 659.817 | 10 / 7 | -0.3 | No |
| 697 | 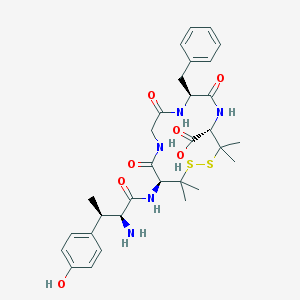 CHEMBL310681 CHEMBL310681 | C31H41N5O7S2 | 659.817 | 10 / 7 | -0.3 | No |
| 701 | 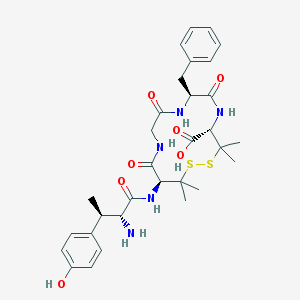 CHEMBL80626 CHEMBL80626 | C31H41N5O7S2 | 659.817 | 10 / 7 | -0.3 | No |
| 706 | 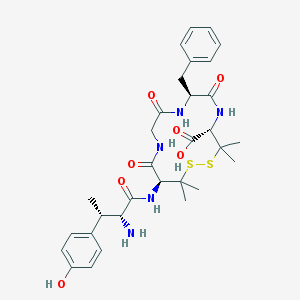 CHEMBL307614 CHEMBL307614 | C31H41N5O7S2 | 659.817 | 10 / 7 | -0.3 | No |
| 463063 | 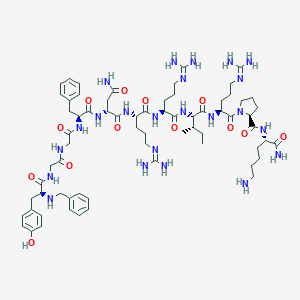 CHEMBL3634916 CHEMBL3634916 | C68H105N23O13 | 1452.74 | 18 / 20 | -3.6 | No |
| 1229 | 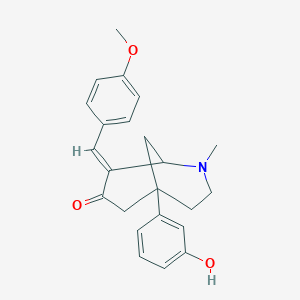 CHEMBL143800 CHEMBL143800 | C23H25NO3 | 363.457 | 4 / 1 | 3.6 | Yes |
| 1438 | 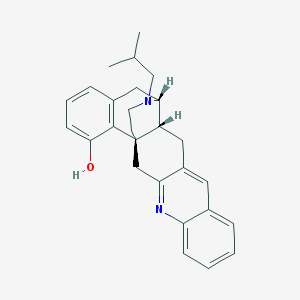 CHEMBL1929362 CHEMBL1929362 | C27H30N2O | 398.55 | 3 / 1 | 5.8 | No |
| 1511 | 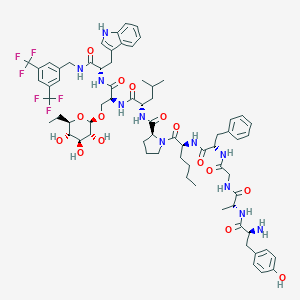 CHEMBL556537 CHEMBL556537 | C70H89F6N11O15 | 1438.53 | 22 / 14 | 5.8 | No |
| 1530 | 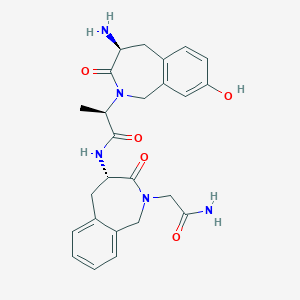 SB-0304 SB-0304 | C25H29N5O5 | 479.537 | 6 / 4 | -0.2 | Yes |
| 1767 | 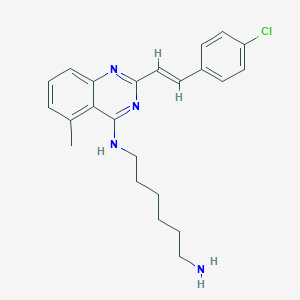 CHEMBL1186493 CHEMBL1186493 | C23H27ClN4 | 394.947 | 4 / 2 | 5.6 | No |
| 1842 | 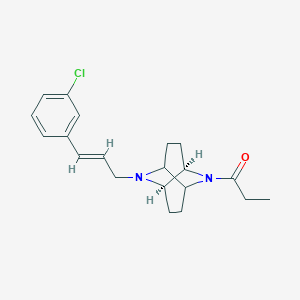 CHEMBL2310837 CHEMBL2310837 | C20H25ClN2O | 344.883 | 2 / 0 | 4.0 | Yes |
| 2053 | 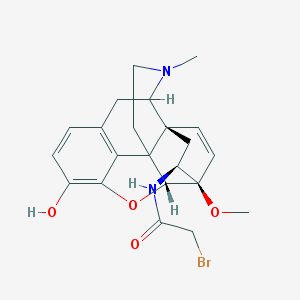 BDBM50027090 BDBM50027090 | C22H25BrN2O4 | 461.356 | 5 / 2 | 2.2 | Yes |
| 2055 | 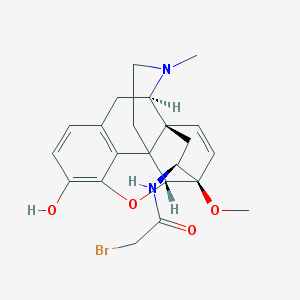 CHEMBL608248 CHEMBL608248 | C22H25BrN2O4 | 461.356 | 5 / 2 | 2.2 | Yes |
| 441748 | 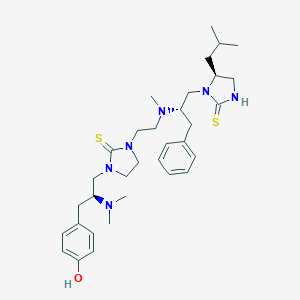 CHEMBL3326662 CHEMBL3326662 | C33H50N6OS2 | 610.924 | 5 / 2 | 5.3 | No |
| 521514 | 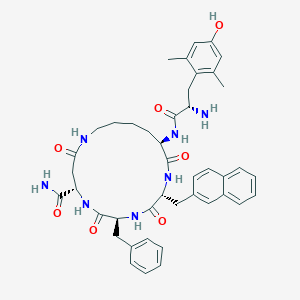 CHEMBL3759981 CHEMBL3759981 | C43H51N7O7 | 777.923 | 8 / 8 | 3.4 | No |
| 463227 | 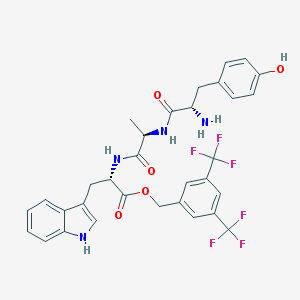 CHEMBL3609619 CHEMBL3609619 | C32H30F6N4O5 | 664.605 | 12 / 5 | 4.5 | No |
| 3169 | 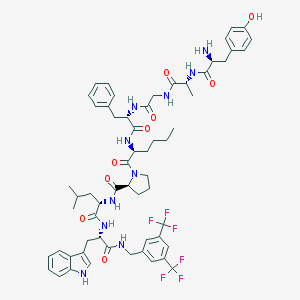 CHEMBL557188 CHEMBL557188 | C60H72F6N10O9 | 1191.29 | 16 / 10 | 7.4 | No |
| 3241 | 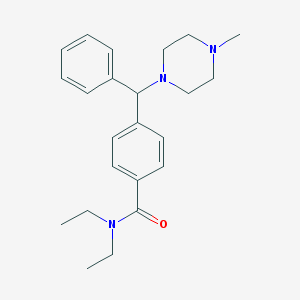 CHEMBL124642 CHEMBL124642 | C23H31N3O | 365.521 | 3 / 0 | 3.5 | Yes |
| 547942 | 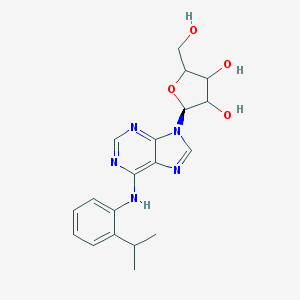 CHEMBL95302 CHEMBL95302 | C19H23N5O4 | 385.424 | 8 / 4 | 2.3 | Yes |
| 3553 | 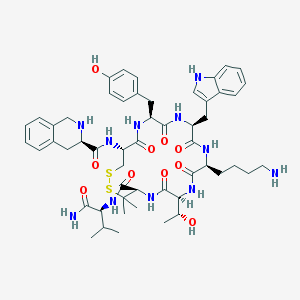 CHEMBL405672 CHEMBL405672 | C53H71N11O10S2 | 1086.34 | 14 / 13 | 2.5 | No |
| 3569 | 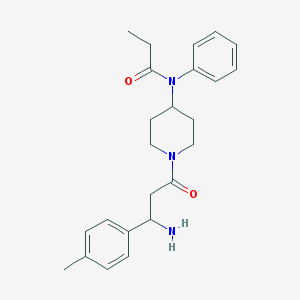 CHEMBL231049 CHEMBL231049 | C24H31N3O2 | 393.531 | 3 / 1 | 2.7 | Yes |
| 3719 | 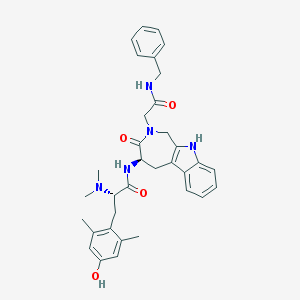 CHEMBL451962 CHEMBL451962 | C34H39N5O4 | 581.717 | 5 / 4 | 4.1 | No |
| 3901 | 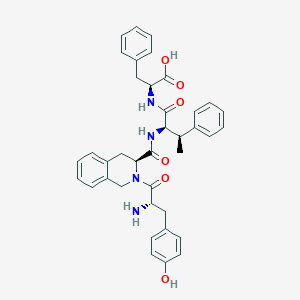 CHEMBL2371018 CHEMBL2371018 | C38H40N4O6 | 648.76 | 7 / 5 | 2.3 | No |
| 3904 | 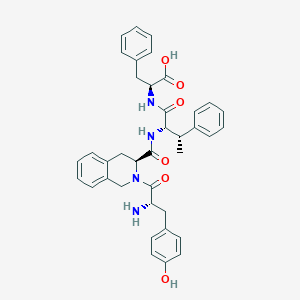 CHEMBL2371027 CHEMBL2371027 | C38H40N4O6 | 648.76 | 7 / 5 | 2.3 | No |
| 3946 | 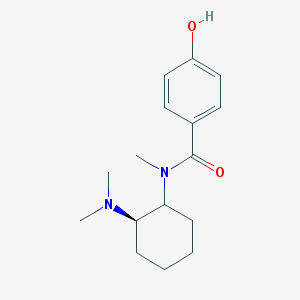 CHEMBL298893 CHEMBL298893 | C16H24N2O2 | 276.38 | 3 / 1 | 2.4 | Yes |
| 3947 | 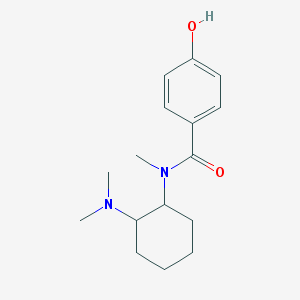 CHEMBL609676 CHEMBL609676 | C16H24N2O2 | 276.38 | 3 / 1 | 2.4 | Yes |
| 4453 | 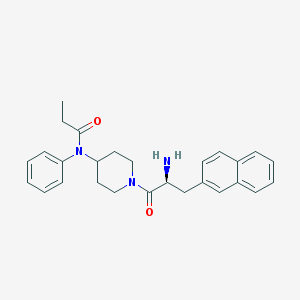 CHEMBL230943 CHEMBL230943 | C27H31N3O2 | 429.564 | 3 / 1 | 4.1 | Yes |
| 4479 | 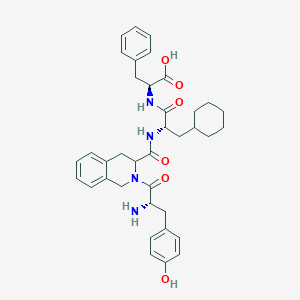 H-Tyr-Tic-Cha-Phe-OH H-Tyr-Tic-Cha-Phe-OH | C37H44N4O6 | 640.781 | 7 / 5 | 3.1 | No |
| 4482 | 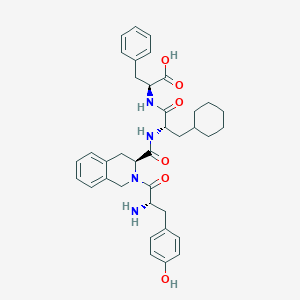 CHEMBL2371935 CHEMBL2371935 | C37H44N4O6 | 640.781 | 7 / 5 | 3.1 | No |
| 4825 | 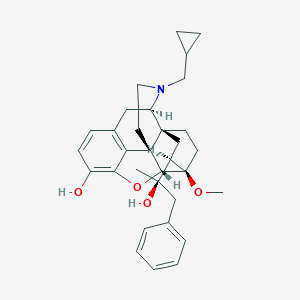 CHEMBL2338752 CHEMBL2338752 | C32H39NO4 | 501.667 | 5 / 2 | 5.2 | No |
| 5321 | 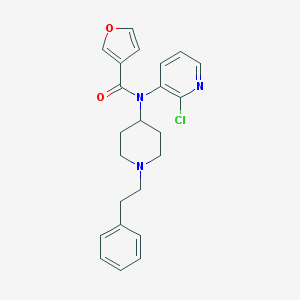 CHEMBL161147 CHEMBL161147 | C23H24ClN3O2 | 409.914 | 4 / 0 | 4.4 | Yes |
| 5387 | 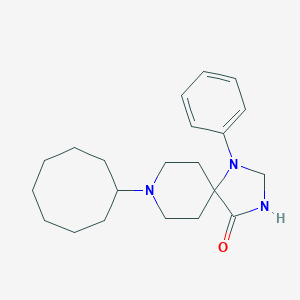 CHEMBL353990 CHEMBL353990 | C21H31N3O | 341.499 | 3 / 1 | 4.2 | Yes |
| 463567 | 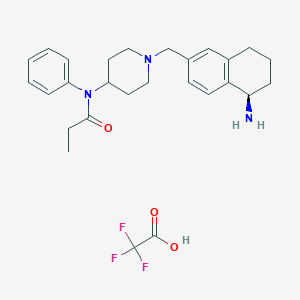 CHEMBL3613884 CHEMBL3613884 | C27H34F3N3O3 | 505.582 | 8 / 2 | N/A | No |
| 5837 | 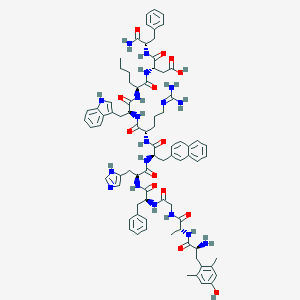 BDBM50321606 BDBM50321606 | C80H98N18O14 | 1535.78 | 17 / 18 | 2.2 | No |
| 5871 | 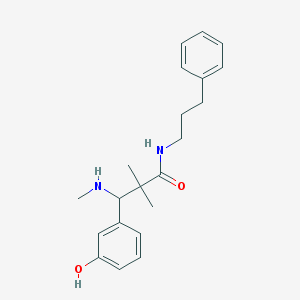 CHEMBL347750 CHEMBL347750 | C21H28N2O2 | 340.467 | 3 / 3 | 3.5 | Yes |
| 5959 | 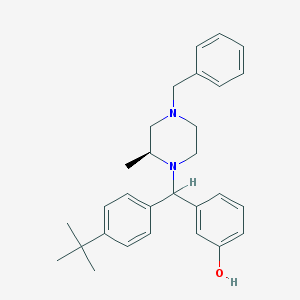 CHEMBL151416 CHEMBL151416 | C29H36N2O | 428.62 | 3 / 1 | 6.5 | No |
| 6029 | 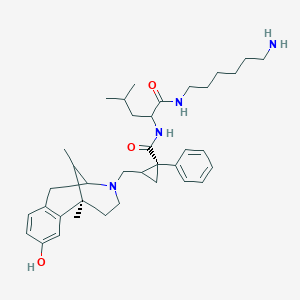 CHEMBL101072 CHEMBL101072 | C37H54N4O3 | 602.864 | 5 / 4 | 5.9 | No |
| 6397 | 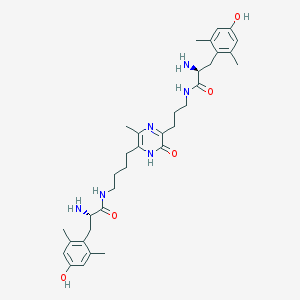 CHEMBL385779 CHEMBL385779 | C34H48N6O5 | 620.795 | 8 / 7 | 2.5 | No |
| 6492 | 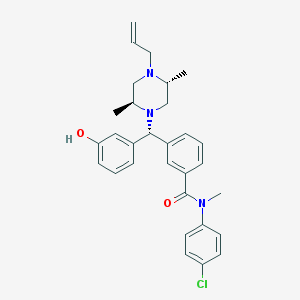 CHEMBL155797 CHEMBL155797 | C30H34ClN3O2 | 504.071 | 4 / 1 | 6.1 | No |
| 521644 | 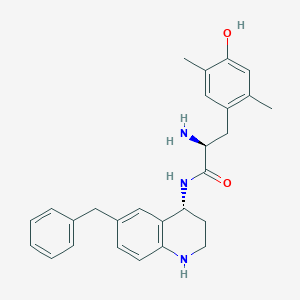 CHEMBL3770677 CHEMBL3770677 | C27H31N3O2 | 429.564 | 4 / 4 | 4.6 | Yes |
| 6950 | 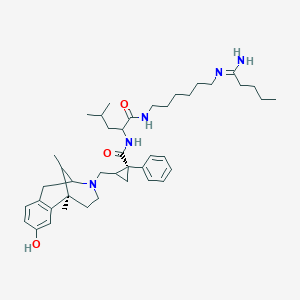 CHEMBL102013 CHEMBL102013 | C42H63N5O3 | 685.998 | 5 / 4 | 7.3 | No |
| 6966 | 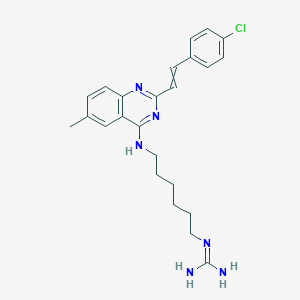 CID 69083551 CID 69083551 | C24H29ClN6 | 436.988 | 4 / 3 | 5.0 | Yes |
| 7461 | 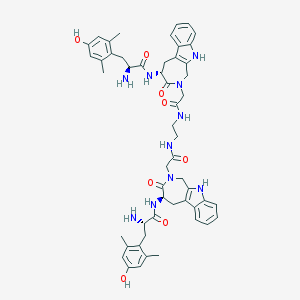 CHEMBL1076738 CHEMBL1076738 | C52H60N10O8 | 953.114 | 10 / 10 | 3.0 | No |
| 7665 | 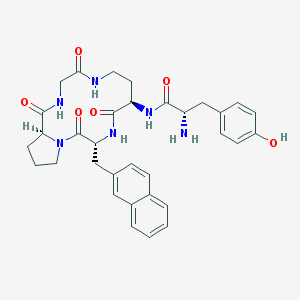 CHEMBL278889 CHEMBL278889 | C33H38N6O6 | 614.703 | 7 / 6 | 1.8 | No |
| 7948 | 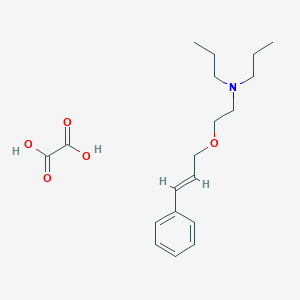 CHEMBL2048983 CHEMBL2048983 | C19H29NO5 | 351.443 | 6 / 2 | N/A | N/A |
| 8144 | 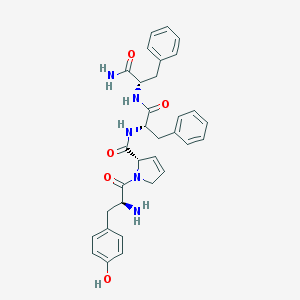 CHEMBL557243 CHEMBL557243 | C32H35N5O5 | 569.662 | 6 / 5 | 2.5 | No |
| 8361 | 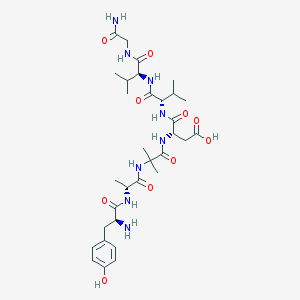 CHEMBL315603 CHEMBL315603 | C32H50N8O10 | 706.798 | 11 / 10 | -3.1 | No |
| 8565 | 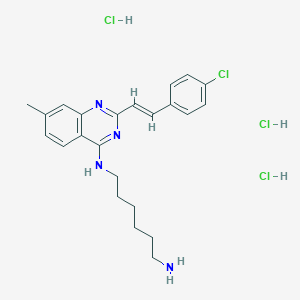 CHEMBL3216028 CHEMBL3216028 | C23H30Cl4N4 | 504.321 | 4 / 5 | N/A | No |
| 8751 | 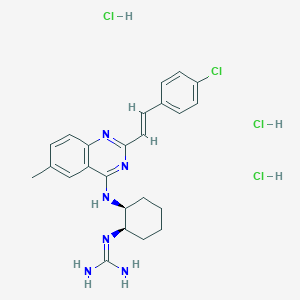 CHEMBL3217121 CHEMBL3217121 | C24H30Cl4N6 | 544.346 | 4 / 6 | N/A | No |
| 8889 | 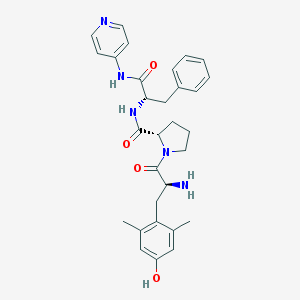 CHEMBL409907 CHEMBL409907 | C30H35N5O4 | 529.641 | 6 / 4 | 2.9 | No |
| 9156 | 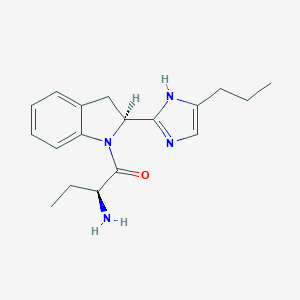 CHEMBL188565 CHEMBL188565 | C18H24N4O | 312.417 | 3 / 2 | 2.2 | Yes |
| 9525 | 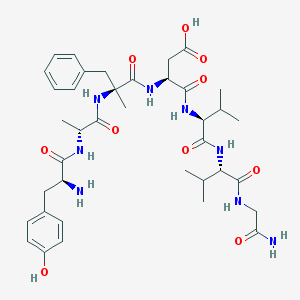 CHEMBL121946 CHEMBL121946 | C38H54N8O10 | 782.896 | 11 / 10 | -1.5 | No |
| 9639 | 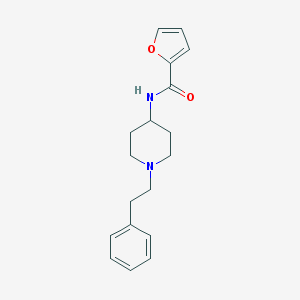 CHEMBL161512 CHEMBL161512 | C18H22N2O2 | 298.386 | 3 / 1 | 3.1 | Yes |
| 9662 | 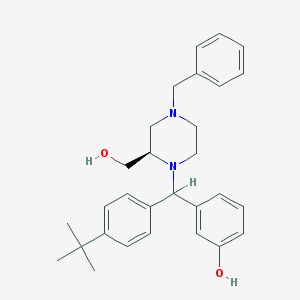 CHEMBL149795 CHEMBL149795 | C29H36N2O2 | 444.619 | 4 / 2 | 5.5 | No |
| 9918 | 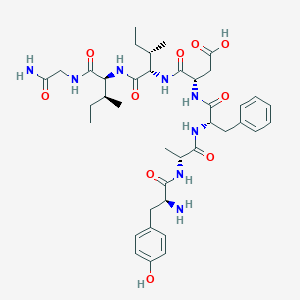 CHEMBL175116 CHEMBL175116 | C39H56N8O10 | 796.923 | 11 / 10 | -1.0 | No |
| 9957 | 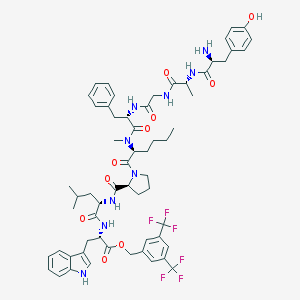 CHEMBL387670 CHEMBL387670 | C61H73F6N9O10 | 1206.3 | 17 / 8 | 8.1 | No |
| 10055 | 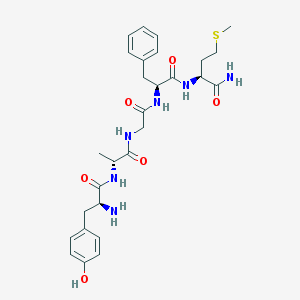 H-Tyr-D-Ala-Gly-Phe-Met-NH2 H-Tyr-D-Ala-Gly-Phe-Met-NH2 | C28H38N6O6S | 586.708 | 8 / 7 | -0.1 | No |
| 11252 | 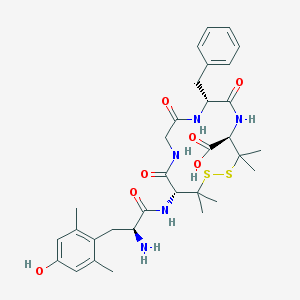 CHEMBL159793 CHEMBL159793 | C32H43N5O7S2 | 673.844 | 10 / 7 | 0.1 | No |
| 11261 | 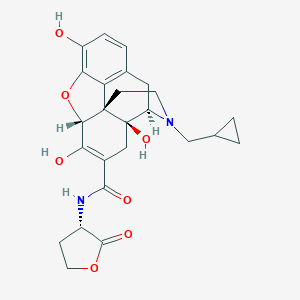 SCHEMBL9880436 SCHEMBL9880436 | C25H28N2O7 | 468.506 | 8 / 4 | 1.3 | Yes |
| 11303 | 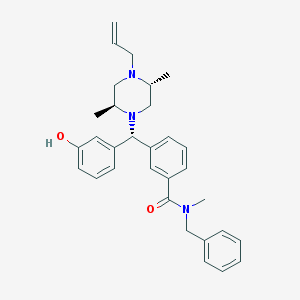 CHEMBL422877 CHEMBL422877 | C31H37N3O2 | 483.656 | 4 / 1 | 5.4 | No |
| 11330 | 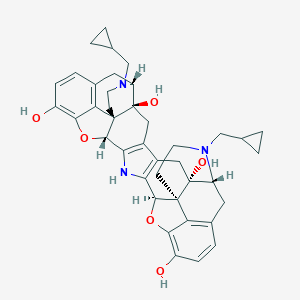 Norbinaltorphimine Norbinaltorphimine | C40H43N3O6 | 661.799 | 8 / 5 | 3.2 | No |
| 11349 | 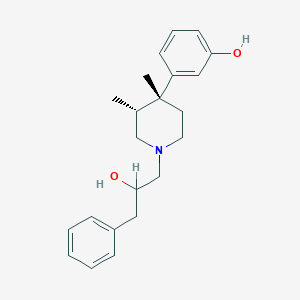 CHEMBL93614 CHEMBL93614 | C22H29NO2 | 339.479 | 3 / 2 | 4.5 | Yes |
| 11612 | 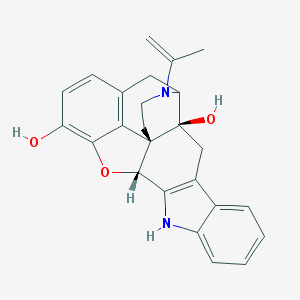 BDBM50098673 BDBM50098673 | C25H24N2O3 | 400.478 | 4 / 3 | 3.7 | Yes |
| 11615 | 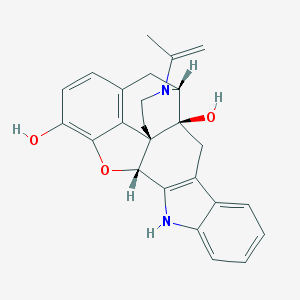 CHEMBL611697 CHEMBL611697 | C25H24N2O3 | 400.478 | 4 / 3 | 3.7 | Yes |
| 11983 | 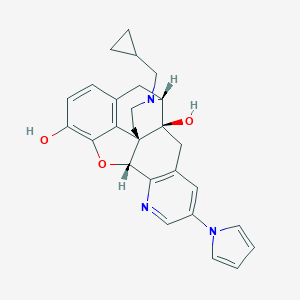 CHEMBL346165 CHEMBL346165 | C27H27N3O3 | 441.531 | 5 / 2 | 2.9 | Yes |
| 12085 | 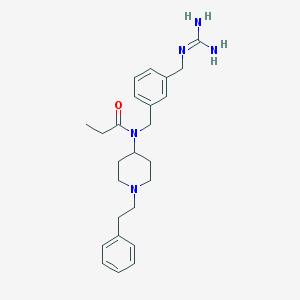 CHEMBL1089542 CHEMBL1089542 | C25H35N5O | 421.589 | 3 / 2 | 2.7 | Yes |
| 12155 | 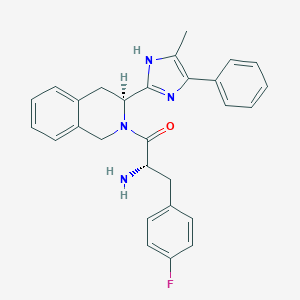 CHEMBL381793 CHEMBL381793 | C28H27FN4O | 454.549 | 4 / 2 | 4.2 | Yes |
| 12189 | 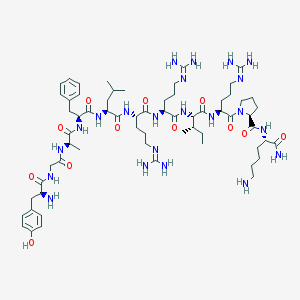 CHEMBL415795 CHEMBL415795 | C64H106N22O12 | 1375.69 | 17 / 19 | -2.4 | No |
| 12194 | 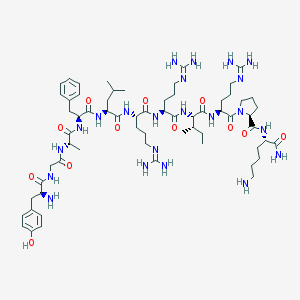 CHEMBL262963 CHEMBL262963 | C64H106N22O12 | 1375.69 | 17 / 19 | -2.4 | No |
| 464402 | 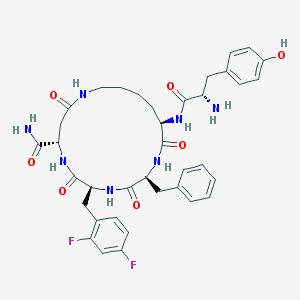 CHEMBL3580743 CHEMBL3580743 | C37H43F2N7O7 | 735.79 | 10 / 8 | 1.6 | No |
| 12984 | 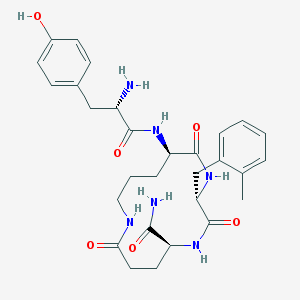 CHEMBL323024 CHEMBL323024 | C29H38N6O6 | 566.659 | 7 / 7 | 0.4 | No |
| 13229 | 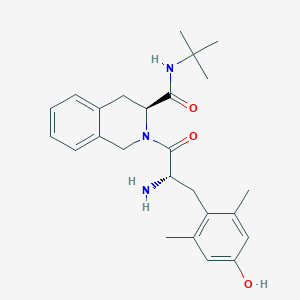 CHEMBL146108 CHEMBL146108 | C25H33N3O3 | 423.557 | 4 / 3 | 3.1 | Yes |
| 13378 | 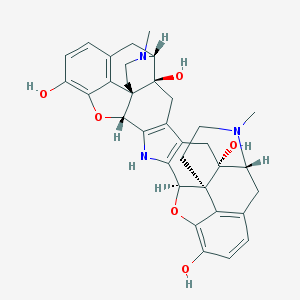 CHEMBL606959 CHEMBL606959 | C34H35N3O6 | 581.669 | 8 / 5 | 1.7 | No |
| 13435 | 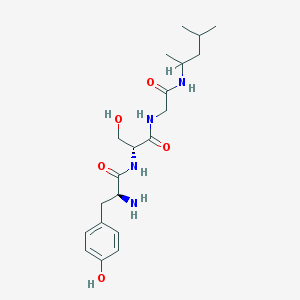 CHEMBL2370599 CHEMBL2370599 | C20H32N4O5 | 408.499 | 6 / 6 | 0.4 | No |
| 536348 | 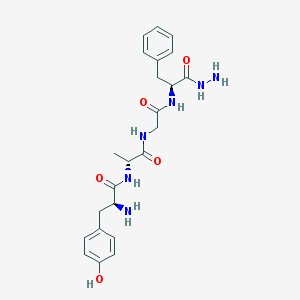 CHEMBL91760 CHEMBL91760 | C23H30N6O5 | 470.53 | 7 / 7 | -0.8 | No |
| 13720 | 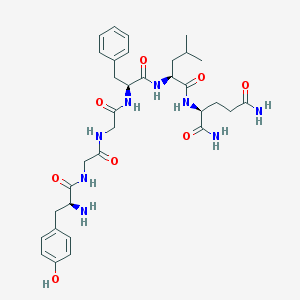 CHEMBL132775 CHEMBL132775 | C33H46N8O8 | 682.779 | 9 / 9 | -0.9 | No |
| 521855 | 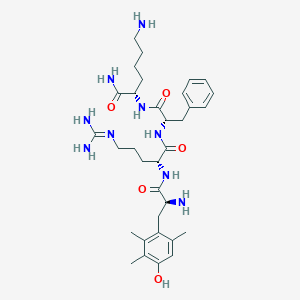 CHEMBL3823054 CHEMBL3823054 | C33H51N9O5 | 653.829 | 8 / 9 | 0.4 | No |
| 14025 | 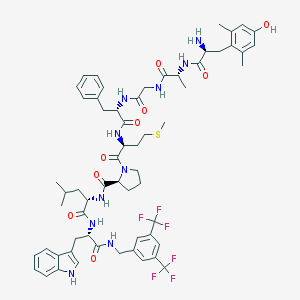 CHEMBL1766208 CHEMBL1766208 | C61H74F6N10O9S | 1237.37 | 17 / 10 | 7.3 | No |
| 14085 | 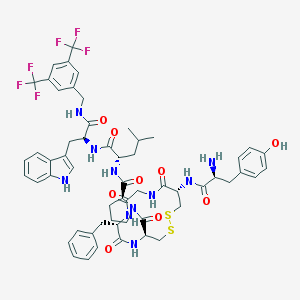 CHEMBL611925 CHEMBL611925 | C57H64F6N10O9S2 | 1211.31 | 18 / 10 | 6.0 | No |
| 14090 | 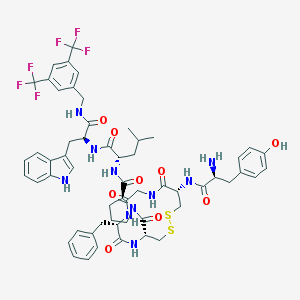 CHEMBL611924 CHEMBL611924 | C57H64F6N10O9S2 | 1211.31 | 18 / 10 | 6.0 | No |
| 14189 | 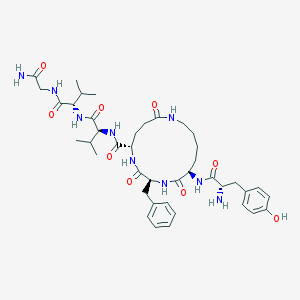 CHEMBL132164 CHEMBL132164 | C41H59N9O9 | 821.977 | 10 / 10 | 1.2 | No |
| 464635 | 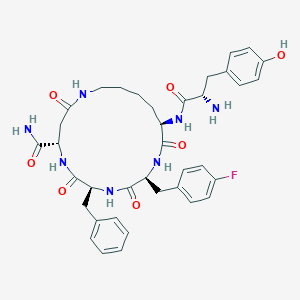 CHEMBL3582484 CHEMBL3582484 | C37H44FN7O7 | 717.799 | 9 / 8 | 1.5 | No |
| 14428 | 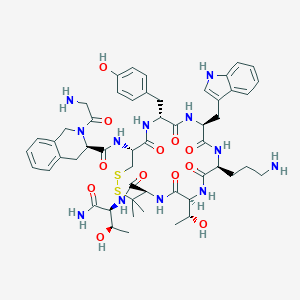 CHEMBL2369473 CHEMBL2369473 | C53H70N12O12S2 | 1131.33 | 16 / 14 | -0.4 | No |
| 14589 | 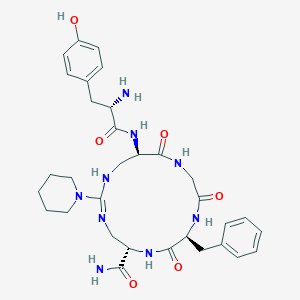 CHEMBL2402930 CHEMBL2402930 | C32H43N9O6 | 649.753 | 8 / 8 | -0.8 | No |
| 14596 | 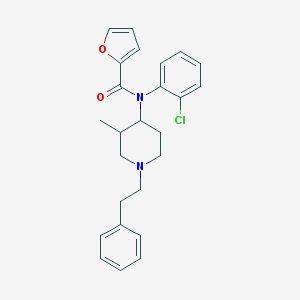 CHEMBL327270 CHEMBL327270 | C25H27ClN2O2 | 422.953 | 3 / 0 | 6.0 | No |
| 15134 | 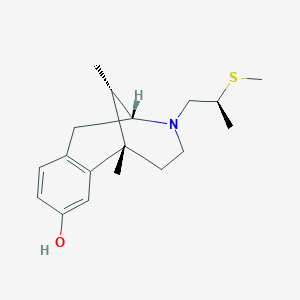 CHEMBL99496 CHEMBL99496 | C18H27NOS | 305.48 | 3 / 1 | 2.9 | Yes |
| 15135 | 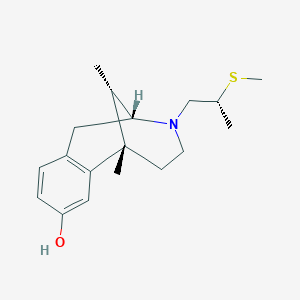 CHEMBL97918 CHEMBL97918 | C18H27NOS | 305.48 | 3 / 1 | 2.9 | Yes |
| 15136 | 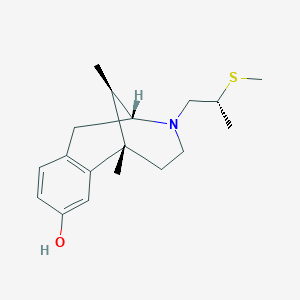 CHEMBL98817 CHEMBL98817 | C18H27NOS | 305.48 | 3 / 1 | 2.9 | Yes |
| 15137 | 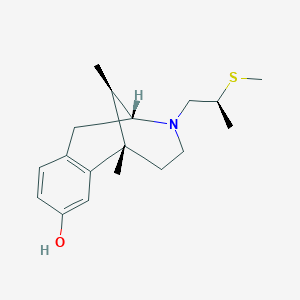 CHEMBL98650 CHEMBL98650 | C18H27NOS | 305.48 | 3 / 1 | 2.9 | Yes |
| 15523 | 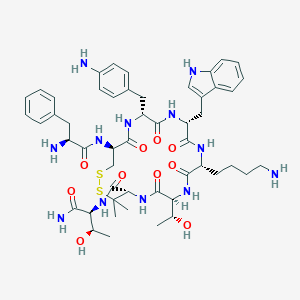 CHEMBL2371213 CHEMBL2371213 | C51H70N12O10S2 | 1075.31 | 15 / 14 | 0.5 | No |
| 15567 | 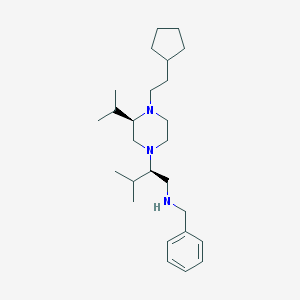 CHEMBL572875 CHEMBL572875 | C26H45N3 | 399.667 | 3 / 1 | 6.5 | No |
| 15716 | 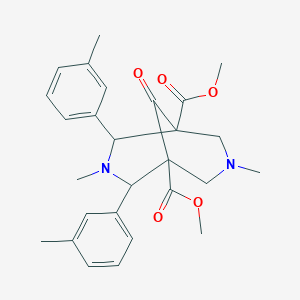 CHEMBL179350 CHEMBL179350 | C27H32N2O5 | 464.562 | 7 / 0 | 3.2 | Yes |
| 15747 | 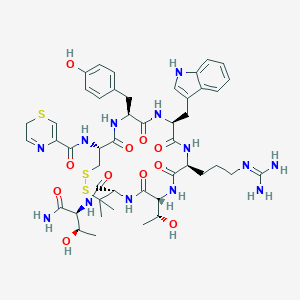 BDBM50085059 BDBM50085059 | C47H63N13O11S3 | 1082.28 | 16 / 14 | -1.0 | No |
| 15852 | 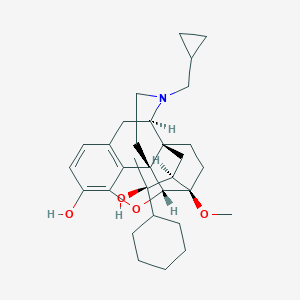 CHEMBL2338749 CHEMBL2338749 | C31H43NO4 | 493.688 | 5 / 2 | 5.6 | No |
| 16132 | 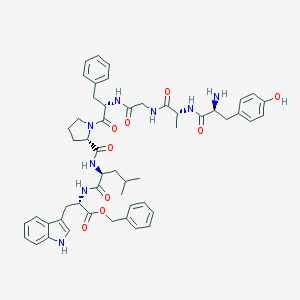 H-Tyr-D-Ala-Gly-Phe-Pro-Leu-Trp-O-Bzl H-Tyr-D-Ala-Gly-Phe-Pro-Leu-Trp-O-Bzl | C52H62N8O9 | 943.115 | 10 / 8 | 5.0 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218